INTRODUCTION Aronia melanocarpa also known as black chokeberry, belongs to the genus Aronia (Rosaceae). In addition to A. melanocarpa, Aronia includes A. arbutifolia or red chokeberry and A. prunifolia or purple chokeberry, both distributed naturally in North American, and an additional cultivated taxon, A. mitschurinii or Mitschurin’s chokeberry, originating from Europe. However, the species boundaries and relationships among the species of Aronia are not clear. Moreover, the taxonomic history of Aroniais complex, as species of this genus have formerly been placed in many different genera, such as Mespilus, Pyrus, Adenorachis, Sorbus, and Photinia. In the present study, we first sequenced and characterized the complete chloroplast (cp) genome of A. melanocarpa and compared its sequence with those of the cp genomes from 13 species of the family Rosaceae. The aims of this study were: (1) to increase our understanding of the structural patterns of complete cp genome of A. melanocarpa; (2) to investigate the phylogenetic relationships of A. melanocarpa with other Rosaceae species based on their cp genomes.
RATIONALE The chloroplast is a unique and essential organelle in green plants with vital roles in photosynthesis and carbon fixation. Comparative analyses of cp genomes between different plant species reveal intra- and inter-species rearrangements that have occurred during evolution, such as inverted repeat (IR) contraction and expansion. Based on these characteristics, the cp genome has been wildly used for species identification, phylogenetic analysis, and exploring the genetic basis of environmental adaptation.
RESULTS The complete A. melanocarpa cp genome was sequenced, analyzed, and compared with that from 13 other species in the Rosaceae. The cp genome is 159 772 bp and has a total guanine-cytosine (GC) content of 36.6%. It exhibits a typical quadripartite structure with four separate regions, including a large single copy (LSC) region of 87 810 bp and a small single copy (SSC) region of 19 200 bp separated by two inverted repeats (IRa and IRb) regions of 26 381 bp each. A total of 132 genes were annotated, including 87 protein-coding genes, 37 tRNAs, and eight rRNAs, with 22 duplicates in the IR regions. In total, 76 simple sequence repeats (SSRs) and 50 long repeats were detected. Phylogenetic analysis indicated that A. melanocarpa is most closely related to A. arbutifolia and forms a sister clade to Cydonia oblonga with weak support.
CONCLUSION We analyzed the complete cp genome of A. melanocarpa by using Illumina high-throughput sequencing technology. The sequence of A. melanocarpa cp genome could be further used for the development of molecular markers. Highly variable regions were detected in intergenic regions, such as trnK-rps16, rps16-trnQ, trnG-atpA, petN-psbM, trnT-psbD, psbZ-trnG, trnT-trnL, ndhC-trnV and accD-psaI, which might be useful for broad applications in genetic research studies as well as phylogenetic studies. Phylogenetic construction results strongly supported that A. melanocarpa was closest related to A. arbutifolia, followed by C. oblonga with weak support. This newly available genomic data for A. melanocarpa will provide a basis for future research on the population genetics and phylogenomics and will benefit the breeding studies and utilization of the genus Aronia.
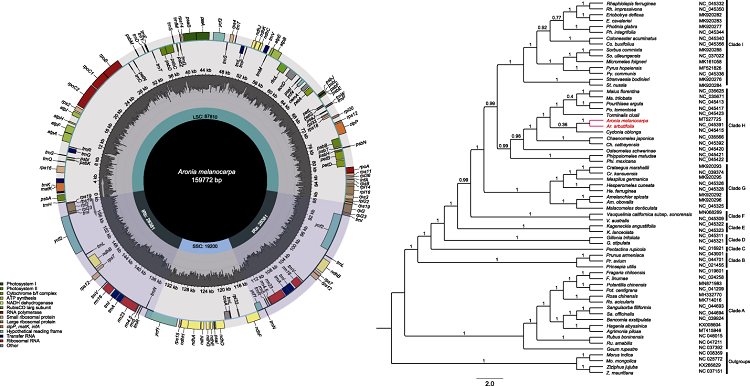
Map of the chloroplast genome of Aronia melanocarpa and phylogenetic analyses among the 60 Rosaceae species using their complete chloroplast genomes. Aronia formed a clade with Dichotomanthes and Pourthiaea based on cpDNA tree. Moreover, A. melanocarpa is most closely related to A. arbutifolia and forms a sister clade to Cydonia oblonga with weak support.

