

Chinese Bulletin of Botany ›› 2024, Vol. 59 ›› Issue (3): 495-514.DOI: 10.11983/CBB23090 cstr: 32102.14.CBB23090
• SPECIAL TOPICS • Previous Articles
Fengpan Wang1, Zhaoxuan Zhong1,2, Lijun Chen1, Jiangping Shu1, Yuehong Yan1,*( )
)
Received:2023-07-08
Accepted:2023-12-19
Online:2024-05-10
Published:2024-05-10
Contact:
E-mail: Fengpan Wang, Zhaoxuan Zhong, Lijun Chen, Jiangping Shu, Yuehong Yan. A Comprehensive Overview of the Studies on the Gene Function in Pteridophytes[J]. Chinese Bulletin of Botany, 2024, 59(3): 495-514.
| 编号 | 物种 | 基因组 大小(M) | 染色体 数目(n) | 全基因 组重复 事件(次) | 重复序列占比(%) | 基因 数目 | 生境 | 是否叶两型 | 孢子类型 | 参考文献 |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 江南卷柏(Selaginella moellendorffii) | 212.6 | 10 | 0 | - | 22285 | 陆生 | 否 | 异型 | Banks et al., |
| 2 | 翠云草(S. kraussiana) | 115 | - | - | - | - | 陆生 | 否 | 异型 | Ge et al., |
| 3 | 卷柏(S. tamariscina) | 301 | - | - | 60.22 | 27761 | 陆生 | 否 | 异型 | Xu et al., |
| 4 | 膜叶卷柏(S. lepidophylla) | 109 | - | - | 27204 | 陆生 | 否 | 异型 | VanBuren et al., | |
| 5 | 细叶满江红(Azolla filiculoides) | 750 | 22 | 2 | 53.6 | 20201 | 水生 | 否 | 异型 | Li et al., |
| 6 | 勺叶槐叶蘋(Salvinia cucullata) | 260 | 9 | 1 | 44.5 | 19914 | 水生 | 否 | 异型 | Li et al., |
| 7 | 台湾水韭(I. taiwanensis) | 1658.3 | 11 | 1 | 38 | 39461 | 水生 | 是 | 异型 | Wickell et al., |
| 8 | 中华水韭(I. sinensis) | 2131.8 | 22 | 2 | 63.15 | 57303 | 水生 | 否 | 异型 | Cui et al., |
| 9 | 荷叶铁线蕨(Adiantum nelumboides) | 1977.1 | 60 | 1 | 81.7 | 69568 | 陆生 | 否 | 同型 | Zhong et al., |
| 10 | 美洲水蕨(Ceratopteris richardii) | 7463.3 | 39 | 2 | 85.2 | 36875 | 水生 | 是 | 同型 | Marchant et al., 2022 |
| 11 | 桫椤(Alsophila spinulosa) | 6230 | 69 | 2 | 75.15 | 67831 | 陆生 | 否 | 同型 | Huang et al., |
| 12 | 铁线蕨(Adiantum capillus-veneris) | 4830 | 30 | 0 | 85.25 | 31244 | 陆生 | 否 | 同型 | Fang et al., |
| 13 | 东北石松(Lycopodium clavatum) | 2304.7 | - | 2 | 85.32 | 23292 | 陆生 | 否 | 同型 | Yu et al., |
Table 1 Pteridophytes with whole-genome sequence published
| 编号 | 物种 | 基因组 大小(M) | 染色体 数目(n) | 全基因 组重复 事件(次) | 重复序列占比(%) | 基因 数目 | 生境 | 是否叶两型 | 孢子类型 | 参考文献 |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 江南卷柏(Selaginella moellendorffii) | 212.6 | 10 | 0 | - | 22285 | 陆生 | 否 | 异型 | Banks et al., |
| 2 | 翠云草(S. kraussiana) | 115 | - | - | - | - | 陆生 | 否 | 异型 | Ge et al., |
| 3 | 卷柏(S. tamariscina) | 301 | - | - | 60.22 | 27761 | 陆生 | 否 | 异型 | Xu et al., |
| 4 | 膜叶卷柏(S. lepidophylla) | 109 | - | - | 27204 | 陆生 | 否 | 异型 | VanBuren et al., | |
| 5 | 细叶满江红(Azolla filiculoides) | 750 | 22 | 2 | 53.6 | 20201 | 水生 | 否 | 异型 | Li et al., |
| 6 | 勺叶槐叶蘋(Salvinia cucullata) | 260 | 9 | 1 | 44.5 | 19914 | 水生 | 否 | 异型 | Li et al., |
| 7 | 台湾水韭(I. taiwanensis) | 1658.3 | 11 | 1 | 38 | 39461 | 水生 | 是 | 异型 | Wickell et al., |
| 8 | 中华水韭(I. sinensis) | 2131.8 | 22 | 2 | 63.15 | 57303 | 水生 | 否 | 异型 | Cui et al., |
| 9 | 荷叶铁线蕨(Adiantum nelumboides) | 1977.1 | 60 | 1 | 81.7 | 69568 | 陆生 | 否 | 同型 | Zhong et al., |
| 10 | 美洲水蕨(Ceratopteris richardii) | 7463.3 | 39 | 2 | 85.2 | 36875 | 水生 | 是 | 同型 | Marchant et al., 2022 |
| 11 | 桫椤(Alsophila spinulosa) | 6230 | 69 | 2 | 75.15 | 67831 | 陆生 | 否 | 同型 | Huang et al., |
| 12 | 铁线蕨(Adiantum capillus-veneris) | 4830 | 30 | 0 | 85.25 | 31244 | 陆生 | 否 | 同型 | Fang et al., |
| 13 | 东北石松(Lycopodium clavatum) | 2304.7 | - | 2 | 85.32 | 23292 | 陆生 | 否 | 同型 | Yu et al., |
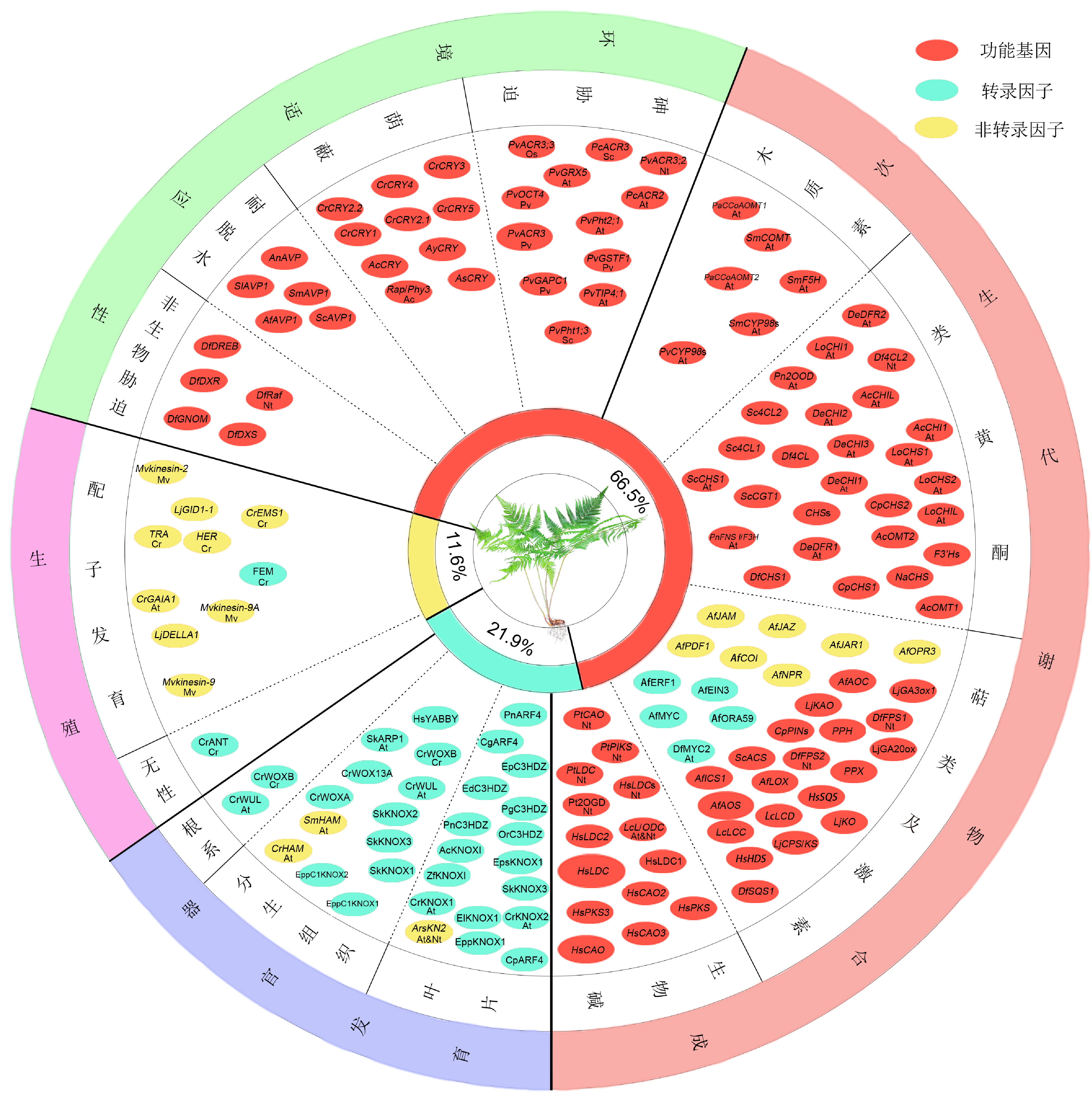
Figure 3 Overview of gene function researches in pteridophytes Ac: Adiantum capillus-veneris; At: Arabidopsis thaliana; Cr: Ceratopteris richardii; Mv: Marsilea vestita; Nt: Nicotiana tabacum; Os: Oryza sativa; Pv: Pteris vittata; Sc: Saccharomyces cerevisiae. Percentage means the ratio of different gene categories (transcription factor, non-transcription factor and functional gene) in all investigated genes.
| [1] | 陈雪菲, 张映雪, 黄贤燕, 王全喜, 曹建国, 王幼芳 (2020). 蕨类植物花青素和原花青素成分及含量分析. 植物科学学报 38, 820-830. |
| [2] |
顾钰峰, 金冬梅, 刘保东, 戴锡玲, 严岳鸿 (2020). 蕨类植物的鳞片特征及演化I: 凤尾蕨科. 植物学报 55, 163-176.
DOI |
| [3] | 黄小芳 (2016). 基于转录组测序的红盖鳞毛蕨类黄酮3'-羟化酶(F3'H)基因的克隆与表达分析. 硕士论文. 上海: 上海师范大学. pp. 1-64. |
| [4] | 李赛男, 王雯静, 张蓓蓓, 张泽坤, 葛祥宇, 杜宇, 张晓雪, 王娟, 史社坡 (2022). 蛇足石杉中赖氨酸脱羧酶基因的克隆、表达及功能鉴定. 药学学报 57, 3437-3445. |
| [5] | 刘爱佳, 陈晓英, 李翠, 闫志刚, 冯世鑫, 缪剑华, 雷明 (2022). 蛇足石杉HsCAO3基因的克隆及转录自激活能力检测. 分子植物育种 20, 809-816. |
| [6] | 刘爱佳, 陈晓英, 李翠, 闫志刚, 冯世鑫, 张占江, 雷明, 缪剑华 (2021). 蛇足石杉HsCAO2基因的克隆、生物信息学分析及载体构建. 中草药 52, 7309-7316. |
| [7] |
严岳鸿, 卫然, 舒江平, 张宪春 (2019). 通过现存蕨类植物多样性透视陆生植物的演化. 生物多样性 27, 1165-1171.
DOI |
| [8] | Alber AV, Renault H, Basilio-Lopes A, Bassard JE, Liu ZH, Ullmann P, Lesot A, Bihel F, Schmitt M, Werck- Reichhart D (2019). Evolution of coumaroyl conjugate 3-hydroxylases in land plants: lignin biosynthesis and defense. Plant J 99, 924-936. |
| [9] | Alejo-Jacuinde G, Herrera-Estrella L (2022). Exploring the high variability of vegetative desiccation tolerance in pteridophytes. Plants 11, 1222. |
| [10] | Anand A, Bass SH, Wu E, Wang N, McBride KE, Annaluru N, Miller M, Hua M, Jones TJ (2018). An improved ternary vector system for Agrobacterium-mediated rapid maize transformation. Plant Mol Biol 97, 187-200. |
| [11] | Araki T, Saga Y, Marugami M, Otaka J, Araya H, Saito K, Yamazaki M, Suzuki H, Kushiro T (2016). Onocerin biosynthesis requires two highly dedicated triterpene cyclases in a fern Lycopodium clavatum. ChemBioChem 17, 288-290. |
| [12] |
Atallah NM, Banks JA (2015). Reproduction and the pheromonal regulation of sex type in fern gametophytes. Front Plant Sci 6, 100.
DOI PMID |
| [13] | Banks JA (1997). Sex determination in the fern Ceratopteris. Trends Plant Sci 2, 175-180. |
| [14] |
Banks JA, Nishiyama T, Hasebe M, Bowman JL, Gribskov M, dePamphilis C, Albert VA, Aono N, Aoyama T, Ambrose BA, Ashton NW, Axtell MJ, Barker E, Barker MS, Bennetzen JL, Bonawitz ND, Chapple C, Cheng CY, Correa LGG, Dacre M, DeBarry J, Dreyer I, Elias M, Engstrom EM, Estelle M, Feng L, Finet C, Floyd SK, Frommer WB, Fujita T, Gramzow L, Gutensohn M, Harholt J, Hattori M, Heyl A, Hirai T, Hiwatashi Y, Ishikawa M, Iwata M, Karol KG, Koehler B, Kolukisaoglu U, Kubo M, Kurata T, Lalonde S, Li KJ, Li Y, Litt A, Lyons E, Manning G, Maruyama T, Michael TP, Mikami K, Miyazaki S, Morinaga SI, Murata T, Mueller-Roeber B, Nelson DR, Obara M, Oguri Y, Olmstead RG, Onodera N, Petersen BL, Pils B, Prigge M, Rensing SA, Riaño-Pachón DM, Roberts AW, Sato Y, Scheller HV, Schulz B, Schulz C, Shakirov EV, Shibagaki N, Shinohara N, Shippen DE, Sørensen I, Sotooka R, Sugimoto N, Sugita M, Sumikawa N, Tanurdzic M, Theißen G, Ulvskov P, Wakazuki S, Weng JK, Willats WWGT, Wipf D, Wolf PG, Yang LX, Zimmer AD, Zhu QH, Mitros T, Hellsten U, Loqué D, Otillar R, Salamov A, Schmutz J, Shapiro H, Lindquist E, Lucas S, Rokhsar D, Grigoriev IV (2011). The Selaginella genome identifies genetic changes associated with the evolution of vascular plants. Science 332, 960-963.
DOI PMID |
| [15] | Berruezo F, de Souza FSJ, Picca PI, Nemirovsky SI, Martínez Tosar L, Rivero M, Mentaberry AN, Zelada AM (2017). Sequencing of small RNAs of the fern Pleopeltis minima (Polypodiaceae) offers insight into the evolution of the microrna repertoire in land plants. PLoS One 12, e0177573. |
| [16] |
Bharathan G, Goliber TE, Moore C, Kessler S, Pham T, Sinha NR (2002). Homologies in leaf form inferred from KNOXI gene expression during development. Science 296, 1858-1860.
DOI PMID |
| [17] | Bui LT, Pandzic D, Youngstrom CE, Wallace S, Irish EE, Szövényi P, Cheng CL (2017). A fern AINTEGUMENTA gene mirrors BABY BOOM in promoting apogamy in Ceratopteris richardii. Plant J 90, 122-132. |
| [18] |
Bunsupa S, Hanada K, Maruyama A, Aoyagi K, Komatsu K, Ueno H, Yamashita M, Sasaki R, Oikawa A, Saito K, Yamazaki M (2016). Molecular evolution and functional characterization of a bifunctional decarboxylase involved in lycopodium alkaloid biosynthesis. Plant Physiol 171, 2432-2444.
DOI PMID |
| [19] | Cai C, Lanman NA, Withers KA, DeLeon AM, Wu Q, Gribskov M, Salt DE, Banks JA (2019). Three genes define a bacterial-like arsenic tolerance mechanism in the arsenic hyperaccumulating fern Pteris vittata. Curr Biol 29, 1625-1633. |
| [20] | Cai SG, Chen G, Wang YY, Huang YQ, Marchant DB, Wang YZ, Yang Q, Dai F, Hills A, Franks PJ, Nevo E, Soltis DE, Soltis PS, Sessa E, Wolf PG, Xue DW, Zhang GP, Pogson BJ, Blatt MR, Chen ZH (2017). Evolutionary conservation of ABA signaling for stomatal closure. Plant Physiol 174, 732-747. |
| [21] | Cai SG, Huang YQ, Chen F, Zhang X, Sessa E, Zhao CC, Marchant DB, Xue DW, Chen G, Dai F, Leebens-Mack JH, Zhang GP, Shabala S, Christie JM, Blatt MR, Nevo E, Soltis PS, Soltis DE, Franks PJ, Wu FB, Chen ZH (2021). Evolution of rapid blue-light response linked to explosive diversification of ferns in angiosperm forests. New Phytol 230, 1201-1213. |
| [22] | Cai ZP, Xie ZY, Wang XC, Zhang SX, Wu Q, Yu XD, Guo Y, Gao SY, Zhang YG, Xu ST, Wang HG, Luo JJ (2022). Excavation of genes responsive to brassinosteroids by transcriptome sequencing in Adiantum flabellulatum gametophytes. Genes 13, 1061. |
| [23] |
Cardoso AA, Randall JM, McAdam SA (2019). Hydraulics regulate stomatal responses to changes in leaf water status in the fern Athyrium filix-femina. Plant Physiol 179, 533-543.
DOI PMID |
| [24] | Chen JJ, Tomes S, Gleave AP, Hall W, Luo ZW, Xu J, Yao JL (2022). Significant improvement of apple (Malus domestica Borkh.) transgenic plant production by pre-transformation with a Baby boom transcription factor. Hortic Res 9, uhab014. |
| [25] | Chen JX, Cao Y, Yan XJ, Chen YS, Ma LQ (2021). Novel PvACR3;2 and PvACR3;3 genes from arsenic-hyperaccumulator Pteris vittata and their roles in manipulating plant arsenic accumulation. J Hazard Mater 415, 125647. |
| [26] | Chen YS, Xu WZ, Shen HL, Yan HL, Xu WX, He ZY, Ma M (2013). Engineering arsenic tolerance and hyperaccumulation in plants for phytoremediation by a PvACR3 transgenic approach. Environ Sci Technol 47, 9355-9362. |
| [27] | Cordle AR, Irish EE, Cheng CL (2007). Apogamy induction in Ceratopteris richardii. Int J Plant Sci 168, 361-369. |
| [28] | Costarelli A, Cannavò S, Cerri M, Pellegrino RM, Reale L, Paolocci F, Pasqualini S (2021). Light and temperature shape the phenylpropanoid profile of Azolla filiculoides fronds. Front Plant Sci 12, 727667. |
| [29] | Cruz R, Melo-de-Pinna GFA, Vasco A, Prado J, Ambrose BA (2020). Class I KNOX is related to determinacy during the leaf development of the fern Mickelia scandens (Dryopteridaceae). Int J Mol Sci 21, 4295. |
| [30] | Cui JT, Zhu YK, Du H, Liu ZH, Shen SQ, Wang TX, Cui WW, Zhang R, Jiang SJ, Wu YM, Gu XF, Yu H, Liang Z (2023). Chromosome-level reference genome of tetraploid Isoetes sinensis provides insights into evolution and adaption of lycophytes. GigaScience 12, giad079. |
| [31] |
de Vries J, Fischer AM, Roettger M, Rommel S, Schluepmann H, Bräutigam A, Carlsbecker A, Gould SB (2016). Cytokinin-induced promotion of root meristem size in the fern Azolla supports a shoot-like origin of euphyllophyte roots. New Phytol 209, 705-720.
DOI PMID |
| [32] | de Vries S, de Vries J, Teschke H, von Dahlen JK, Rose LE, Gould SB (2018). Jasmonic and salicylic acid response in the fern Azolla filiculoides and its cyanobiont. Plant Cell Environ 41, 2530-2548. |
| [33] |
Debernardi JM, Tricoli DM, Ercoli MF, Hayta S, Ronald P, Palatnik JF, Dubcovsky J (2020). A GRF-GIF chimeric protein improves the regeneration efficiency of transgenic plants. Nat Biotechnol 38, 1274-1279.
DOI PMID |
| [34] | DiTusa SF, Fontenot EB, Wallace RW, Silvers MA, Steele TN, Elnagar AH, Dearman KM, Smith AP (2016). A member of the phosphate transporter 1 (Pht1) family from the arsenic-hyperaccumulating fern Pteris vittata is a high-affinity arsenate transporter. New Phytol 209, 762-772. |
| [35] |
Fang YH, Qin X, Liao QG, Du R, Luo XZ, Zhou Q, Li Z, Chen HC, Jin WT, Yuan YN, Sun PB, Zhang R, Zhang J, Wang L, Cheng SF, Yang XY, Yan YH, Zhang XT, Zhang ZH, Bai SN, Van de Peer Y, Lucas WJ, Huang SW, Yan JB (2022). The genome of homosporous maidenhair fern sheds light on the euphyllophyte evolution and defences. Nat Plants 8, 1024-1037.
DOI PMID |
| [36] | Feng HY, Li XY, Sun D, Chen YS, Xu GH, Cao Y, Ma LQ (2021). Expressing phosphate transporter PvPht2;1 enhances P transport to the chloroplasts and increases arsenic tolerance in Arabidopsis thaliana. Environ Sci Technol 55, 2276-2284. |
| [37] |
Frangedakis E, Saint-Marcoux D, Moody LA, Rabbinowitsch E, Langdale JA (2017). Nonreciprocal complementation of KNOX gene function in land plants. New Phytol 216, 591-604.
DOI PMID |
| [38] | Gao R, Yu D, Chen LL, Wang WZ, Sun LL, Chang Y (2019). Cloning and functional analysis of squalene synthase gene from Dryopteris fragrans (L.) Schott. Protein Expres Purif 155, 95-103. |
| [39] | Ge YC, Liu J, Zeng MH, He JF, Qin P, Huang H, Xu L (2016). Identification of WOX family genes in Selaginella kraussiana for studies on stem cells and regeneration in lycophytes. Front Plant Sci 7, 93. |
| [40] | Geng Y, Guo L, Han H, Liu X, Banks JA, Wisecaver JH, Zhou Y (2021). Conservation and diversification of HAIRY MERISTEM gene family in land plants. Plant J 106, 366-378. |
| [41] | Geng Y, Yan A, Zhou Y (2022). Positional cues and cell division dynamics drive meristem development and archegonium formation in Ceratopteris gametophytes. Commun Biol 5, 650. |
| [42] | Güngör E, Brouwer P, Dijkhuizen LW, Shaffar DC, Nierop KGJ, de Vos RCH, Sastre Toraño J, van Der Meer IM, Schluepmann H (2021). Azolla ferns testify: seed plants and ferns share a common ancestor for leucoanthocyanidin reductase enzymes. New Phytol 229, 1118-1132. |
| [43] | Harrison CJ, Corley SB, Moylan EC, Alexander DL, Scotland RW, Langdale JA (2005). Independent recruitment of a conserved developmental mechanism during leaf evolution. Nature 434, 509-514. |
| [44] | He ZY, Yan HL, Chen YS, Shen HL, Xu WX, Zhang HY, Shi L, Zhu YG, Ma M (2016). An aquaporin PvTIP4;1 from Pteris vittata may mediate arsenite uptake. New Phytol 209, 746-761. |
| [45] | Hendrix SD (1980). An evolutionary and ecological perspective of the insect fauna of ferns. Am Nat 115, 171-196. |
| [46] |
Hidalgo O, Pellicer J, Christenhusz M, Schneider H, Leitch AR, Leitch IJ (2017). Is there an upper limit to genome size? Trends Plant Sci 22, 567-573.
DOI PMID |
| [47] | Hong YF, Wang Z, Li MH, Su YJ, Wang T (2022). First multi-organ full-length transcriptome of tree fern Alsophila spinulosa highlights the stress-resistant and light-adapted genes. Front Genet 12, 784546. |
| [48] |
Huang X, Wang WL, Gong T, Wickell D, Kuo LY, Zhang XT, Wen JL, Kim H, Lu FC, Zhao HS, Chen S, Li H, Wu WQ, Yu CJ, Chen S, Fan W, Chen S, Bao XQ, Li L, Zhang D, Jiang LY, Khadka D, Yan XJ, Liao ZY, Zhou GK, Guo YL, Ralph J, Sederoff RR, Wei HR, Zhu P, Li FW, Ming R, Li QZ (2022). The flying spider-monkey tree fern genome provides insights into fern evolution and arborescence. Nat Plants 8, 500-512.
DOI PMID |
| [49] |
Imaizumi T, Kanegae T, Wada M (2000). Cryptochrome nucleocytoplasmic distribution and gene expression are regulated by light quality in the fern Adiantum capillus- veneris. Plant Cell 12, 81-95.
DOI PMID |
| [50] | Indriolo E, Na G, Ellis D, Salt DE, Banks JA (2010). A vacuolar arsenite transporter necessary for arsenic tolerance in the arsenic hyperaccumulating fern Pteris vittata is missing in flowering plants. Plant Cell 22, 2045-2057. |
| [51] | Jiménez A, Quintanilla LG, Pajarón S, Pangua E (2008). Reproductive and competitive interactions among gametophytes of the allotetraploid fern Dryopteris corleyi and its two diploid parents. Ann Bot 102, 353-359. |
| [52] | Kawai H, Kanegae T, Christensen S, Kiyosue T, Sato Y, Imaizumi T, Kadota A, Wada M (2003). Responses of ferns to red light are mediated by an unconventional photoreceptor. Nature 421, 287-290. |
| [53] |
Kawai-Toyooka H, Kuramoto C, Orui K, Motoyama K, Kikuchi K, Kanegae T, Wada M (2004). DNA interference: a simple and efficient gene-silencing system for high-throughput functional analysis in the fern Adiantum. Plant Cell Physiol 45, 1648-1657.
PMID |
| [54] | Kinosian SP, Wolf PG (2022). The biology of C. richardii as a tool to understand plant evolution. eLife 11, e75019. |
| [55] | Kumar RP, Siddique S (2022). 22-Hydroxyhopane, a novel multitargeted phytocompound against SARS-CoV-2 from Adiantum latifolium Lam. Nat Prod Res 36, 4276-4281. |
| [56] | Lane TS, Rempe CS, Davitt J, Staton ME, Peng YH, Soltis DE, Melkonian M, Deyholos M, Leebens-Mack JH, Chase M, Rothfels CJ, Stevenson D, Graham SW, Yu J, Liu T, Pires JC, Edger PP, Zhang Y, Xie YL, Zhu Y, Carpenter E, Wong GKS, Stewart CN Jr (2016). Diversity of ABC transporter genes across the plant kingdom and their potential utility in biotechnology. BMC Biotechnol 16, 47. |
| [57] | Li C, Huang WJ, Han XX, Zhao GH, Zhang WY, He WJ, Nie B, Chen XF, Zhang TJ, Bai WH, Zhang XP, He JJ, Zhao C, Fernie AR, Tschaplinski TJ, Yang XH, Yan SJ, Wang L (2023). Diel dynamics of multi-omics in elkhorn fern provide new insights into weak CAM photosynthesis. Plant Commun 4, 100594. |
| [58] | Li FW, Brouwer P, Carretero-Paulet L, Cheng SF, De Vries J, Delaux PM, Eily A, Koppers N, Kuo LY, Li Z, Simenc M, Small I, Wafula E, Angarita S, Barker MS, Bräutigam A, DePamphilis C, Gould S, Hosmani PS, Huang YM, Huettel B, Kato Y, Liu X, Maere S, McDowell R, Mueller LA, Nierop KGJ, Rensing SA, Robison T, Rothfels CJ, Sigel EM, Song Y, Timilsena PR, Van de Peer Y, Wang HL, Wilhelmsson PKI, Wolf PG, Xu X, Der JP, Schluepmann H, Wong GKS, Pryer KM (2018a). Fern genomes elucidate land plant evolution and cyanobacterial symbioses. Nat Plants 4, 460-472. |
| [59] | Li QC, Wang HB, Wang HJ, Li Y, Wang ZZ, Zhang XM (2018b). Effect of arsenate on endogenous levels of cytokinins with different existing forms in two Pteris species. Plant Physiol Biochem 132, 652-659. |
| [60] | Li SS, Li Y, Sun LL, Hu BZ, Chang Y (2015). Identification and expression analysis of 4-coumarate: coenzyme a ligase gene family in Dryopteris fragrans. Cell Mol Biol 61, 25-33. |
| [61] | Li XY, Sun D, Feng HY, Chen JX, Chen YS, Li HB, Cao Y, Ma LQ (2020). Efficient arsenate reduction in As-hyperaccumulator Pteris vittata are mediated by novel arsenate reductases PvHAC1 and PvHAC2. J Hazard Mater 399, 122895. |
| [62] |
Liu W, Xu L (2018). Recruitment of IC-WOX genes in root evolution. Trends Plant Sci 23, 490-496.
DOI PMID |
| [63] | Ma LQ, Komar KM, Tu C, Zhang WH, Cai Y, Kennelley ED (2001). A fern that hyperaccumulates arsenic. Nature 409, 579. |
| [64] |
Marchant DB (2019). Ferns with benefits: incorporating Ceratopteris into the genomics era. Am Fern J 109, 183-191.
DOI |
| [65] | Marchant DB, Chen G, Cai SG, Chen F, Schafran P, Jenkins J, Shu SQ, Plott C, Webber J, Lovell JT, He GF, Sandor L, Williams M, Rajasekar S, Healey A, Barry K, Zhang YW, Sessa E, Dhakal RR, Wolf PG, Harkess A, Li FW, Rössner C, Becker A, Gramzow L, Xue DW, Wu YH, Tong T, Wang YY, Dai F, Hua SJ, Wang H, Xu SC, Xu F, Duan HL, Theißen G, McKain MR, Li Z, Mckibben MTW, Barker MS, Schmitz RJ, Stevenson DW, Zumajo-Cardona C, Ambrose BA, Leebens-Mack JH, Grimwood J, Schmutz J, Soltis PS, Soltis DE, Chen ZH (2022). Dynamic genome evolution in a model fern. Nat Plants 8, 1038-1051. |
| [66] |
Marks RA, Hotaling S, Frandsen PB, VanBuren R (2021). Representation and participation across 20 years of plant genome sequencing. Nat Plants 7, 1571-1578.
DOI PMID |
| [67] | Mathews S, Ma LQ, Rathinasabapathi B, Natarajan S, Saha UK (2010). Arsenic transformation in the growth media and biomass of hyperaccumulator Pteris vittata L. Bioresour Technol 101, 8024-8030. |
| [68] | McAdam SAM, Brodribb TJ (2012). Fern and lycophyte guard cells do not respond to endogenous abscisic acid. Plant Cell 24, 1510-1521. |
| [69] |
McAdam SAM, Brodribb TJ (2013). Ancestral stomatal control results in a canalization of fern and lycophyte adaptation to drought. New Phytol 198, 429-441.
DOI PMID |
| [70] |
McAdam SAM, Brodribb TJ, Banks JA, Hedrich R, Atallah NM, Cai C, Geringer MA, Lind C, Nichols DS, Stachowski K, Geiger D, Sussmilch FC (2016). Abscisic acid controlled sex before transpiration in vascular plants. Proc Natl Acad Sci USA 113, 12862-12867.
DOI PMID |
| [71] |
McFarland FL, Collier R, Walter N, Martinell B, Kaeppler SM, Kaeppler HF (2023). A key to totipotency: Wuschel- like homeobox 2a unlocks embryogenic culture response in maize (Zea mays L.). Plant Biotechnol J 21, 1860-1872.
DOI PMID |
| [72] | Muthukumar B, Joyce BL, Elless MP, Stewart CN Jr (2013). Stable transformation of ferns using spores as targets: Pteris vittata and Ceratopteris thalictroides. Plant Physiol 163, 648-658. |
| [73] | Nett RS, Dho Y, Low YY, Sattely ES (2021). A metabolic regulon reveals early and late acting enzymes in neuroactive lycopodium alkaloid biosynthesis. Proc Natl Acad Sci USA 118, e2102949118. |
| [74] | Niu M, Fu J, Ni R, Xiong RL, Zhu TT, Lou HX, Zhang P, Li JX, Cheng AX (2021). Functional and structural investigation of chalcone synthases based on integrated metabolomics and transcriptome analysis on flavonoids and anthocyanins biosynthesis of the fern Cyclosorus parasiticus. Front Plant Sci 12, 757516. |
| [75] | Niu XM, Xu YC, Li ZW, Bian YT, Hou XH, Chen JF, Zou YP, Jiang J, Wu Q, Ge S, Balasubramanian S, Guo YL (2019). Transposable elements drive rapid phenotypic variation in Capsella rubella. Proc Natl Acad Sci USA 116, 6908-6913. |
| [76] |
Ostria-Gallardo E, Larama G, Berríos G, Fallard A, Gutiérrez-Moraga A, Ensminger I, Bravo LA (2020). A comparative gene co-expression analysis using self-organizing maps on two congener filmy ferns identifies specific desiccation tolerance mechanisms associated to their microhabitat preference. BMC Plant Biol 20, 56.
DOI PMID |
| [77] | Plackett ARG, Emms DM, Kelly S, Hetherington AM, Langdale JA (2021). Conditional stomatal closure in a fern shares molecular features with flowering plant active stomatal responses. Curr Biol 31, 4560-4570. |
| [78] |
Plackett ARG, Huang LD, Sanders HL, Langdale JA (2014). High-efficiency stable transformation of the model fern species Ceratopteris richardii via microparticle bombardment. Plant Physiol 165, 3-14.
DOI PMID |
| [79] |
Plackett ARG, Rabbinowitsch EH, Langdale JA (2015). Protocol: genetic transformation of the fern Ceratopteris richardii through microparticle bombardment. Plant Methods 11, 37.
DOI PMID |
| [80] | Popov M, Zemanová V, Sácký J, Pavlík M, Leonhardt T, Matoušek T, Kaňa A, Pavlíková D, Kotrba P (2021). Arsenic accumulation and speciation in two cultivars of Pteris cretica L. and characterization of arsenate reductase PcACR2 and arsenite transporter PcACR3 genes in the hyperaccumulating cv. Albo-lineata. Ecotoxicol Environ Saf 216, 112196. |
| [81] |
Pradeep Kumar R, Dinesh Babu KV, Evans DA (2019). Isolation, characterization and mode of action of a larvicidal compound, 22-hydroxyhopane from Adiantum latifolium Lam. against Oryctes rhinoceros Linn. Pestic Biochem Physiol 153, 161-170.
DOI PMID |
| [82] |
Prats KA, Brodersen CR (2021). Desiccation and rehydration dynamics in the epiphytic resurrection fern Pleopeltis polypodioides. Plant Physiol 187, 1501-1518.
DOI PMID |
| [83] | Putzier CC, Polich SB, Verhoeven AS (2022). Sustained zeaxanthin-dependent thermal dissipation is induced by desiccation and low temperatures in the fern Polypodium virginianum. Physiol Plant 174, e13743. |
| [84] |
Rodríguez-Pelayo C, Ambrose BA, Vasco A, Alzate JF, Pabón-Mora N (2022). Evolution and expression of LEAFY genes in ferns and lycophytes. EvoDevo 13, 2.
DOI PMID |
| [85] | Ruiz-Ruano FJ, Navarro-Domínguez B, Camacho JPM, Garrido-Ramos MA (2022). Transposable element landscapes illuminate past evolutionary events in the endangered fern Vandenboschia speciosa. Genome 65, 95-103. |
| [86] |
Rutherford G, Tanurdzic M, Hasebe M, Banks JA (2004). A systemic gene silencing method suitable for high throughput, reverse genetic analyses of gene function in fern gametophytes. BMC Plant Biol 4, 6.
PMID |
| [87] |
Sánchez-Bermejo E, Castrillo G, Del Llano B, Navarro C, Zarco-Fernández S, Martinez-Herrera DJ, Leo-del Puerto Y, Muñoz R, Cámara C, Paz-Ares J, Alonso- Blanco C, Leyva A (2014). Natural variation in arsenate tolerance identifies an arsenate reductase in Arabidopsis thaliana. Nat Commun 5, 4617.
DOI PMID |
| [88] |
Sano R, Juárez CM, Hass B, Sakakibara K, Ito M, Banks JA, Hasebe M (2005). KNOX homeobox genes potentially have similar function in both diploid unicellular and multicellular meristems, but not in haploid meristems. Evol Dev 7, 69-78.
PMID |
| [89] |
Shukla AK, Upadhyay SK, Mishra M, Saurabh S, Singh R, Singh H, Thakur N, Rai P, Pandey P, Hans AL, Srivastava S, Rajapure V, Yadav SK, Singh MK, Kumar J, Chandrashekar K, Verma PC, Singh AP, Nair KN, Bhadauria S, Wahajuddin M, Singh S, Sharma S, Omkar, Upadhyay RS, Ranade SA, Tuli R, Singh PK (2016). Expression of an insecticidal fern protein in cotton protects against whitefly. Nat Biotechnol 34, 1046-1051.
DOI PMID |
| [90] |
Siengalewicz P, Mulzer J, Rinner U (2013). Lycopodium alkaloids-synthetic highlights and recent developments. Alkaloids Chem Biol 72, 1-151.
PMID |
| [91] |
Siriwardana NS, Lamb RS (2012). The poetry of reproduction: the role of LEAFY in Arabidopsis thaliana flower formation. Int J Dev Biol 56, 207-221.
DOI PMID |
| [92] | Song CH, Guan YL, Zhang DR, Tang X, Chang Y (2022). Integrated mRNA and miRNA transcriptome analysis suggests a regulatory network for UV-B-controlled terpenoid synthesis in fragrant woodfern (Dryopteris fragrans). Int J Mol Sci 23, 5708. |
| [93] | Sripinyowanich S, Kil EJ, Petchsri S, Jo Y, Choi H, Cho WK, Lee S (2021). De novo transcriptome assembly of two Microsorum fern species identifies enzymes required for two upstream pathways of phytoecdysteroids. Int J Mol Sci 22, 2085. |
| [94] | Stout SC, Clark GB, Archer-Evans S, Roux SJ (2003). Rapid and efficient suppression of gene expression in a single-cell model system, Ceratopteris richardii. Plant Physiol 131, 1165-1168. |
| [95] |
Strain E, Hass B, Banks JA (2001). Characterization of mutations that feminize gametophytes of the fern Ceratopteris. Genetics 159, 1271-1281.
DOI PMID |
| [96] | Sun J, Li GS (2020). Leaf dorsoventrality candidate gene CpARF4 has conserved expression pattern but divergent tasiR-ARF regulation in the water fern Ceratopteris pteridoides. Am J Bot 107, 1470-1480. |
| [97] | Sun JY, Morita H, Chen GS, Noguchi H, Abe I (2012). Molecular cloning and characterization of copper amine oxidase from Huperzia serrata. Bioorg Med Chem Lett 22, 5784-5790. |
| [98] | Sun LL, Li Y, Li SS, Wu XJ, Hu BZ, Chang Y (2014). Identification and characterisation of DfCHS, a chalcone synthase gene regulated by temperature and ultraviolet in Dryopteris fragrans. Cell Mol Biol 60, 1-7. |
| [99] | Sun YQ, Shang LG, Zhu QH, Fan LJ, Guo LB (2022). Twenty years of plant genome sequencing: achievements and challenges. Trends Plant Sci 27, 391-401. |
| [100] | Sundaram S, Wu S, Ma LQ, Rathinasabapathi B (2009). Expression of a Pteris vittata glutaredoxin PvGRX5 in transgenic Arabidopsis thaliana increases plant arsenic tolerance and decreases arsenic accumulation in the leaves. Plant Cell Environ 32, 851-858. |
| [101] | Sztein AE, Cohen JD, Slovin JP, Cooke TJ (1995). Auxin metabolism in representative land plants. Am J Bot 82, 1514-1521. |
| [102] |
Tanaka J, Yano K, Aya K, Hirano K, Takehara S, Koketsu E, Ordonio RL, Park SH, Nakajima M, Ueguchi-Tanaka M, Matsuoka M (2014). Antheridiogen determines sex in ferns via a spatiotemporally split gibberellin synthesis pathway. Science 346, 469-473.
DOI PMID |
| [103] | Tomei EJ, Wolniak SM (2016). Kinesin-2 and kinesin-9 have a typical functions during ciliogenesis in the male gametophyte of Marsilea vestita. BMC Cell Biol 17, 29. |
| [104] |
VanBuren R, Wai CM, Ou SJ, Pardo J, Bryant D, Jiang N, Mockler TC, Edger P, Michael TP (2018). Extreme haplotype variation in the desiccation-tolerant clubmoss Selaginella lepidophylla. Nat Commun 9, 13.
DOI PMID |
| [105] | Vasco A, Ambrose BA (2020). Simple and divided leaves in ferns: exploring the genetic basis for leaf morphology differences in the genus Elaphoglossum (Dryopteridaceae). Int J Mol Sci 21, 5180. |
| [106] |
Vasco A, Smalls TL, Graham SW, Cooper ED, Wong GKS, Stevenson DW, Moran RC, Ambrose BA (2016). Challenging the paradigms of leaf evolution: class III HD-Zips in ferns and lycophytes. New Phytol 212, 745-758.
DOI PMID |
| [107] | Wang FG, Wang AH, Bai CK, Jin DM, Nie LY, Harris AJ, Che L, Wang JJ, Li SY, Xu L, Shen H, Gu YF, Shang H, Duan L, Zhang XC, Chen HF, Yan YH (2022). Genome size evolution of the extant lycophytes and ferns. Plant Divers 44, 141-152. |
| [108] | Wang J, Ding N, Wu Y, Shi XP, Qi BW, Liu X, Wang XH, Li J, Tu PF, Shi SP (2020). Enzymatic synthesis of 2-hydroxy-4H-quinolizin-4-one scaffolds by integrating coenzyme a ligases and a type III PKS from Huperzia serrata. RSC Adv 10, 23566-23572. |
| [109] | Wang J, Wang XH, Liu X, Li J, Shi XP, Song YL, Zeng KW, Zhang L, Tu PF, Shi SP (2016). Synthesis of unnatural 2-substituted quinolones and 1,3-diketones by a member of type III polyketide synthases from Huperzia serrata. Org Lett 18, 3550-3553. |
| [110] | Weng JK, Akiyama T, Bonawitz ND, Li X, Ralph J, Chapple C (2010a). Convergent evolution of syringyl lignin biosynthesis via distinct pathways in the lycophyte Selaginella and flowering plants. Plant Cell 22, 1033-1045. |
| [111] | Weng JK, Akiyama T, Ralph J, Chapple C (2011). Independent recruitment of an O-methyltransferase for syringyl lignin biosynthesis in Selaginella moellendorffii. Plant Cell 23, 2708-2724. |
| [112] | Weng JK, Li X, Stout J, Chapple C (2008). Independent origins of syringyl lignin in vascular plants. Proc Natl Acad Sci USA 105, 7887-7892. |
| [113] | Weng JK, Mo HP, Chapple C (2010b). Over-expression of F5H in COMT-deficient Arabidopsis leads to enrichment of an unusual lignin and disruption of pollen wall formation. Plant J 64, 898-911. |
| [114] |
Wickell D, Kuo LY, Yang HP, Dhabalia Ashok A, Irisarri I, Dadras A, de Vries S, de Vries J, Huang YM, Li Z, Barker MS, Hartwick NT, Michael TP, Li FW (2021). Underwater CAM photosynthesis elucidated by Isoetes genome. Nat Commun 12, 6348.
DOI PMID |
| [115] | Wilson CL (1953). The telome theory. Bot Rev 19, 417-437. |
| [116] | Wyder S, Rivera A, Valdés AE, Cañal MJ, Gagliardini V, Fernández H, Grossniklaus U (2020). Differential gene expression profiling of one- and two-dimensional apogamous gametophytes of the fern Dryopteris affinis ssp. affinis. Plant Physiol Biochem 148, 302-311. |
| [117] | Xu BF, Lei L, Zhu XC, Zhou YQ, Xiao YL (2017). Identification and characterization of L-lysine decarboxylase from Huperzia serrata and its role in the metabolic pathway of lycopodium alkaloid. Phytochemistry 136, 23-30. |
| [118] |
Xu XY, McGrath SP, Zhao FJ (2007). Rapid reduction of arsenate in the medium mediated by plant roots. New Phytol 176, 590-599.
DOI PMID |
| [119] | Xu ZC, Xin TY, Bartels D, Li Y, Gu W, Yao H, Liu S, Yu HY, Pu XD, Zhou JG, Xu J, Xi CC, Lei HT, Song JY, Chen SL (2018). Genome analysis of the ancient tracheophyte Selaginella tamariscina reveals evolutionary features relevant to the acquisition of desiccation tolerance. Mol Plant 11, 983-994. |
| [120] |
Yadav SK, Archana, Singh R, Singh PK, Vasudev PG (2019). Insecticidal fern protein Tma12 is possibly a lytic polysaccharide monooxygenase. Planta 249, 1987-1996.
DOI PMID |
| [121] | Yan HL, Gao YW, Wu LL, Wang LY, Zhang T, Dai CH, Xu WX, Feng L, Ma M, Zhu YG, He ZY (2019). Potential use of the Pteris vittata arsenic hyperaccumulation-regulation network for phytoremediation. J Hazard Mater 368, 386-396. |
| [122] | Yang MQ, You WJ, Wu SW, Fan Z, Xu BF, Zhu ML, Li X, Xiao YL (2017). Global transcriptome analysis of Huperzia serrata and identification of critical genes involved in the biosynthesis of huperzine A. BMC Genomics 18, 245. |
| [123] | Yobi A, Wone BWM, Xu WX, Alexander DC, Guo LN, Ryals JA, Oliver MJ, Cushman JC (2013). Metabolomic profiling in Selaginella lepidophylla at various hydration states provides new insights into the mechanistic basis of desiccation tolerance. Mol Plant 6, 369-385. |
| [124] |
Yokota T, Ohnishi T, Shibata K, Asahina M, Nomura T, Fujita T, Ishizaki K, Kohchi T (2017). Occurrence of brassinosteroids in non-flowering land plants, liverwort, moss, lycophyte and fern. Phytochemistry 136, 46-55.
DOI PMID |
| [125] | Youngstrom CE, Geadelmann LF, Irish EE, Cheng CL (2019). A fern WUSCHEL-RELATED HOMEOBOX gene functions in both gametophyte and sporophyte generations. BMC Plant Biol 19, 416. |
| [126] | Youngstrom CE, Withers KA, Irish EE, Cheng CL (2022). Vascular function of the T3/modern clade WUSCHEL- Related HOMEOBOX transcription factor genes predate apical meristem-maintenance function. BMC Plant Biol 22, 210. |
| [127] |
Yu J, Zhang YY, Liu W, Wang H, Wen ST, Zhang YJ, Xu L (2020). Molecular evolution of auxin-mediated root initiation in plants. Mol Biol Evol 37, 1387-1393.
DOI PMID |
| [128] | Yu JG, Tang JY, Wei R, Lan MF, Xiang RC, Zhang XC, Xiang QP (2023). The first homosporous lycophyte genome revealed the association between the recent dynamic accumulation of LTR-RTs and genome size variation. Plant Mol Biol 112, 325-340. |
| [129] |
Yu Y, Jia TR, Chen XM (2017). The ‘how’ and ‘where’ of plant microRNAs. New Phytol 216, 1002-1017.
DOI PMID |
| [130] | Zhang J, Liu JY, Zheng FB, Yu M, Shabala S, Song WY (2022). Comparative analysis of arsenic transport and tolerance mechanisms: evolution from prokaryote to higher plants. Cells 11, 2741. |
| [131] | Zhang XS, Ni R, Wang PY, Zhu TT, Sun CJ, Lou HX, Cheng AX (2019). Isolation and functional characterization of two Caffeoyl Coenzyme A 3-O-methyltransferases from the fern species Polypodiodes amoena. Plant Physiol Biochem 136, 169-177. |
| [132] | Zhong Y, Liu YB, Wu W, Chen JF, Sun CY, Liu HM, Shu JP, Ebihara A, Yan YH, Zhou RC, Schneider H (2022). Genomic insights into genetic diploidization in the homosporous fern Adiantum nelumboides. Genome Biol Evol 14, evac127. |
| [133] | Zhou Y, Gao L, Wang B, Wang T (2013). Molecular cloning and characterization of three cryptochrome genes from the fern Asplenium yunnanense. Plant Physiol Biochem 67, 71-76. |
| [134] | Zumajo-Cardona C, Vasco A, Ambrose BA (2019). The evolution of the KANADI gene family and leaf development in lycophytes and ferns. Plants 8, 313. |
| [1] | 黄 承玲, Han Li Rong, Ling Qing Hong, Xiong Yang Sheng, Ling Tian Xiao, Xia Guowei Xia Guowei, Ren Chen Zheng, Wei Zhou. Study on genetic conservation of Rhododendron liboense based on SNP molecular markers,a plant species with extremely small populations [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [2] | Lu Zi-Jia, Wang Tian-Rui, Zheng Si-Si, Cao Jian-Guo, Kozlowski Gregor, Song Yi-Gang. Environmental adaptive genetic variation and genetic vulnerability of relict plant Pterocarya hupehensis [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [3] | Chuanyong Wang, Dian Zhuang, Zhengda Song, Henghua Zhai, Naiwei Li, Fan Zhang. Structural and Comparative Analysis of the Complete Chloroplast Genome and Phylogenetic Inference of the Aronia melanocarpa [J]. Chinese Bulletin of Botany, 2025, 60(4): 1-0. |
| [4] | Ziyun Wang, Yanwen Lv, Yu Xiao, Chao Wu, Xinsheng Hu. Advances in Regulation and Evolutionary Mechanisms of Plant Gene Expression [J]. Chinese Bulletin of Botany, 2025, 60(4): 1-0. |
| [5] | Jingjing Li, Yanfei Li, Anqi Wang, Jiaying Wang, Chengyan Deng, Min Lu, Jianying Ma, Silan Dai. Establishment of Regeneration and Genetic Transformation System for Chrysanthemum Cultivar ‘Wandai Fengguang’ [J]. Chinese Bulletin of Botany, 2025, 60(4): 1-0. |
| [6] | LU Zhen, XIE Guang-Jie, Qaisar KHAN, QIN Ying, HUANG Yu-Yan, GUO Dao-Jun, YANG Ting-Ting, YANG Li-Tao, XING Yong-Xiu, LI Yang-Rui, WANG Zhen. Burkholderia strains enhance the tolerance of sugarcane to aluminum stress by improving the physiological adaptability and regulating the expression of aluminum responsive genes [J]. Chin J Plant Ecol, 2025, 49(3): 475-487. |
| [7] | Zhang Ruli, Li Dezhu, Zhang Yuxiao. Population Genetic Structure and Climate Adaptation Analysis of Brachystachyum densiflorum [J]. Chinese Bulletin of Botany, 2025, 60(3): 407-424. |
| [8] | Tong Miao, Wang Huan, Zhang Wenshuang, Wang Chao, Song Jianxiao. Distribution characteristics of antibiotic resistance genes in soil bacterial communities exposed to heavy metal pollution [J]. Biodiv Sci, 2025, 33(3): 24101-. |
| [9] | Xiong Lianglin, Liang Guolu, Guo Qigao, Jing Danlong. Advances in the Regulation of Alternative Splicing of Genes in Plants in Response to Abiotic Stress [J]. Chinese Bulletin of Botany, 2025, 60(3): 435-448. |
| [10] | Zeng Wendan, Yan Huabing, Wu Zhengdan, Shang Xiaohong, Cao Sheng, Lu Liuying, Xiao Liang, Shi Pingli, Cheng Dong, Long Ziyuan, Li Jieyu. Agrobacterium rhizogenes-mediated Transformation System of Pueraria lobata Hairy Roots [J]. Chinese Bulletin of Botany, 2025, 60(3): 425-434. |
| [11] | Yang Li, Qu Xitong, Chen Zihang, Zou Tingting, Wang Quanhua, Wang Xiaoli. Identification of the Spinach AT-hook Gene Family and Analysis of Expression Profiles [J]. Chinese Bulletin of Botany, 2025, 60(3): 377-392. |
| [12] | Lin Zhen, Xiang Jiabao, Cai Hejiayi, Gao Bei, Yang Jintao, Li Junyi, Zhou Qingsong, Huang Xiaolei, Deng Jun. Mitochondrial genomic data for seven Hemipteran species [J]. Biodiv Sci, 2025, 33(2): 24434-. |
| [13] | Liu Zhixiang, Xie Hua, Zhang Hui, Huang Xiaolei. Functional diversity and regulation of cuticular hydrocarbons in social insects [J]. Biodiv Sci, 2025, 33(2): 24302-. |
| [14] | Zhigang Yang, Pengcheng Zhang, Haiwen Chang, Liru Kang, Yi Zuo, Haoxin Xiang, Fengying Han. Genetic Diversity Analysis of Pepper Germplasms Based on Morphological Traits and SSR Markers [J]. Chinese Bulletin of Botany, 2025, 60(2): 218-234. |
| [15] | YIN Si, YANG Yi-Ting, LU Rui-Ling, NIAN Rui, HAO Zhuan, GAO Yong. Phylogeographic study of natural populations of Amorphophallus yunnanensis (Araceae) in China [J]. Chin J Plant Ecol, 2025, 49(2): 308-319. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||