

Chinese Bulletin of Botany ›› 2023, Vol. 58 ›› Issue (6): 882-892.DOI: 10.11983/CBB23100 cstr: 32102.14.CBB23100
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Qiwei Jia1,†, Qianqian Zhong1,†, Yujia Gu1, Tianqi Lu1, Wei Li1, Shuai Yang1, Chaoyu Zhu1, Chengxiang Hu1, Sanfeng Li2, Yuexing Wang2,*( ), Yuchun Rao1,*(
), Yuchun Rao1,*( )
)
Received:2023-07-28
Accepted:2023-11-02
Online:2023-11-01
Published:2023-11-27
Contact:
* E-mail: wangyuexing@caas.cn;ryc@zjnu.cn
About author:† These authors contributed equally to this paper
Qiwei Jia, Qianqian Zhong, Yujia Gu, Tianqi Lu, Wei Li, Shuai Yang, Chaoyu Zhu, Chengxiang Hu, Sanfeng Li, Yuexing Wang, Yuchun Rao. Mapping of QTL for Cell Wall Related Components in Rice Stem and Analysis of Candidate Genes[J]. Chinese Bulletin of Botany, 2023, 58(6): 882-892.
| Primer name | Sequence (5′-3′) | Tm (°C) | Length (bp) | Size (bp) |
|---|---|---|---|---|
| LOC_Os01g04930-F-qrt | GAGGGGGAACTGGTTCATGG | 60.00 | 20 | 164 |
| LOC_Os01g04930-R-qrt | CGAGGTCCCTGAAGTGGTTC | 60.00 | 20 | |
| LOC_Os01g08440-F-qrt | AGCAACGACCTCTTCAAGCA | 59.90 | 20 | 178 |
| LOC_Os01g08440-R-qrt | TCCTCAGCACCCACAAGAAC | 59.90 | 20 | |
| LOC_Os01g09850-F-qrt | CTCCTCCTACCAGCAACGTG | 60.10 | 20 | 180 |
| LOC_Os01g09850-R-qrt | GACCCAAGAAGTCCCTGGTG | 60.00 | 20 | |
| LOC_Os02g56460-F-qrt | CTGCTGGAGCGACTACGAAT | 59.90 | 20 | 182 |
| LOC_Os02g56460-R-qrt | CGTGGTTGGTGCTGAAGTTG | 60.00 | 20 | |
| LOC_Os02g58480-F-qrt | ACCGTCGGACTACCTCAAGA | 60.00 | 20 | 187 |
| LOC_Os02g58480-R-qrt | GTCGGTCAGCTTGGAAGACA | 60.00 | 20 | |
| LOC_Os02g58590-F-qrt | GCCTCGCACAAGCAGAAAAA | 60.00 | 20 | 183 |
| LOC_Os02g58590-R-qrt | AGGTCTGCACGCTCCTTTTT | 60.00 | 20 | |
| LOC_Os04g52280-F-qrt | GGATGAGGGTGACGGTGATC | 59.90 | 20 | 179 |
| LOC_Os04g52280-R-qrt | TGCCAGGTAAGAGTGCATGG | 60.00 | 20 | |
| LOC_Os08g14760-F-qrt | AATACCAGTCGCCTTCGTGG | 60.10 | 20 | 186 |
| LOC_Os08g14760-R-qrt | AGATGTTGCAGCTGCTTCCT | 60.00 | 20 | |
| LOC_Os11g04400-F-qrt | CAGTGCACCCATGGAGGATT | 60.00 | 20 | 183 |
| LOC_Os11g04400-R-qrt | TTCTCCAACATCTCGCCGAC | 60.10 | 20 | |
| LOC_Os12g29300-F-qrt | GTCTTCTTCGACTGCACCGA | 60.00 | 20 | 155 |
| LOC_Os12g29300-R-qrt | TCTTCTCCGAGTAGGCGACA | 60.00 | 20 | |
| LOC_Os12g36890-F-qrt | CTTCACCTCCGTGTTCCTCC | 60.00 | 20 | 162 |
| LOC_Os12g36890-R-qrt | CCGGACCACTTGATCTCGAG | 59.90 | 20 | |
| LOC_Os12g41720-F-qrt | CAACTACGTCCGCATCCAGG | 60.80 | 20 | 163 |
| LOC_Os12g41720-R-qrt | CCGGTTTGACTTCTCCGACA | 60.00 | 20 | |
| LOC_Os12g41780-F-qrt | TTCACTGGCATCCCGAAGTC | 60.00 | 20 | 187 |
| LOC_Os12g41780-R-qrt | CAATGCCTGATCTTCGCAGC | 60.00 | 20 |
Table 1 Primer sequences for qRT-PCR
| Primer name | Sequence (5′-3′) | Tm (°C) | Length (bp) | Size (bp) |
|---|---|---|---|---|
| LOC_Os01g04930-F-qrt | GAGGGGGAACTGGTTCATGG | 60.00 | 20 | 164 |
| LOC_Os01g04930-R-qrt | CGAGGTCCCTGAAGTGGTTC | 60.00 | 20 | |
| LOC_Os01g08440-F-qrt | AGCAACGACCTCTTCAAGCA | 59.90 | 20 | 178 |
| LOC_Os01g08440-R-qrt | TCCTCAGCACCCACAAGAAC | 59.90 | 20 | |
| LOC_Os01g09850-F-qrt | CTCCTCCTACCAGCAACGTG | 60.10 | 20 | 180 |
| LOC_Os01g09850-R-qrt | GACCCAAGAAGTCCCTGGTG | 60.00 | 20 | |
| LOC_Os02g56460-F-qrt | CTGCTGGAGCGACTACGAAT | 59.90 | 20 | 182 |
| LOC_Os02g56460-R-qrt | CGTGGTTGGTGCTGAAGTTG | 60.00 | 20 | |
| LOC_Os02g58480-F-qrt | ACCGTCGGACTACCTCAAGA | 60.00 | 20 | 187 |
| LOC_Os02g58480-R-qrt | GTCGGTCAGCTTGGAAGACA | 60.00 | 20 | |
| LOC_Os02g58590-F-qrt | GCCTCGCACAAGCAGAAAAA | 60.00 | 20 | 183 |
| LOC_Os02g58590-R-qrt | AGGTCTGCACGCTCCTTTTT | 60.00 | 20 | |
| LOC_Os04g52280-F-qrt | GGATGAGGGTGACGGTGATC | 59.90 | 20 | 179 |
| LOC_Os04g52280-R-qrt | TGCCAGGTAAGAGTGCATGG | 60.00 | 20 | |
| LOC_Os08g14760-F-qrt | AATACCAGTCGCCTTCGTGG | 60.10 | 20 | 186 |
| LOC_Os08g14760-R-qrt | AGATGTTGCAGCTGCTTCCT | 60.00 | 20 | |
| LOC_Os11g04400-F-qrt | CAGTGCACCCATGGAGGATT | 60.00 | 20 | 183 |
| LOC_Os11g04400-R-qrt | TTCTCCAACATCTCGCCGAC | 60.10 | 20 | |
| LOC_Os12g29300-F-qrt | GTCTTCTTCGACTGCACCGA | 60.00 | 20 | 155 |
| LOC_Os12g29300-R-qrt | TCTTCTCCGAGTAGGCGACA | 60.00 | 20 | |
| LOC_Os12g36890-F-qrt | CTTCACCTCCGTGTTCCTCC | 60.00 | 20 | 162 |
| LOC_Os12g36890-R-qrt | CCGGACCACTTGATCTCGAG | 59.90 | 20 | |
| LOC_Os12g41720-F-qrt | CAACTACGTCCGCATCCAGG | 60.80 | 20 | 163 |
| LOC_Os12g41720-R-qrt | CCGGTTTGACTTCTCCGACA | 60.00 | 20 | |
| LOC_Os12g41780-F-qrt | TTCACTGGCATCCCGAAGTC | 60.00 | 20 | 187 |
| LOC_Os12g41780-R-qrt | CAATGCCTGATCTTCGCAGC | 60.00 | 20 |
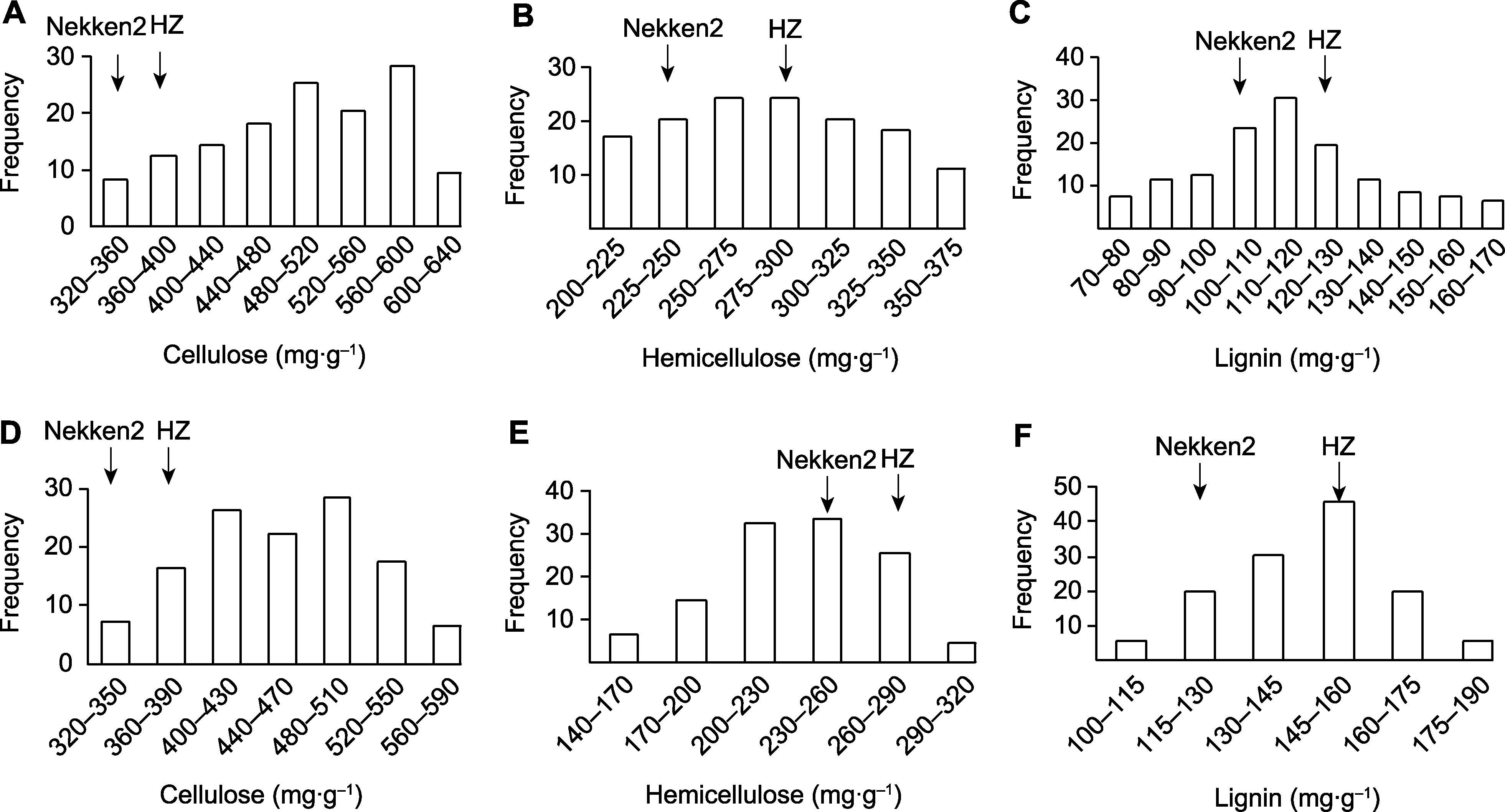
Figure 2 The contents of stem cell wall related components in rice recombinant inbred line population (A) The content of cellulose components in 2020; (B) The content of hemicellulose components in 2020; (C) The content of lignin components in 2020; (D) The content of cellulose components in 2021; (E) The content of hemicellulose components in 2021; (F) The content of lignin components in 2021
| Year | Cell wall component | QTL | Chromosome | Physical distance (bp) | Position of support (cM) | Likelihood of odd (LOD) |
|---|---|---|---|---|---|---|
| 2020 | Cellulose | qCel202 | 2 | 35381094-35599368 | 151.67-152.60 | 3.48 |
| qCel204 | 4 | 29276416-29507404 | 125.50-126.49 | 2.82 | ||
| Hemicellulose | qHem201 | 1 | 2070966-6568005 | 8.88-28.16 | 5.02 | |
| qHem203 | 3 | 29806859-30321190 | 127.77-129.99 | 2.53 | ||
| qHem208 | 8 | 4587694-8917564 | 19.67-38.23 | 3.48 | ||
| qHem2012-1 | 12 | 15484322-15779533 | 66.38-67.64 | 2.94 | ||
| qHem2012-2 | 12 | 22426037-27516985 | 96.13-117.95 | 5.52 | ||
| Lignin | qLig202 | 2 | 33583953-35902152 | 143.97-153.90 | 3.03 | |
| qLig204-1 | 4 | 27558155-28380156 | 118.13-121.66 | 3.19 | ||
| qLig204-2 | 4 | 31066903-31212801 | 133.18-133.80 | 3.25 | ||
| qLig2011 | 11 | 1449031-5404218 | 6.21-23.17 | 3.46 | ||
| qLig2012-1 | 12 | 24319945-24511904 | 104.25-105.66 | 3.45 | ||
| qLig2012-2 | 12 | 27434341-27516085 | 117.60-117.95 | 2.60 | ||
| 2021 | Cellulose | qCel211 | 1 | 2192312-2834929 | 9.39-12.15 | 3.34 |
| qCel2112 | 12 | 25510919-27516085 | 109.35-117.95 | 3.94 | ||
| Hemicellulose | qHem211 | 1 | 5699636-6152981 | 24.43-26.37 | 3.34 | |
| qHem213-1 | 3 | 8841716-8989609 | 37.90-38.53 | 2.55 | ||
| qHem213-2 | 3 | 20915784-21229182 | 89.66-91.00 | 3.07 | ||
| qHem214 | 4 | 2647701-5256057 | 11.34-22.53 | 3.81 | ||
| qHem218 | 8 | 1430252-1451568 | 6.13-6.22 | 2.50 | ||
| qHem2112-1 | 12 | 7276559-7573232 | 31.19-32.46 | 2.55 | ||
| qHem2112-2 | 12 | 17268215-18172857 | 74.02-77.90 | 3.40 | ||
| Lignin | qLig215-1 | 5 | 327464-853216 | 1.40-3.65 | 2.90 | |
| qLig215-2 | 5 | 25519701-27453301 | 109.39-117.68 | 3.35 |
Table 2 QTL analysis of stem cell wall related components in rice recombinant inbred line population
| Year | Cell wall component | QTL | Chromosome | Physical distance (bp) | Position of support (cM) | Likelihood of odd (LOD) |
|---|---|---|---|---|---|---|
| 2020 | Cellulose | qCel202 | 2 | 35381094-35599368 | 151.67-152.60 | 3.48 |
| qCel204 | 4 | 29276416-29507404 | 125.50-126.49 | 2.82 | ||
| Hemicellulose | qHem201 | 1 | 2070966-6568005 | 8.88-28.16 | 5.02 | |
| qHem203 | 3 | 29806859-30321190 | 127.77-129.99 | 2.53 | ||
| qHem208 | 8 | 4587694-8917564 | 19.67-38.23 | 3.48 | ||
| qHem2012-1 | 12 | 15484322-15779533 | 66.38-67.64 | 2.94 | ||
| qHem2012-2 | 12 | 22426037-27516985 | 96.13-117.95 | 5.52 | ||
| Lignin | qLig202 | 2 | 33583953-35902152 | 143.97-153.90 | 3.03 | |
| qLig204-1 | 4 | 27558155-28380156 | 118.13-121.66 | 3.19 | ||
| qLig204-2 | 4 | 31066903-31212801 | 133.18-133.80 | 3.25 | ||
| qLig2011 | 11 | 1449031-5404218 | 6.21-23.17 | 3.46 | ||
| qLig2012-1 | 12 | 24319945-24511904 | 104.25-105.66 | 3.45 | ||
| qLig2012-2 | 12 | 27434341-27516085 | 117.60-117.95 | 2.60 | ||
| 2021 | Cellulose | qCel211 | 1 | 2192312-2834929 | 9.39-12.15 | 3.34 |
| qCel2112 | 12 | 25510919-27516085 | 109.35-117.95 | 3.94 | ||
| Hemicellulose | qHem211 | 1 | 5699636-6152981 | 24.43-26.37 | 3.34 | |
| qHem213-1 | 3 | 8841716-8989609 | 37.90-38.53 | 2.55 | ||
| qHem213-2 | 3 | 20915784-21229182 | 89.66-91.00 | 3.07 | ||
| qHem214 | 4 | 2647701-5256057 | 11.34-22.53 | 3.81 | ||
| qHem218 | 8 | 1430252-1451568 | 6.13-6.22 | 2.50 | ||
| qHem2112-1 | 12 | 7276559-7573232 | 31.19-32.46 | 2.55 | ||
| qHem2112-2 | 12 | 17268215-18172857 | 74.02-77.90 | 3.40 | ||
| Lignin | qLig215-1 | 5 | 327464-853216 | 1.40-3.65 | 2.90 | |
| qLig215-2 | 5 | 25519701-27453301 | 109.39-117.68 | 3.35 |
| Gene ID | Chromosome | Function | Cloned or not |
|---|---|---|---|
| LOC_Os01g04930 | 1 | MYB transcription factor | Not cloned |
| LOC_Os01g08440 | 1 | UDP glucoside transferase | Not cloned |
| LOC_Os01g09850 | 1 | C2H2-type zinc finger transcription factor | Cloned (Huang et al., |
| LOC_Os02g56460 | 2 | Cinnamyl CoA reductase | Cloned (Kawasaki et al., |
| LOC_Os02g58480 | 2 | Sucrose synthase | Not cloned |
| LOC_Os02g58590 | 2 | N-acetylglucosaminyltransferase I | Cloned (Fanata et al., |
| LOC_Os04g52280 | 4 | Cinnamyl alcohol dehydrogenase | Cloned (Li et al., |
| LOC_Os08g14760 | 8 | 4-coumarate:coenzyme A ligase | Cloned (Gu et al., |
| LOC_Os11g04400 | 11 | GRAS family transcription factor | Not cloned |
| LOC_Os12g29300 | 12 | CESA10-cellulase synthesis | Not cloned |
| LOC_Os12g36890 | 12 | Cellulose synthase-like | Cloned (Hu et al., |
| LOC_Os12g41720 | 12 | Patatin-related phospholipase A | Not cloned |
| LOC_Os12g41780 | 12 | Glycosyltransferase | Not cloned |
Table 3 Rice stem cell wall related components candidate genes and their function
| Gene ID | Chromosome | Function | Cloned or not |
|---|---|---|---|
| LOC_Os01g04930 | 1 | MYB transcription factor | Not cloned |
| LOC_Os01g08440 | 1 | UDP glucoside transferase | Not cloned |
| LOC_Os01g09850 | 1 | C2H2-type zinc finger transcription factor | Cloned (Huang et al., |
| LOC_Os02g56460 | 2 | Cinnamyl CoA reductase | Cloned (Kawasaki et al., |
| LOC_Os02g58480 | 2 | Sucrose synthase | Not cloned |
| LOC_Os02g58590 | 2 | N-acetylglucosaminyltransferase I | Cloned (Fanata et al., |
| LOC_Os04g52280 | 4 | Cinnamyl alcohol dehydrogenase | Cloned (Li et al., |
| LOC_Os08g14760 | 8 | 4-coumarate:coenzyme A ligase | Cloned (Gu et al., |
| LOC_Os11g04400 | 11 | GRAS family transcription factor | Not cloned |
| LOC_Os12g29300 | 12 | CESA10-cellulase synthesis | Not cloned |
| LOC_Os12g36890 | 12 | Cellulose synthase-like | Cloned (Hu et al., |
| LOC_Os12g41720 | 12 | Patatin-related phospholipase A | Not cloned |
| LOC_Os12g41780 | 12 | Glycosyltransferase | Not cloned |
| [1] | 白勇, 巩威, 刘天昀, 朱玉贤 (2003). 水稻肉桂酰辅酶A还原酶基因的克隆与表达分析. 科学通报 48, 1780-1784. |
| [2] |
陈桂华, 邓化冰, 张桂莲, 唐文帮, 黄璜 (2016). 水稻茎秆性状与抗倒性的关系及配合力分析. 中国农业科学 49, 407-417.
DOI |
| [3] | 代建秀, 唐子清, 吴春文, 史培康, 韦良艳, 施洁, 黎晟, 罗继景, 蔡中全 (2017). 水稻特异粗茎相关性状QTL的初步定位分析. 分子植物育种 15, 1395-1402. |
| [4] | 丰安徽 (2017). 控制水稻茎秆粗细的OsTB1等位基因发掘与功能分析. 硕士论文. 成都: 四川农业大学. pp. 1-47. |
| [5] | 冯春燕 (2010). 植物4-香豆酸:辅酶A连接酶(4CL)研究进展. 现代农业科技 (8), 39-40. |
| [6] | 韩雷锋, 周燃, 周涛, 林翠香, 甘泉, 倪大虎, 石英尧, 宋丰顺 (2023). 水稻抗倒伏和产量性状的相关性分析及QTLs定位. 生物学杂志 40(2), 65-70. |
| [7] | 胡文冉, 李晓东, 周小云, 杨洋, 李波, 李晓荣 (2019). 棉花肉桂醇脱氢酶基因在棉纤维中的表达及对其结构组分的影响. 西北植物学报 39, 1114-1120. |
| [8] |
金佳怡, 罗怿婷, 杨惠敏, 芦涛, 叶涵斐, 谢继毅, 王珂欣, 陈芊羽, 方媛, 王跃星, 饶玉春 (2023). 水稻叶绿素含量QTL定位与候选基因表达分析. 植物学报 58, 394-403.
DOI |
| [9] | 鞠晓晨, 胡杰, 高冠军, 张庆路, 何予卿 (2016). 水稻茎秆抗倒伏相关QTL定位与分析. 分子植物育种 14, 475-481. |
| [10] | 李丰成 (2015). 植物细胞壁结构特征与生物质高效利用分子机理研究. 博士论文. 武汉: 华中农业大学. pp. 1-136. |
| [11] |
刘畅, 李来庚 (2016). 水稻抗倒伏性状的分子机理研究进展. 中国水稻科学 30, 216-222.
DOI |
| [12] | 刘子玮 (2018). 水稻GRAS转录因子家族的系统分析与功能研究. 硕士论文. 南京: 南京大学. pp. 1-44. |
| [13] | 卢雯瑩, 崔贺云, 高杉, 崔顺梅, 吴营照 (2020). 肉桂醇脱氢酶基因研究进展及其在小麦抗倒伏中的应用. 天津农业科学 26(10), 7-10. |
| [14] | 罗兵, 韩永笑, 张淇鑫, 郭浩, 李红梅, 王静, 杨志刚, 孙海燕 (2018). 水稻UDP-N-乙酰葡萄糖胺酰基转移酶基因(OsLpxA) RNA干扰载体的构建及遗传转化. 分子植物育种 16, 5311-5317. |
| [15] | 穆平, 李自超, 李春平, 张洪亮, 王象坤 (2004). 水、旱条件下水稻茎秆主要抗倒伏性状的QTL分析. 遗传学报 31, 717-723. |
| [16] | 苏仕华, 王珏, 孙成亮, 成英, 卢红 (2008). 水稻倒伏对产量影响的调查与分析. 北方水稻 38, 41-43. |
| [17] | 王士强, 顾春梅, 沈巧梅, 赵黎明, 王丽萍, 王贺 (2011). 水稻倒伏发生规律及防御技术的研究进展. 北方水稻 41, 69-72. |
| [18] | 王晓飞, 陆展华, 刘维, 卢东柏, 王石光, 巫浩翔, 方志强, 何秀英 (2022). “绿色革命”以来水稻抗倒伏研究进展. 广东农业科学 49(3), 1-13. |
| [19] |
魏和平, 芦涛, 贾绮玮, 邓飞, 朱浩, 岂泽华, 王玉玺, 叶涵斐, 殷文晶, 方媛, 穆丹, 饶玉春 (2022). 水稻抽穗期QTL定位及候选基因分析. 植物学报 57, 588-595.
DOI |
| [20] | 吴丹, 唐冬英, 李新梅, 李丽, 赵小英, 刘选明 (2015). F-box蛋白在植物生长发育中的功能研究进展. 生命科学研究 19, 362-367. |
| [21] | 肖应辉, 罗丽华, 闫晓燕, 高艳红, 王春明, 江玲, 矢野昌裕, 翟虎渠, 万建民 (2005). 水稻品种倒伏指数QTL分析. 作物学报 31, 348-354. |
| [22] |
辛良杰, 李秀彬 (2009). 近年来我国南方双季稻区复种的变化及其政策启示. 自然资源学报 24, 58-65.
DOI |
| [23] | 许作鹏 (2017). 水稻茎秆强度相关性状QTL分析及基因克隆. 硕士论文. 扬州: 扬州大学. pp. 19-78. |
| [24] | 杨波, 杨文钰 (2011). 水稻抗倒伏研究进展. 耕作与栽培 (2), 1-5, 9. |
| [25] | 杨德卫, 朱镇, 张亚东, 林静, 陈涛, 赵凌, 朱文银, 王才林 (2009). 基于CSSL的水稻穗颈长度QTL的代换作图. 遗传 31, 741-747. |
| [26] | 杨窑龙, 饶玉春, 李赓觅, 黄李超, 冷语佳, 张光恒, 高振宇, 胡江, 朱丽, 郭龙彪, 钱前, 曾大力 (2011). 水稻茎秆相关性状遗传分析. 分子植物育种 9, 160-168. |
| [27] | 袁新捷 (2021). 水稻抗倒伏研究及相关性状QTL初步定位分析. 硕士论文. 武汉: 华中农业大学. pp. 9-61. |
| [28] | 赵新勇, 邵在胜, 吴艳珍, 赵轶鹏, 王余龙, 王云霞, 杨连新 (2018). 花后人为模拟倒伏对超级稻生长、产量和品质的影响. 中国生态农业学报 26, 980-989. |
| [29] | 周雷 (2018). 一个水稻茎粗抗倒QTL的定位及其候选基因分析. 硕士论文. 成都: 四川农业大学. pp. 2-40. |
| [30] | Fanata WID, Lee KH, Son BH, Yoo JY, Harmoko R, Ko KS, Ramasamy NK, Kim KH, Oh DB, Jung HS, Kim JY, Lee SY, Lee KO (2013). N-glycan maturation is crucial for cytokinin-mediated development and cellulose synthesis in Oryza sativa. Plant J 73, 966-979. |
| [31] |
Gui JS, Shen JH, Li LG (2011). Functional characterization of evolutionarily divergent 4-coumarate: coenzyme A ligases in rice. Plant Physiol 157, 574-586.
DOI URL |
| [32] |
Haigler CH, Ivanova-Datcheva M, Hogan PS, Salnikov VV, Hwang S, Martin K, Delmer DP (2001). Carbon partitioning to cellulose synthesis. Plant Mol Biol 47, 29-51.
PMID |
| [33] |
Hu J, Zhu L, Zeng DL, Gao ZY, Guo LB, Fang YX, Zhang GH, Dong GJ, Yan MX, Liu J, Qian Q (2010). Identification and characterization of NARROW AND ROLLED LEAF 1, a novel gene regulating leaf morphology and plant architecture in rice. Plant Mol Biol 73, 283-292.
DOI URL |
| [34] |
Huang P, Yoshida H, Yano K, Kinoshita S, Kawai K, Koketsu E, Hattori M, Takehara S, Huang J, Hirano K, Ordonio RL, Matsuoka M, Ueguchi-Tanaka M (2018). OsIDD2, a zinc finger and INDETERMINATE DOMAIN protein, regulates secondary cell wall formation. J Integr Plant Biol 60, 130-143.
DOI |
| [35] |
Ishimaru K, Togawa E, Ookawa T, Kashiwagi T, Madoka Y, Hirotsu N (2008). New target for rice lodging resistance and its effect in a typhoon. Planta 227, 601-609.
DOI PMID |
| [36] |
Kashiwagi T, Togawa E, Hirotsu N, Ishimaru K (2008). Improvement of lodging resistance with QTLs for stem diameter in rice (Oryza sativa L.). Theor Appl Genet 117, 749-757.
DOI PMID |
| [37] |
Kawasaki T, Koita H, Nakatsubo T, Hasegawa K, Wakabayashi K, Takahashi H, Umemura K, Umezawa T, Shimamoto K (2006). Cinnamoyl-CoA reductase, a key enzyme in lignin biosynthesis, is an effector of small GTPase Rac in defense signaling in rice. Proc Natl Acad Sci USA 103, 230-235.
DOI PMID |
| [38] |
Li M, Xiong GY, Li R, Cui JJ, Tang D, Zhang BC, Pauly M, Cheng ZK, Zhou YH (2009a). Rice cellulose synthase- like D4 is essential for normal cell-wall biosynthesis and plant growth. Plant J 60, 1055-1069.
DOI URL |
| [39] |
Li XJ, Yang Y, Yao JL, Chen GX, Li XH, Zhang QF, Wu CY (2009b). FLEXIBLE CULM 1 encoding a cinnamyl- alcohol dehydrogenase controls culm mechanical strength in rice. Plant Mol Biol 69, 685-697.
DOI URL |
| [40] |
Liu GM, Zhang K, Ai J, Deng XJ, Hong YY, Wang XM (2015). Patatin-related phospholipase A, pPLAIIIα, modulates the longitudinal growth of vegetative tissues and seeds in rice. J Exp Bot 66, 6945-6955.
DOI PMID |
| [41] |
Livak KJ, Schmittgen TD (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2-ΔΔCT method. Methods 25, 402-408.
DOI PMID |
| [42] | McCouch SR, Cho YG, Yano M, Paul E, Blinstrub M, Morishima H, Kinoshita T (1997). Report on QTL nomenclature. Rice Genet Newsl 14, 11-13. |
| [43] |
Nixon BT, Mansouri K, Singh A, Du J, Davis JK, Lee JG, Slabaugh E, Vandavasi VG, O'Neill H, Roberts EM, Roberts AW, Yingling YG, Haigler CH (2016). Comparative structural and computational analysis supports eighteen cellulose synthases in the plant cellulose synthesis complex. Sci Rep 6, 28696.
DOI PMID |
| [44] |
Paredez AR, Somerville CR, Ehrhardt DW (2006). Visualization of cellulose synthase demonstrates functional association with microtubules. Science 312, 1491-1495.
DOI PMID |
| [45] |
Peng S, Cassman KG, Virmani SS, Sheehy J, Khush GS (1999). Yield potential trends of tropical rice since the release of IR8 and the challenge of increasing rice yield potential. Crop Sci 39, 1552-1559.
DOI URL |
| [46] |
Rosinski JA, Atchley WR (1998). Molecular evolution of the MYB family of transcription factors: evidence for polyphyletic origin. J Mol Evol 46, 74-83.
DOI PMID |
| [47] |
Song XQ, Zhang BC, Zhou YH (2011). Golgi-localized UDP-glucose transporter is required for cell wall integrity in rice. Plant Signal Behav 6, 1097-1100.
DOI PMID |
| [48] |
Van Soest PJ, Robertson JB, Lewis BA (1991). Methods for dietary fiber, neutral detergent fiber, and nonstarch polysaccharides in relation to animal nutrition. J Dairy Sci 74, 3583-3597.
DOI PMID |
| [49] |
Wang L, Wang AH, Huang XH, Zhao Q, Dong GJ, Qian Q, Sang T, Han B (2011). Mapping 49 quantitative trait loci at high resolution through sequencing-based genotyping of rice recombinant inbred lines. Theor Appl Genet 122, 327-340.
DOI PMID |
| [50] |
Zhang BL, Zhao TM, Yu WG, Kuang BQ, Yao Y, Liu TL, Chen XY, Zhang WH, Wu AM (2014). Functional conservation of the glycosyltransferase gene GT47A in the monocot rice. J Plant Res 127, 423-432.
DOI URL |
| [1] | Zhao Ling, Guan Ju, Liang Wenhua, Zhang Yong, Lu Kai, Zhao Chunfang, Li Yusheng, Zhang Yadong. Mapping of QTLs for Heat Tolerance at the Seedling Stage in Rice Based on a High-density Bin Map [J]. Chinese Bulletin of Botany, 2025, 60(3): 342-353. |
| [2] | Jinjin Lian, Luyao Tang, Yinuo Zhang, Jiaxing Zheng, Chaoyu Zhu, Yuhan Ye, Yuexing Wang, Wennan Shang, Zhenghao Fu, Xinxuan Xu, Richeng Wu, Mei Lu, Changchun Wang, Yuchun Rao. Genetic Locus Mining and Candidate Gene Analysis of Antioxidant Traits in Rice [J]. Chinese Bulletin of Botany, 2024, 59(5): 738-751. |
| [3] | Chaoyu Zhu, Chengxiang Hu, Zhenan Zhu, Zhining Zhang, Lihai Wang, Jun Chen, Sanfeng Li, Jinjin Lian, Luyao Tang, Qianqian Zhong, Wenjing Yin, Yuexing Wang, Yuchun Rao. Mapping of QTLs Associated with Rice Panicle Traits and Candidate Gene Analysis [J]. Chinese Bulletin of Botany, 2024, 59(2): 217-230. |
| [4] | Jing Xia, Yuchun Rao, Danyun Cao, Yi Wang, Linxin Liu, Yating Xu, Wangshu Mou, Dawei Xue. Research Progress on the Regulatory Mechanisms of OsACS and OsACO in Rice Ethylene Biosynthesis [J]. Chinese Bulletin of Botany, 2024, 59(2): 291-301. |
| [5] | Jiayi Jin, Yiting Luo, Huimin Yang, Tao Lu, Hanfei Ye, Jiyi Xie, Kexin Wang, Qianyu Chen, Yuan Fang, Yuexing Wang, Yuchun Rao. QTL Mapping and Expression Analysis on Candidate Genes Related to Chlorophyll Content in Rice [J]. Chinese Bulletin of Botany, 2023, 58(3): 394-403. |
| [6] | Weijun Ye, Yin Zhang, Peiran Wang, Lingling Zhang, Dongfeng Tian, Zejiang Wu, Bin Zhou. QTLs Analysis for Five Yield-related Traits in Mungbean [J]. Chinese Bulletin of Botany, 2023, 58(1): 150-158. |
| [7] | Wei Heping, Lu Tao, Jia Qiwei, Deng Fei, Zhu Hao, Qi Zehua, Wang Yuxi, Ye Hanfei, Yin Wenjing, Fang Yuan, Mu Dan, Rao Yuchun. QTL Mapping of Candidate Genes for Heading Date in Rice [J]. Chinese Bulletin of Botany, 2022, 57(5): 588-595. |
| [8] | Liu Xiaolong, Ji Ping, Yang Hongtao, Ding Yongdian, Fu Jialing, Liang Jiangxia, Yu Congcong. Priming Effect of Abscisic Acid on High Temperature Stress During Rice Heading-flowering Stage [J]. Chinese Bulletin of Botany, 2022, 57(5): 596-610. |
| [9] | Kairu Yang, Qiwei Jia, Jiayi Jin, Hanfei Ye, Sheng Wang, Qianyu Chen, Yian Guan, Chenyang Pan, Dedong Xin, Yuan Fang, Yuexing Wang, Yuchun Rao. Cloning and Functional Analysis of Rice Yellow Green Leaf Regulatory Gene YGL18 [J]. Chinese Bulletin of Botany, 2022, 57(3): 276-287. |
| [10] | Hong Yu, Jiayang Li. The Gold Will Glitter Wherever it is: Convergent Selection in Maize and Rice [J]. Chinese Bulletin of Botany, 2022, 57(2): 153-156. |
| [11] | Hanfei Ye, Wenjing Yin, Yian Guan, Kairu Yang, Qianyu Chen, Shuying Yu, Xudong Zhu, Dedong Xin, Wei Zhang, Yuexing Wang, Yuchun Rao. QTL Mapping and Candidate Gene Analysis of Vitamin E in Rice Grain [J]. Chinese Bulletin of Botany, 2022, 57(2): 157-170. |
| [12] | Jiaxin Li, Xia Li, Yinfeng Xie. Mechanism on Drought Tolerance Enhanced by Exogenous Trehalose in C4-PEPC Rice [J]. Chinese Bulletin of Botany, 2021, 56(3): 296-314. |
| [13] | Chenyang Pan, Yue Zhang, Han Lin, Qianyu Chen, Kairu Yang, Jiaji Jiang, Mengjia Li, Tao Lu, Kexin Wang, Mei Lu, Sheng Wang, Hanfei Ye, Yuchun Rao, Haitao Hu. QTL Mapping and Candidate Gene Analysis on Rice Leaf Water Potential [J]. Chinese Bulletin of Botany, 2021, 56(3): 275-283. |
| [14] | Chenyang Pan, Hanfei Ye, Weiyong Zhou, Sheng Wang, Mengjia Li, Mei Lu, Sanfeng Li, Xudong Zhu, Yuexing Wang, Yuchun Rao, Gaoxing Dai. QTL Mapping of Candidate Genes Involved in Cd Accumulation in Rice Grain [J]. Chinese Bulletin of Botany, 2021, 56(1): 25-32. |
| [15] | Wei Xuan, Guohua Xu. Genomic Basis of Rice Adaptation to Soil Nitrogen Status [J]. Chinese Bulletin of Botany, 2021, 56(1): 1-5. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||