-
Hosted by:Chinese Academy of Sciences
Sponsored by:Institute of Botany, Chinese Academy of Sciences, Botanical Society of China
Co-hosted by:Key Laboratory of Soybean Molecular Design Breeding, Northeast Institute of Geography and Agroecology, Chinese Academy of Sciences
Institute of Biotechnology and Germplasm Resources, Yunnan AgriculturalAcademy
Fujian Agriculture and Forestry University
Hunan Provincial Key Laboratory of Phytohormones and Growth Development, Hunan Agricultural University
State Key Laboratory of Crops Biology, Shandong Agricultural University

WeChat:zwxb_2009
 Table of Content
Table of Content- Establishment of Regeneration System in vitro for Hydrangea macrophylla cv. ‘Chikushi-no-kaze’
- Yuyan Jin, Shuangshuang Chen, Jing Feng, Xintong Liu, Xiangyu Qi, Huijie Chen, Yan Dong, Yanming Deng
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25068 cstr: 32102.14.CBB25068
-
 Abstract
(
316 )
Abstract
(
316 )
 PDF (1066KB)
(
178
)
PDF (1066KB)
(
178
)
 Save
Save
- References | Related Articles | Metrics
- Establishment of a Regeneration System for Changnienia amoena
- Shengfei Yang, Yuye Deng, Shiyun Cai, Yafei Liu, Yin Peng, Yuanjie Ding
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25081 cstr: 32102.14.CBB25081
-
 Abstract
(
308 )
Abstract
(
308 )
 PDF (1646KB)
(
1281
)
PDF (1646KB)
(
1281
)
 Save
Save
- References | Related Articles | Metrics
- The Mechanism of Manganese Accumulation Mediated by SpMTP10 Isolated from Sedum plumbizincicola
- Siying Chen, Jinglin Wang, Yingyi Li, Xiangxin Lu, Peihong Zhang, Qinghong Qiu, Yan Gao, Tianyu Gu, Jiashi Peng
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25053 cstr: 32102.14.CBB25053
-
 Abstract
(
195 )
Abstract
(
195 )
 PDF (2180KB)
(
760
)
PDF (2180KB)
(
760
)
 Save
Save
- References | Related Articles | Metrics
- Genome-wide Identification of the AP2/ERF Gene Family and Its Expression Patterns under Salt Stress in the Desert Plant Reaumuria soongarica
- Mengxuan Zhu, Yuan Liu, Haoyu Zhao, Zhenyun Sun, Wei Gong, Zhenhua Dang
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25066
-
 Abstract
(
331 )
Abstract
(
331 )
 PDF (1892KB)
(
462
)
PDF (1892KB)
(
462
)
 Save
Save
- References | Related Articles | Metrics
- Comparative Analysis of Transcriptome of Adventitious Roots under Different Hydroponic Conditions of Lycium barbarum
- Yang Gaier, Zhang Xuan, Wang Jiadong, Zhang Bo, Duan Linyuan, Li Xiang
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25077 cstr: 32102.14.CBB25077
-
 Abstract
(
356 )
Abstract
(
356 )
 PDF (1960KB)
(
4482
)
PDF (1960KB)
(
4482
)
 Save
Save
- Related Articles | Metrics
- Analysis of the Function of WRI1 in Heat Stress of Arabidopsis Seedlings
- Xuya Gu, Zhangman Lin, Siyuan Hu, Xuening Li, Xiaolin Qin, Zhengxi Wu, Nuo Li, Mingyin Feng, Ruihua Huang
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25085 cstr: 32102.14.CBB25085
-
 Abstract
(
365 )
Abstract
(
365 )
 PDF (1550KB)
(
550
)
PDF (1550KB)
(
550
)
 Save
Save
- References | Related Articles | Metrics
-
Effect of Salt Stress on Carbohydrate Secretion by the Root System of Thinopyrum ponticum
- Congcong Yi, Dan Li, Yanjie Li, Xiuwen Lin, Yingshuai Lu, Xiaopeng Chen
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25046 cstr: 32102.14.CBB25046
-
 Abstract
(
262 )
Abstract
(
262 )
 PDF (1122KB)
(
94
)
PDF (1122KB)
(
94
)
 Save
Save
- Related Articles | Metrics
- Molecular Mechanisms of Glycosyltransferases CsUGT73B4 and CsUGT85K5 in Tea Plants in Response to Cold Stress
- Gai Xinyue, Wang Xiaodong, Fan Yangen, Li Bin, Fu Xiaodong, Sun Ping, Huang Xiaoqin
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25087
-
 Abstract
(
334 )
Abstract
(
334 )
 PDF (2009KB)
(
115
)
PDF (2009KB)
(
115
)
 Save
Save
- Related Articles | Metrics
- Cloning and Functional Validation of the Key Gene LbCER3 Involved in Waxy Cuticle Synthesis of Lycium barbarum
- Juanhong Zhao, Zhigang Li, Guoqi Zheng, Zhanfei Liu, Lirong Kou, Wenxiu Wang, Tiantian Zhou, Juan Yang
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25132 cstr: 32102.14.CBB25132
-
 Abstract
(
164 )
Abstract
(
164 )
 PDF (2401KB)
(
528
)
PDF (2401KB)
(
528
)
 Save
Save
- Related Articles | Metrics
-
The Regulatory Network and Mechanism of Plant Floral Organ Development Mediated by Auxin Response Factors
- Xinyu Liu, Silan Dai
- Chinese Bulletin of Botany. 2026, 61(3): 1-0. doi: 10.11983/CBB25169
-
 Abstract
(
239 )
Abstract
(
239 )
 PDF (1627KB)
(
220
)
PDF (1627KB)
(
220
)
 Save
Save
- Related Articles | Metrics
INTRODUCTION: Hydrangea is essential in landscaping, ecology, and medical care, with significant development prospects. ‘Chikushi-no-kaze’ is an ideal low-maintenance variety for micro-landscaping and potted Hydrangea, widely favored by consumers. However, the extremely low setting rate of Hydrangea and poor seed development under natural conditions render traditional reproduction methods inadequate for meeting the demands of large-scale annual production in the market. The breeding of Hydrangea plantlets through tissue culture technology is currently the most efficient method for producing high-quality plantlets.
RATIONALE: Regeneration efficiency in plant tissue culture is a key factor in achieving factory seedling production. Therefore, this study investigated the regeneration efficiency of isolated leaves from tissue culture plantlets of ‘Chikushi-no-kaze’ under optimal culture conditions at each key stage, considering different leaf positions, dark culture durations, and other factors. The aim was to establish an efficient regeneration technology system, which provides technical guidance for large-scale plantlet production and serves as a reference for establishing a genetic transformation system for Hydrangea in the future.
RESULTS: The 3rd to 5th leaves (middle mature leaves) of ‘Chikushi-no-kaze’ tissue culture plantlets were identified as the optimal sampling leaves. A dark culture duration of 10-14 days was conducive to callus formation. The most suitable medium for the induction and regeneration of adventitious buds was MS + 3.0 mg·L-1 CPPU + 0.1 mg·L-12,4-D, with induction and regeneration rates of 97.78% and 93.33%, respectively. The optimal medium for adventitious bud proliferation was MS + 2.0 mg·L-1 6-BA + 0.1 mg·L-1 IBA, yielding a proliferation coefficient of 8.33. For elongation growth, the optimal medium was MS + 1.0 mg·L-1 6-BA + 0.1 mg·L-1 IBA, resulting in an average stem length of 4.10 cm. The optimal medium for rooting culture was MS + 0.3 mg·L-1 IBA, achieving a rooting rate of 87.20%.
CONCLUSION: This study initially established a technical system for in vitro leaf regeneration of big leaf Hydrangea 'Chikushi-no-kaze', which effectively solved the problem of low efficiency of adventitious bud regeneration of Hydrangea, helped to achieve efficient reproduction and recycling. Lay the foundation for large-scale production and genetic improvement of Hydrangea.
INTRODUCTION: Changnienia amoena is listed as a national second-class protected plant. It is a rare orchid species unique to China, as well as a valuable medicinal plant and a potential wild potted flower germplasm resource with great development potential. Currently, wild resources of C. amoena are rapidly declining, making conservation efforts crucial.
RATIONALE: Research on C. amoena regeneration technology has mainly focused on different combinations of plant growth regulators, and a complete regeneration system has not yet been established. Tissue culture technology plays a crucial role in the conservation of wild C. amoena resources, and research on its regeneration system can promote the sustainable development and utilization of these resources.
RESULTS: The results showed that: (1) 75% ethanol had a significant effect on the sterilization of C. amoena pseudobulbs, and the best sterilization treatment for C. amoena pseudobulbs was to use 75% ethanol for 30 seconds, followed by 0.1% mercuric acid solution for 12 minutes; (2) The pseudobulbs were able to induce adventitious buds under the different blocking treatments, but the effect of the adventitious buds was that the complete pseudobulbs> the halved pseudobulbs> the cruciform block pseudobulbs and the rate of germination was much higher than that of the blocking treatment; (3) Considering the browning rate, contamination rate, survival rate and induced germination rate, the optimal collection time for C. amoena pseudobulbs to be used as explants for adventitious shoots induced in May; (4) The optimal medium for the adventitious shoots of C. amoena pseudobulbs was 1/2 MS+1.0mg∙L–1 6-BA+ 0.5mg∙L–1 NAA, with an induction rate of 75.56%; the optimal rooting medium is1/2 MS+0.4 mg∙L–1 6-BA+1.0 mg∙L–1 NAA, with a rooting rate of up to 93.33%; (5) After treatment with a 50 mg∙L–1 6-BA solution, the histocultured plantlets were transplanted into humus soil and covered with plastic film for hardening. The survival rate was as high as 94.44%.
CONCLUSION: In this study, the pseudobulbs of C. amoena were used as explants to explore the effects of sterilization conditions, the way of explant cutting, different collection times, and the concentration of plant growth regulators on the sterilization effect of explants, inducing adventitious shoots and rooting of histocultured plantlets, and to conduct a preliminary study on the regeneration system of C. amoena, which can help to protect the germplasm resources of C. amoena, and provide a theoretical basis and technical references for the future propagation of C. amoena.
INTRODUCTION: Manganese (Mn) is an essential micronutrient for plant growth and primarily act as enzyme cofactors and participate in the redox processes. However, excessive absorption of Mn by plant can also induce toxicity damages. Therefore, plants need to tightly regulate the uptake, homeostasis, and distribution of Mn to cope with stresses caused by its deficiency or excess. In these processes, the cation diffusion facilitator (CDF) family transporters, which in plants are also known as metal tolerance proteins (MTP), had been shown to be crucial for Mn homeostasis. Therefore, identifying MTP family genes and elucidating their underlying mechanisms for Mn accumulation would not only provide the novel insights about basic scientific issues of plant Mn accumulation, but also gene resources for crops improvement and Mn pollution bioremediation.
RATIONALE: Sedum plumbizincicola is a recently discovered Cd/Zn hyperaccumulator that grows in mining areas. The soil in its natural habitat contains more than 10 000 mg·kg-1 of Mn, suggesting that S. plumbizincicola may have efficient Mn transport and detoxification capabilities. Based on the transcriptome sequencing results of S. plumbizincicola obtained previously, a member of the MTP family gene named SpMTP10 was cloned and its role in mediating Mn accumulation was investigated in this study.
RESULTS: Phylogenetic analysis with orthologs from Arabidopsis and rice revealed that SpMTP10 belongs to the Mn-CDF subfamily and is most closely related to AtMTP10, AtMTP9 and OsMTP9, with the highest sequence identity of 72% to AtMTP10. SpMTP10 is mainly expressed in the roots of S. plumbizincicola and its expression level is not affected by Mn treatment. Expression of SpMTP10 in yeast can greatly enhance the tolerance of transformants to excessive Mn stress, and increase the Mn accumulation in transformants. However, under conditions of excessive cadmium (Cd), zinc (Zn), copper (Cu), and iron (Fe) stress, the yeast transformants exhibited no significant changes in tolerance. Subsequent subcellular localization analysis revealed that SpMTP10 was localized to the endoplasmic reticulum (ER) membrane. Compared with wild-type plants, transgenic Arabidopsis overexpressing SpMTP10 demonstrated reduced Mn accumulation in roots but increased Mn accumulation in shoots, rendering the plants more sensitive to excessive Mn stress.
CONCLUSION: In conclusion, SpMTP10 likely enhances yeast tolerance to excessive Mn toxicity by promoting Mn sequestration in the ER. In plants, Mn transport mediated by SpMTP10 into the ER may facilitate intercellular migration of Mn in the ER lumen via plasmodesmata, thereby promoting Mn movement toward vascular tissues in roots and subsequent long-distance transport to shoots.
INTRODUCTION: Reaumuria soongorica, a perennial small shrub, is widely distributed in arid desert regions of northwestern China. It exhibits exceptional drought and salt tolerance, making it an ideal model for studying molecular mechanisms of plant stress resistance.
RATIONALE: We used the R. soongorica genome as a reference to identify members of the AP2/ERF gene family in this species, and analyzed their phylogeny, gene structure, conserved motifs, cis-acting elements, gene duplication events, as well as the expression patterns of these family members under salt stress.
RESULTS: Seventy AP2/ERF genes were identified from the R. soongorica genome. Phylogenetic analysis classified these genes into four subfamilies: AP2, ERF, DREB, and RAV. Cis-acting element analysis revealed multiple regulatory elements associated with light responsiveness, stress adaptation, growth regulation, and hormone signaling in the promoter regions of R. soongorica AP2/ERF genes. These genes exhibited an uneven distribution across all 11 chromosomes of the R. soongorica genome, with 63 genes (90% of the total) originating from gene duplication events. Evolutionary analysis suggested that whole-genome duplication (WGD) and dispersed duplication were the primary drivers of family expansion. Under salt stress, AP2/ERF genes showed divergent expression patterns in R. soongorica seedlings, with six genes displaying significant differential expression (|log2FC|≥1, P<0.05), implicating their potential roles in salt stress response.
CONCLUSION: This study identified and characterized the AP2/ERF gene family in R. soongorica at the genomic level, thereby establishing a foundation for elucidating its functional roles in this species' adaptation to arid and saline environments.
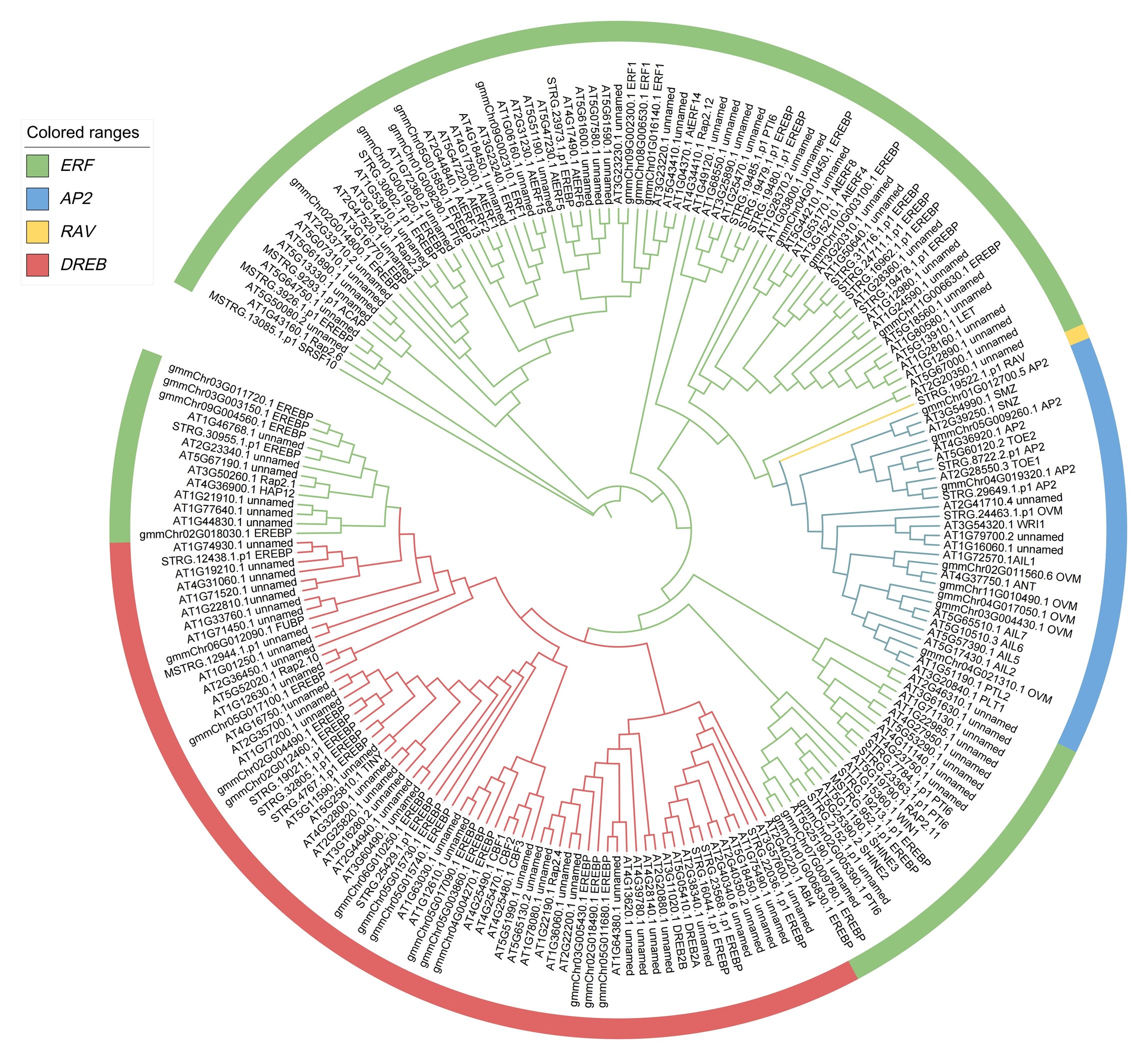
Phylogenetic tree of the AP2/ERF gene family in R. soongorica.
INTRODUCTION: This study investigates the transcriptional levels of adventitious root formation in different Lycium barbarum genotypes with varying root-forming abilities and aims to identify key genes involved in this process. The findings will provide a theoretical basis for in-depth research on the molecular mechanisms underlying adventitious root formation in wolfberry.
RATIONALE: Three wolfberry genotypes with different root-forming abilities were used as experimental materials. A hydroponic experiment was conducted to analyze the transcriptional differences during adventitious root formation.
RESULTS: Transcriptome sequencing identified 6448 differentially expressed genes (DEGs), with the L-vs-H group having the highest number of DEGs at 4413, including 2583 upregulated and 1830 downregulated genes. A total of 281 transcription factors were identified, mainly from the MYB, AP2/ERF, and bHLH families, with distinct expression patterns. GO enrichment analysis revealed that 1714 DEGs were enriched in 32 GO terms. KEGG enrichment analysis indicated that DEGs were mainly enriched in the phenylpropanoid biosynthesis and plant hormone signal transduction pathways. Among these, MYB19 (Lba07g01820) is a core gene in the phenylpropanoid pathway, and TIR1 (Lba08g00069) is a core gene in the plant hormone signal transduction pathway. Both genes play crucial roles in adventitious root formation in wolfberry. qRT-PCR validated the reliability of the transcriptome data.
CONCLUSION: This study elucidates the molecular mechanisms of adventitious root formation in wolfberry and lays a theoretical foundation for the genetic improvement and efficient propagation of wolfberry and other woody plants.
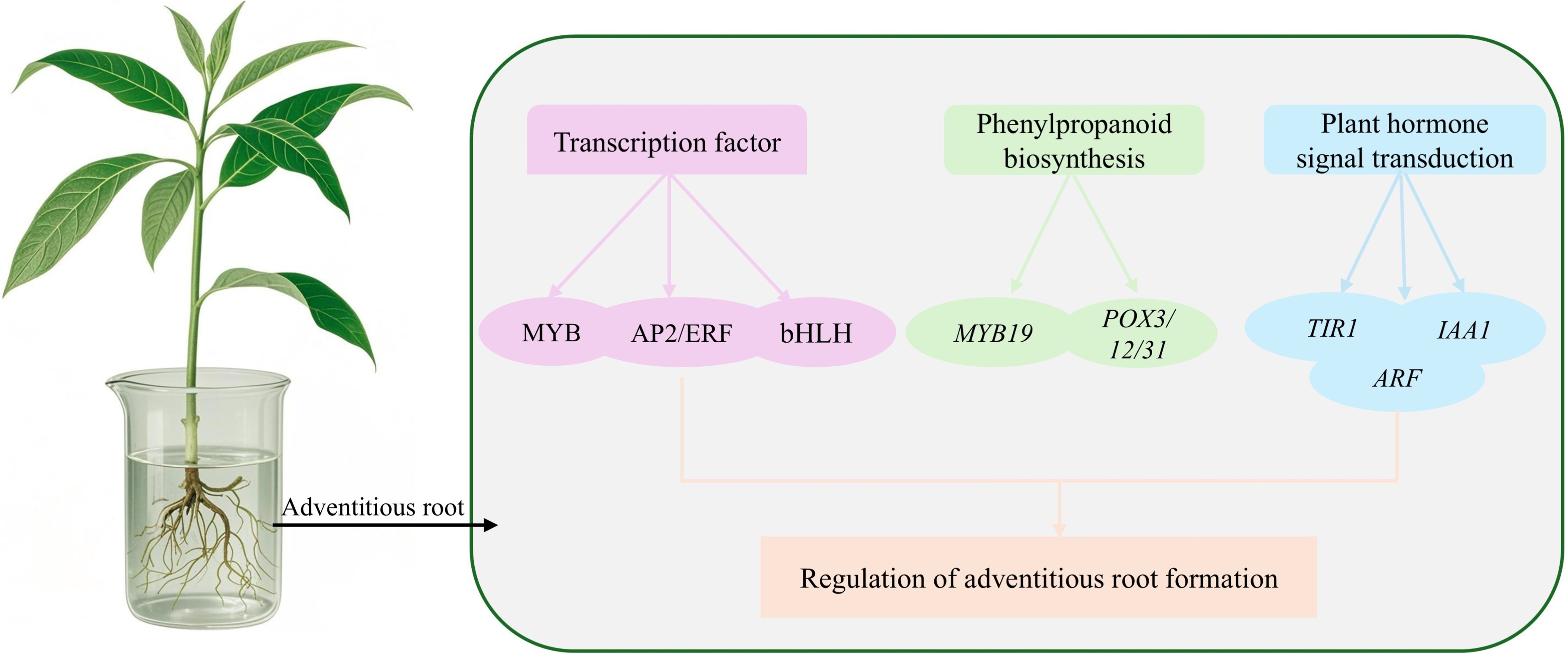
Schematic diagram of transcriptomic analysis of adventitic root generation under hydroponics of Lycium barbarum
INTRODUCTION: Heat stress severely impairs plant growth and crop productivity. WRINKLED1 (WRI1), an AP2/EREBP-class transcription factor in Arabidopsis thaliana, orchestrates carbon partitioning between glycolysis and fatty acid biosynthesis, playing pivotal roles in development and stress adaptation. Elucidating its molecular function under high-temperature stress is critical for improving thermotolerance in crops.
RATIONALE: While WRI1's metabolic regulatory function is established, its role in heat response remains unexplored. To decipher the molecular mechanism of WRI1-mediated thermotolerance, we integrated genetic approaches (wild-type, wri1-4, and WRI1-OE) with RT-qPCR and phenotyping under controlled heat stress.
RESULTS:In this study, histochemical GUS staining of pWRI1::GUS transgenic lines demonstrated constitutive WRI1 expression throughout Arabidopsis seedlings, with significantly enhanced transcription in cotyledons under heat stress (HS) (P<0.05). Prolonged HS induced gradual transcriptional attenuation, though levels remained elevated versus optimal temperature (22°C). RT-qPCR confirmed thermo-responsive WRI1 upregulation (peak:1h HS, 3-fold induction), followed by threshold-dependent decline, indicating acute early-phase responsiveness. Endogenous immunoassays revealed reduced WRI1 protein accumulation under HS, suggesting HS-impaired protein stability or post-translational regulatory mechanisms. Thermotolerance phenotyping of WT, wri1-4, and WRI1-OE lines showed superior HS survival in WRI1-OE, with acquired thermotolerance exceeding basal thermotolerance across genotypes, confirming WRI1-mediated positive thermoregulation. The survival rate of WRI1-OE overexpression seedlings reached approximately 75%-85%, whereas that of wild-type and complementary lines was less than 10%. WRI1-OE reduced reactive oxygen species (ROS) accumulation, while RT-qPCR excluded direct transcriptional regulation of HSF/HSP genes (e.g., HSFA2, HSP101). Differential gene expression across genotypes nevertheless indicated WRI1's auxiliary role in thermotolerance via ROS scavenging and indirect proteostasis maintenance.
CONCLUSION: This study establishes WRI1 as a master regulator of thermotolerance, functioning through synergistic activation of chaperone networks (HSFA2-HSPs) and reactive oxygen species (ROS) scavenging. The discovery of its cross-pathway coordination mechanism provides novel insights into plant thermal adaptation, while positioning WRI1 as an ideal target for breeding climate-resilient crops.
INTRODUCTION: The carbohydrates secreted by plant roots play a crucial role in modulating the carbon source supply and metabolic activity of soil microorganisms.
RATIONALE: In order to unveil the response mechanism of root-secreted carbohydrates in salt-tolerant plants to the intensity of salt stress, this study focused on the salt-tolerant forage grass Thinopyrum ponticum. Four levels of salt stress intensity were conducted by adding NaCl, including control, light, moderate, and severe stress conditions (with soil salt contents of 0%, 0.2%, 0.4%, and 0.6% respectively). Untargeted metabolomics techniques were employed to analyze the compositional alterations of root-secreted carbohydrates and their correlations with the rhizosphere soil properties were evaluated.
RESULTS: The results demonstrated that: (1) Compared to the control, salt stress significantly decreased the contents of five types of carbohydrates, namely glycosides, aminoglycosides, D-glucosamine, 2-O-Methyl-L-fucose and galactoside, while markedly increasing glucose content; (2) Glycosides were the predominant carbohydrates secreted by Thinopyrum ponticum roots, accounting for 79.63% ± 8.19% of the total relative abundance, whereas the remaining five carbohydrate types ranged from 3.40% ± 0.53% to 4.57% ± 1.61%; (3) Root-secreted carbohydrates exhibited significant differences between the control and mild, moderate, or severe salt stress treatments, but no significant distinctions were observed among the latter three treatments; (4) Soil electrical conductivity and nitrate nitrogen content were identified as key factors responsible for the down-regulation of the five carbohydrate types, while soil pH was the primary factor driving the up-regulation of glucose.
CONCLUSION: Thinopyrum ponticum can adapt to salt-stressed environments by regulating the content of carbohydrates secreted by its roots. Meanwhile, the abundance of carbohydrates secreted by the roots is also influenced by the soil properties.
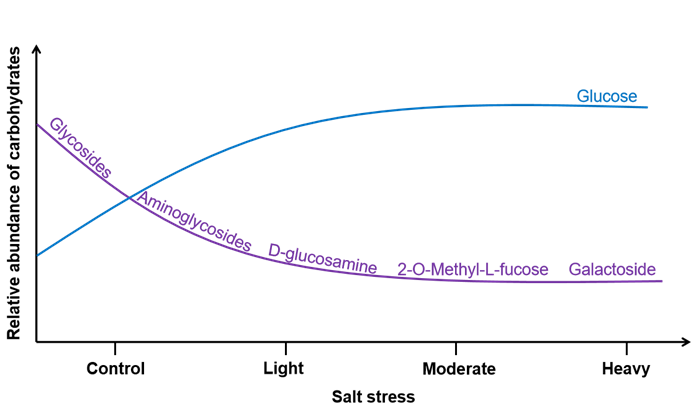

Conceptual map of the influence of salt stress on carbohydrate secretion by the roots of Thinopyrum ponticum
INTRODUCTION: Glycosyltransferases are key enzymes involved in various biological processes in plants. To date, glycosyltransferases have been classified into 138 families, among which the glycosyltransferase family I is the largest. This family primarilyuses UDP-glucose as the glycosyl donor and is referred to as the UGT family. In recent years, the role of UGT genes in plant responses to abiotic stresses has gradually been revealed. However, currently, only a few UGT genes have been clearly identified as being involved in anthocyanin biosynthesis.
RATIONALE: To explore the mechanisms of anthocyanin and its glycosylation-related genes in tea plants under low-temperature stress, this study used FudingDabai Tea as the experimental material. Based on the transcriptome sequencing data, two genes, CsUGT73B4 (CSS0039619) and CsUGT85K5 (CSS0047548), which significantly upregulate anthocyanin expression, were identified. We further investigated the changes in these genes under low-temperature stress and explored the relationship between CsUGT73B4 and CsUGT85K5 genes and anthocyanin metabolism under low-temperature stress. Additionally, the functions of these two genes were verified through heterologous overexpression in Arabidopsis thaliana and gene silencing experiments.
RESULTS: Under low-temperature stress, the expression of CsUGT73B4 and CsUGT85K5 genes was positively correlated with anthocyanin accumulation. In Arabidopsis thaliana plants overexpressing these genes, anthocyanin content and antioxidant enzyme activities (SOD, POD, CAT) were significantly increased, while malondialdehyde (MDA) content and relative conductivity were reduced. Silencing of CsUGT85K5 led to feedback inhibition of CsUGT73B4 expression, revealing that these two genes act in concert to regulate the anthocyanin metabolic pathway.
CONCLUSION: In summary, the CsUGT73B4 and CsUGT85K5 genes are involved in the response to low-temperature stress, regulate anthocyanin metabolism, and may enhance the cold tolerance of tea plants.
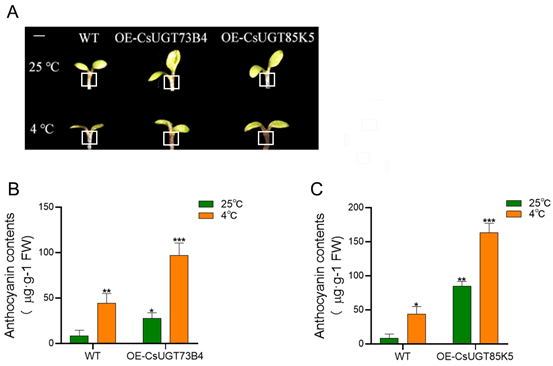
The effect of low-temperature stress on anthocyanin in WT、OE-CsUGT73B4 and OE-CsUGT85K5
(A) Observation of anthocyanin accumulation phenotype in Arabidopsis, the apical meristem position was marked with a white thread (The white line bar=1mm); (B) Effect of low-temperature stress on anthocyanin content in WT and OE-CsUGT73B4 Arabidopsis; (C) Effect of low-temperature stress on anthocyanin content in WT and OE-CsUGT85K5 Arabidopsis. *P<0.05, **P<0.01, ***P<0.001.
INTRODUCTION: Waxes are protective substances found on the surface of plants, especially on fruits and leaves, which help prevent non-stomatal water loss from plant tissues. Lycium barbarum is a characteristic economic forest species of Ningxia. Its fruit is a berry with a high water content, and the surface of the fruit skin is covered with wax, which hinders the normal loss of moisture during the drying process. Alkanes are the main components of the wax on L. barbarum, and the CER3 gene is a key gene involved in alkane biosynthesis. Therefore, an in-depth study of the function of CER3 in L. barbarum provides a theoretical basis for revealing the metabolic synthesis of wax in goji fruits and the molecular mechanism of LbCER3.
RATIONALE: Research on the waxy coating of L. barbarum fruit skin has mainly focused on the structure and composition of the epicuticular wax, with relatively little attention given to its molecular mechanisms. This studyusing the main cultivated varieties of L. barbarum, Ningqi No.1 and Ningqi No.5, as test materials, this study employed methods such as GC-MS, transcriptomics, and molecular biology to investigate the changes in the content and composition of cuticular waxes on the outer and inner layers of the fruit at different developmental stages. It also involved the screening, cloning, and functional validation of key genes related to fruit wax.
RESULTS: During the development and maturation of the fruit, the content of inner and outer cuticular waxes in Ningqi No.1 and Ningqi No.5 gradually decreased, with the total wax content in Ningqi No.1 being significantly higher than that in Ningqi No.5. The outer and inner wax components of both varieties were the same, primarily consisting of alkanes, esters, and acids. Transcriptomic analysis and weighted gene co-expression network analysis (WGCNA) identified LbCER3 as a key gene. The full-length CDS of the LbCER3 gene is 1 884 bp, encoding 628 amino acids, and contains a C-terminal WAX2 domain (Wax2_C) and a fatty acid hydroxylase domain (FA_hydroxylase). An LbCER3 overexpression vector was constructed and successfully transformed into tomato, resulting in a significant reduction in chlorophyll leaching rate in the T1 generation transgenic lines, along with a notable increase in leaf wax content and significant increases in the wax components of alkanes and acids.
CONCLUSION: These findings indicate that LbCER3 is a key gene in the synthesis of wax in L. barbarum.
 Home
Home