INTRODUCTION: Glycosyltransferases are key enzymes involved
in various biological processes in plants. To date, glycosyltransferases have
been classified into 138 families, among which the glycosyltransferase family I
is the largest. This family primarilyuses UDP-glucose as the glycosyl donor and
is referred to as the UGT family. In recent years, the role of UGT genes in
plant responses to abiotic stresses has gradually been revealed. However,
currently, only a few UGT genes have been clearly identified as being involved
in anthocyanin biosynthesis.
RATIONALE: To explore the mechanisms of anthocyanin and its
glycosylation-related genes in tea plants under low-temperature stress, this
study used FudingDabai Tea as the experimental material. Based on the
transcriptome sequencing data, two genes, CsUGT73B4 (CSS0039619) and CsUGT85K5 (CSS0047548), which significantly upregulate anthocyanin expression, were
identified. We further investigated the changes in these genes under
low-temperature stress and explored the relationship between CsUGT73B4 and CsUGT85K5 genes and anthocyanin metabolism under low-temperature
stress. Additionally, the functions of these two genes were verified through
heterologous overexpression in Arabidopsis thaliana and gene silencing
experiments.
RESULTS: Under low-temperature stress, the expression of CsUGT73B4 and CsUGT85K5 genes was positively correlated with anthocyanin
accumulation. In Arabidopsis thaliana plants overexpressing these genes,
anthocyanin content and antioxidant enzyme activities (SOD, POD, CAT) were
significantly increased, while malondialdehyde (MDA) content and relative
conductivity were reduced. Silencing of CsUGT85K5 led to feedback inhibition of
CsUGT73B4 expression, revealing that these two genes act in concert to regulate
the anthocyanin metabolic pathway.
CONCLUSION: In summary, the CsUGT73B4 and CsUGT85K5 genes are involved in the response to low-temperature stress, regulate
anthocyanin metabolism, and may enhance the cold tolerance of tea plants.
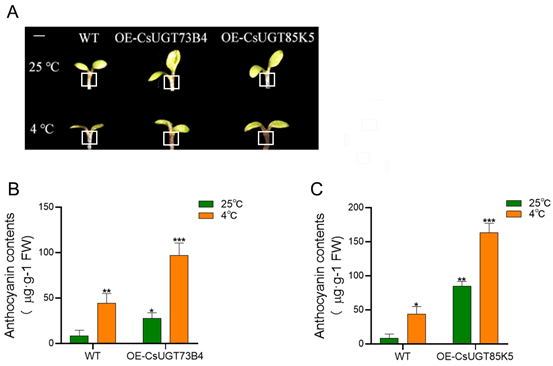
The
effect of low-temperature stress on anthocyanin in WT、OE-CsUGT73B4
and OE-CsUGT85K5
(A) Observation of anthocyanin accumulation phenotype in Arabidopsis,
the apical meristem position was marked with a white thread (The white line bar=1mm); (B) Effect of low-temperature stress on anthocyanin content in WT and
OE-CsUGT73B4 Arabidopsis; (C) Effect of low-temperature stress on
anthocyanin content in WT and OE-CsUGT85K5 Arabidopsis. *P<0.05,
**P<0.01, ***P<0.001.

 Home
Home




