

Chinese Bulletin of Botany ›› 2019, Vol. 54 ›› Issue (3): 385-395.DOI: 10.11983/CBB18151 cstr: 32102.14.CBB18151
• SPECIAL TOPICS • Previous Articles Next Articles
Yuekai Su,Jingren Qiu,Han Zhang,Zhenqiao Song,Jianhua Wang( )
)
Received:2018-07-02
Accepted:2018-08-09
Online:2019-05-01
Published:2019-11-24
Contact:
Jianhua Wang
Yuekai Su,Jingren Qiu,Han Zhang,Zhenqiao Song,Jianhua Wang. Recent Progress in Evolutionary Technology of CRISPR/Cas9 System for Plant Genome Editing[J]. Chinese Bulletin of Botany, 2019, 54(3): 385-395.
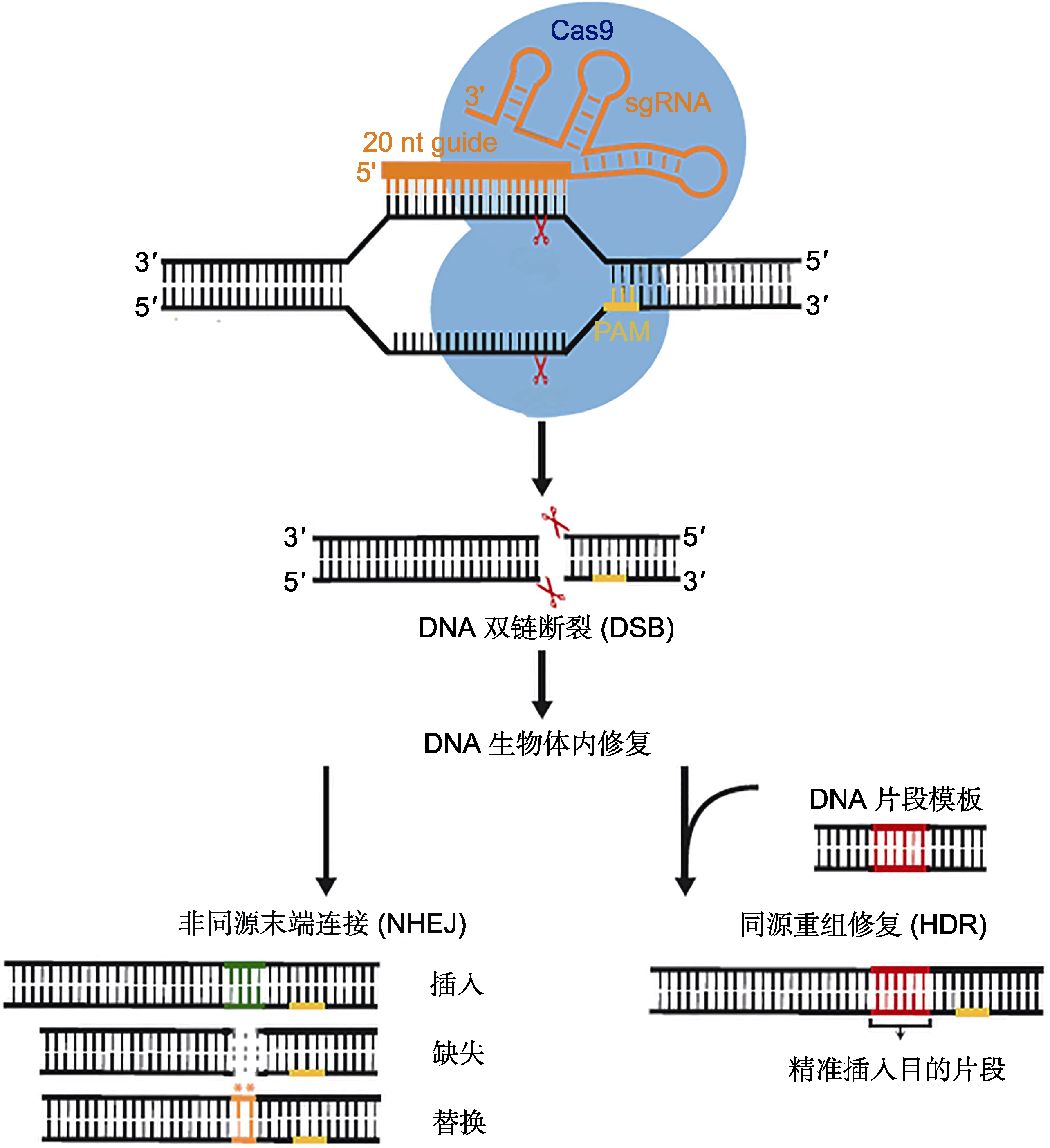
Figure 1 The schematic diagram of CRISPR/Cas9 genome editing (Jiang and Doudna, 2017)The Cas9 protein from Streptococcus pyogenes (SpCas9) and associated guide RNA (sgRNA), containing a 20 nt recognition sequence, will cleave a target sequence located upstream of the protospacer adjacent motif (PAM) region, resulting in double-strand break (DSB). Ultimately use the repair in the organism to achieve genome editing.
| [46] | Liang Z, Chen K, Zhang Y, Liu J, Yin K, Qiu JL, Gao C ( 2018). Genome editing of bread wheat using biolistic delivery of CRISPR/Cas9 in vitro transcripts or ribonucleo proteins. Nat Protoc 13, 413-430. |
| [47] |
Lowder LG, Zhou J, Zhang Y, Malzahn A, Zhong Z, Hsieh TF, Voytas DF, Zhang Y, Qi Y ( 2018). Robust transcriptional activation in plants using multiplexed CRISPR-Act 2.0 and mTALE-Act systems. Mol Plant 11, 245-256.
DOI URL |
| [48] |
Lu HP, Liu SM, Xu SL, Chen WY, Zhou X, Tan YY, Huang JZ, Shu QY ( 2017). CRISPR-S: an active interference element for a rapid and inexpensive selection of genome-edited, transgene-free rice plants. Plant Biotechnol J 15, 1371-1373.
DOI URL PMID |
| [49] |
Ma X, Zhang Q, Zhu Q, Liu W, Chen Y, Qiu R, Wang B, Yang Z, Li H, Lin Y, Xie Y, Shen R, Chen S, Wang Z, Chen Y, Guo J, Chen L, Zhao X, Dong Z, Liu YG ( 2015). A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8, 1274-1284.
DOI URL PMID |
| [50] | Marraffini LA, Sontheimer EJ ( 2008). CRISPR interference limits horizontal gene transfer in Staphylococci by targeting DNA. Science 322, 1843-1845. |
| [51] |
Meng XB, Hu XX, Liu Q, Song XG, Gao CX, Li JY, Wang KJ ( 2018). Robust genome editing of CRISPR-Cas9 at NAG PAMs in rice. Sci China Life Sci 61, 122-125.
DOI URL |
| [52] |
Mikami M, Toki S, Endo M ( 2016). Precision targeted mutagenesis via Cas9 paired nickases in rice. Plant Cell Physiol 57, 1058-1068.
DOI URL PMID |
| [53] | Miki D, Zhang W, Zeng W, Feng Z, Zhu JK ( 2018). CRISPR/Cas9-mediated gene targeting in Arabidopsis using sequential transformation. Nat Commun 9, 1967. |
| [54] |
Miller JC, Tan S, Qiao G, Barlow KA, Wang J, Xia DF, Meng X, Paschon DE, Leung E, Hinkley SJ, Dulay GP, Hua KL, Ankoudinova I, Cost GJ, Urnov FD, Zhang HS, Holmes MC, Zhang L, Gregory PD, Rebar EJ ( 2011). A tale nuclease architecture for efficient genome editing. Nat Biotechnol 29, 143-148.
DOI |
| [55] |
Mussolino C, Cathomen T ( 2013). RNA guides genome engineering. Nat Biotechnol 31, 208-209.
DOI |
| [56] | Nekrasov V, Staskawicz B, Weigel D, Jones JDG, Kamoun S ( 2013). Targeted mutagenesis in the model plant Nicotiana benthamiana using Cas9 RNA-guided endonuclease. Nat Biotechnol 31, 691-693. |
| [57] |
Peng J, Wang Y, Jiang JY, Zhou XY, Song L, Wang LL, Ding C, Qin J, Liu LP, Wang WH, Liu JQ, Huang XX, Wei H, Zhang P ( 2015). Production of human albumin in pigs through CRISPR/Cas9-mediated knockin of human cDNA into swine albumin locus in the zygotes. Sci Rep 5, 16705.
DOI URL PMID |
| [58] | Puchta H, Dujon B, Hohn B ( 1993). Homologous recombination in plant cells is enhanced by in vivo induction of double strand breaks into DNA by a site-specific endonuclease. Nucleic Acids Res 21, 5034-5040. |
| [59] | Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, Zetsche B, Shalem O, Wu X, Makarova KS, Koonin EV, Sharp PA, Zhang F ( 2015). In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186-191. |
| [60] |
Redondo P, Prieto J, Muñoz IG, Alibés A, Stricher F, Serrano L, Cabaniols JP, Daboussi F, Arnould S, Perez C, Duchateau P, Pâques F, Blanco FJ, Montoya G ( 2008). Molecular basis of xeroderma pigmentosum group C DNA recognition by engineered meganucleases. Nature 456, 107-111.
DOI URL PMID |
| [61] |
Ren B, Yan F, Kuang Y, Li N, Zhang D, Zhou X, Lin H, Zhou H ( 2018). Improved base editor for efficiently inducing genetic variations in rice with CRISPR/Cas9-guided hyperactive hAID mutant. Mol Plant 11, 623-626.
DOI URL PMID |
| [62] |
Schiml S, Fauser F, Puchta H ( 2015). The CRISPR/Cas system can be used as nuclease for in planta gene targeting and as paired nickases for directed mutagenesis in Arabidopsis resulting in heritable progeny. Plant J 80, 1139-1150.
DOI URL PMID |
| [63] |
Shan JW, Gao CX ( 2015). Research progress of genome editing and derivative technologies in plants. Hereditas 37, 953-973.
DOI URL PMID |
| [64] |
Shan Q, Wang Y, Li J, Zhang Y, Chen K, Liang Z, Zhang K, Liu J, Xi JJ, Qiu JL, Gao C ( 2013). Targeted genome modification of crop plants using a CRISPR-Cas system. Nat Biotechnol 31, 686-688.
DOI URL PMID |
| [65] | Steinert J, Schiml S, Fauser F, Puchta H ( 2015). Highly efficient heritable plant genome engineering using Cas9 orthologues from Streptococcus thermophilus and Staphylococcus aureus. Plant J 84, 1295-1305. |
| [66] |
Sun Y, Zhang X, Wu C, He Y, Ma Y, Hou H, Guo X, Du W, Zhao Y, Xia L ( 2016). Engineering herbicide-resistant rice plants through CRISPR/Cas9-mediated homologous recombination of acetolactate synthase. Mol Plant 9, 628-631.
DOI URL PMID |
| [67] |
Symington LS, Gautier J ( 2011). Double-strand break end resection and repair pathway choice. Annu Rev Genet 45, 247-271.
DOI URL PMID |
| [1] |
李红, 谢卡斌 ( 2017). 植物CRISPR基因组编辑技术的新进展. 生物工程学报 33, 1700-1711.
DOI URL |
| [2] | 冉毅东, 梁振, 张毅, 高彩霞 ( 2017). 植物基因组编辑试剂材料的导入及转化系统的研究现状及前景. 中国科学: 生命科学 47, 1159-1176. |
| [68] |
Ungerer J, Pakrasi HB ( 2016). Cpf1 is a versatile tool for CRISPR genome editing across diverse species of cyanobacteria. Sci Rep 6, 39681.
DOI URL PMID |
| [69] |
Waltz E ( 2016). Gene-edited CRISPR mushroom escapes US regulation. Nature 532, 293.
DOI URL PMID |
| [3] |
王影, 李相敢, 邱丽娟 ( 2018). CRISPR/Cas9基因组定点编辑中脱靶现象的研究进展. 植物学报 53, 528-541.
DOI URL |
| [4] |
Altpeter F, Springer NM, Bartley LE, Blechl AE, Brutnell TP, Citovsky V, Conrad LJ, Gelvin SB, Jackson DP, Kausch AP, Lemaux PG, Medford JI, Orozco-Cárdenas ML, Tricoli DM, Van Eck J, Voytas DF, Walbot V, Wang K, Zhang ZJ, Stewart Jr CN ( 2016). Advancing crop transformation in the era of genome editing. Plant Cell 28, 1510-1520.
DOI URL PMID |
| [70] |
Wang M, Lu Y, Botella JR, Mao Y, Hua K, Zhu JK ( 2017). Gene targeting by homology-directed repair in rice using a geminivirus-based CRISPR/Cas9 system. Mol Plant 10, 1007-1010.
DOI URL PMID |
| [71] |
Woo JW, Kim J, Kwon SI, Corvalán C, Cho SW, Kim H, Kim SG, Kim ST, Choe S, Kim JS ( 2015). DNA-free genome editing in plants with preassembled CRISPRCas9 ribonucleoproteins. Nat Biotechnol 33, 1162-1164.
DOI URL PMID |
| [5] |
Aman R, Ali Z, Butt H, Mahas A, Aljedaani F, Khan MZ, Ding S, Mahfouz M ( 2018). RNA virus interference via CRISPR/Cas13a system in plants. Genome Biol 19, 1.
DOI URL |
| [6] | Baazim H ( 2014). RNA-guided transcriptional regulation in plants via dCas9 chimeric proteins. Ph.D. thesis. Thuwal, Kingdom of Saudi Arabia: King Abdullah University of Science and Technology. pp. 32-48. |
| [72] |
Xie K, Minkenberg B, Yang Y ( 2015). Boosting CRISPR/ Cas9 multiplex editing capability with the endogenous tRNA-processing system. Proc Natl Acad Sci USA 112, 3570-3575.
DOI URL PMID |
| [73] |
Xing HL, Dong L, Wang ZP, Zhang HY, Han CY, Liu B, Wang XC, Chen QJ ( 2014). A CRISPR/Cas9 toolkit for multiplex genome editing in plants. BMC Plant Biol 14, 327.
DOI URL PMID |
| [7] |
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S, Romero DA, Horvath P ( 2007). Crispr provides acquired resistance against viruses in prokaryotes. Science 315, 1709-1712.
DOI URL PMID |
| [8] |
Brouns SJ, Jore MM, Lundgren M, Westra ER, Slijkhuis RJ, Snijders AP, Dickman MJ, Makarova KS, Koonin EV, van der Oost J ( 2008). Small CRISPR RNAs guide antiviral defense in prokaryotes. Science 321, 960-964.
DOI URL |
| [74] |
Xing YY, Yang J, Ren J ( 2016). Application of CRISPR/ Cas9 mediated genome editing in farm animals. Hereditas (Beijing) 38, 217-226.
DOI URL PMID |
| [75] | Xu R, Li H, Qin R, Wang L, Li L, Wei P, Yang J ( 2014). Gene targeting using the Agrobacterium tumefaciens- |
| [9] | Chamberlain JR, Schwarze U, Wang PR, Hirata RK, Hankenson KD, Pace JM, Underwood RA, Song KM, Sussman M, Byers PH, Russell DW ( 2004). Gene targeting in stem cells from individuals with Osteogenesis imperfecta. Science 303, 1198-1201. |
| [10] |
Cheng AW, Jillette N, Lee P, Plaskon D, Fujiwara Y, Wang W, Taghbalout A, Wang H ( 2016). Casilio: a versatile CRISPR-Cas9-Pumilio hybrid for gene regulation and genomic labelling. Cell Res 26, 254-257.
DOI URL PMID |
| [76] | mediated CRISPR-Cas system in rice.Rice 7, 5. |
| [77] |
Xu R, Qin R, Li H, Li D, Li L, Wei P, Yang J ( 2017). Generation of targeted mutant rice using a CRISPR-Cpf1 system. Plant Biotechnol J 15, 713-717.
DOI URL PMID |
| [78] |
Yan F, Kuang YJ, Ren B, Wang JW, Zhang DW, Lin HH, Yang B, Zhou XP, Zhou HB ( 2018). High-efficient A·T to G·C base editing by Cas9n-guided tRNA adenosine deaminase in rice. Mol Plant 11, 631-634.
DOI URL |
| [11] |
Cong L, Ran FA, Cox D, Lin S, Barretto R, Habib N, Hsu PD, Wu X, Jiang W, Marraffini LA, Zhang F ( 2013). Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819-823.
DOI URL PMID |
| [12] |
Corrigan-Curay J, O’Reilly M, Kohn DB, Cannon PM, Bao G, Bushman FD, Carroll D, Cathomen T, Joung JK, Roth D, Sadelain M, Scharenberg AM, VON Kalle C, Zhang F, Jambou R, Rosenthal E, Hassani M, Singh A, Porteus MH ( 2015). Genome editing technologies: defining a path to clinic: genomic editing: establishing preclinical toxicology standards, Bethesda, Maryland 10 June 2014. Mol Ther 23, 796.
DOI URL PMID |
| [79] |
Yu Z, Chen Q, Chen W, Zhang X, Mei F, Zhang P, Zhao M, Wang X, Shi N, Jackson S, Hong Y ( 2018). Multigene editing via CRISPR/Cas9 guided by a single-sgRNA seed in Arabidopsis. J Integr Plant Biol 60, 376-381.
DOI URL PMID |
| [80] |
Zetsche B, Gootenberg JS, Abudayyeh OO, Slaymaker IM, Makarova KS, Essletzbichler P, Volz SE, Joung J, van der Oost J, Regev A, Koonin EV, Zhang F ( 2015). Cpf1 is a single RNA-guided Endonuclease of a Class 2 CRISPR- Cas system. Cell 163, 759-771.
DOI URL PMID |
| [13] |
Deltcheva E, Chylinski K, Sharma CM, Gonzales K, Chao Y, Pirzada ZA, Eckert MR, Vogel J, Charpentier E ( 2011). CRISPR RNA maturation by trans-encoded small RNA and host factor RNase III. Nature 471, 602-607.
DOI URL PMID |
| [14] | Deveau H, Barrangou R, Garneau JE, Labonté J, Fremaux C, Boyaval P, Romero DA, Horvath P, Moineau S ( 2008). Phage response to CRISPR-encoded resistance in Streptococcus thermophilus. J Bacteriol 190, 1390-1400. |
| [15] |
Doron S, Melamed S, Ofir G, Leavitt A, Lopatina A, Keren M, Amitai G, Sorek R ( 2018). Systematic discovery of antiphage defense systems in the microbial pangenome. Science 359, eaar4120.
DOI URL PMID |
| [16] |
Durai S, Mani M, Kandavelou K, Wu J, Porteus MH, Chandra-segaran S ( 2005). Zinc finger nucleases: custom-designed molecular scissors for genome engineering of plant and mammalian cells. Nucleic Acids Res 33, 5978-5990.
DOI URL PMID |
| [17] |
Endo M, Mikami M, Toki S ( 2015). Multigene knockout utilizing off-target mutations of the CRISPR/Cas9 system in rice. Plant Cell Physiol 56, 41.
DOI URL PMID |
| [18] |
Feng C, Su H, Bai H, Wang R, Liu Y, Guo X, Liu C, Zhang J, Yuan J, Birchler JA, Han F ( 2018). High efficiency genome editing using a dmc1 promoter-controlled CRISPR/Cas9 system in maize. Plant Biotechnol J 16, 1848-1857.
DOI URL PMID |
| [19] |
Fu Y, Sander JD, Reyon D, Cascio VM, Joung JK ( 2014). Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat Biotechnol 32, 279-284.
DOI URL PMID |
| [20] | Gaj T, Gersbach CA, Barbas III CF ( 2013). ZFN, TALEN, and CRISPR/Cas-based methods for genome enginee- |
| [21] | ring.Trends Biotechnol 31, 397-405. |
| [22] |
Gasiunas G, Barrangou R, Horvath P, Siksnys V ( 2012). Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc Natl Acad Sci USA 109, E2579-E2586.
DOI URL |
| [23] |
Gaudelli NM, Komor AC, Rees HA, Packer MS, Badran AH, Bryson DI, Liu DR ( 2017). Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464-471.
DOI URL PMID |
| [24] |
He Y, Zhu M, Wang L, Wu J, Wang Q, Wang R, Zhao Y ( 2018). Programmed self-elimination of the CRISPR/Cas9 construct greatly accelerates the isolation of edited and transgene-free rice plants. Mol Plant 11, 1210-1213.
DOI URL PMID |
| [25] |
Hendel A, Bak RO, Clark JT, Kennedy AB, Ryan DE, Roy S, Steinfeld I, Lunstad BD, Kaiser RJ, Wilkens AB, Bacchetta R, Tsalenko A, Dellinger D, Bruhn L, Porteus MH ( 2015). Chemically modified guide RNAs enhance CRISPR-Cas genome editing in human primary cells. Nat Biotechnol 33, 985-989.
DOI URL PMID |
| [26] | Horvath P, Romero DA, Coûté-Monvoisin AC, Richards M, Deveau H, Moineau S, Boyaval P, Fremaux C, Barrangou R ( 2008). Diversity, activity, and evolution of CRISPR loci in Streptococcus thermophilus. J Bacteriol 190, 1401. |
| [27] |
Hu JH, Miller SM, Geurts MH, Tang W, Chen L, Sun N, Zeina CM, Gao X, Rees HA, Lin Z, Liu DR ( 2018). Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57-63.
DOI URL PMID |
| [28] |
Hu X, Meng X, Liu Q, Li J, Wang K ( 2017 a). Increasing the efficiency of CRISPR-Cas9-VQR precise genome editing in rice. Plant Biotechnol J 16, 292-297.
DOI URL PMID |
| [29] |
Hu X, Wang C, Fu Y, Liu Q, Jiao X, Wang K ( 2016). Expanding the range of CRISPR/Cas9 genome editing in rice. Mol Plant 9, 943-945.
DOI URL PMID |
| [30] |
Hu X, Wang C, Liu Q, Fu Y, Wang K ( 2017 b). Targeted mutagenesis in rice using CRISPR-Cpf1 system. J Genet Genomics 44, 71-73.
DOI URL PMID |
| [31] |
Hur JK, Kim K, Been KW, Baek G, Ye S, Hur JW, Ryu SM, Lee YS, Kim JS ( 2016). Targeted mutagenesis in mice by electroporation of Cpf1 ribonucleo proteins. Nat Biotechnol 34, 807-808.
DOI URL PMID |
| [32] | Ishino Y, Shinagawa H, Makino K, Amemura M, Nakata A ( 1987). Nucleotide sequence of the iap gene, responsible for alkaline phosphatase isozyme conversion in Escherichia coli, and identification of the gene product. J Bacteriol 169, 5429-5433. |
| [33] |
Jansen R, van Embden JD, Gaastra W, Schouls LM ( 2002). Identification of a novel family of sequence repeats among prokaryotes. OMICS 6, 23-33.
DOI URL PMID |
| [34] |
Jiang F, Doudna JA ( 2017). CRISPR-Cas9 structures and mechanisms. Ann Rev Biophy 46, 505-529.
DOI URL PMID |
| [35] |
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E ( 2012). A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816-821.
DOI URL |
| [36] |
Jinek M, East A, Cheng A, Lin S, Ma E, Doudna J ( 2013). RNA-programmed genome editing in human cells. eLife 2, e00471.
DOI URL PMID |
| [37] |
Kai H, Tao X, Yuan F, Dong W, Zhu JK ( 2018). Precise A·T to G·C base editing in the rice genome. Mol Plant 11, 627-630.
DOI URL |
| [38] |
Kim Y, Cheong SA, Lee JG, Lee SW, Lee MS, Baek IJ, Sung YH ( 2016). Generation of knockout mice by Cpf1-mediated gene targeting. Nat Biotechnol 34, 808-810.
DOI URL PMID |
| [39] |
Kweon J, Jang AH, Kim DE, Yang JW, Yoon M, Rim SH, Kim JS, Kim Y ( 2017). Fusion guide RNAs for orthogonal gene manipulation with Cas9 and Cpf1. Nat Commun 8, 1723.
DOI URL PMID |
| [40] |
Le Blanc C, Zhang F, Mendez J, Lozano Y, Chatpar K, Irish VF, Jacob Y ( 2017). Increased efficiency of targeted mutagenesis by CRISPR/Cas9 in plants using heat stress. Plant J 93, 377-386.
DOI URL PMID |
| [41] |
Li C, Zong Y, Wang YP, Jin S, Zhang DB, Song QN, Zhang R, Gao CX ( 2018). Expanded base editing in rice and wheat using a Cas9-adenosine deaminase fusion. Genome Biol 19, 59.
DOI URL |
| [42] |
Li J, Meng X, Zong Y, Chen K, Zhang H, Liu J, Li J, Gao C ( 2016). Gene replacements and insertions in rice by intron targeting using CRISPR-Cas9. Nat Plants 2, 16139.
DOI URL PMID |
| [43] | Li JF, Norville JE, Aach J, McCormack M, Zhang D, Bush J, Church GM, Sheen J ( 2013). Multiplex and homologous recombination-mediated genome editing in Arabidopsis and Nicotiana benthamiana using guide RNA and Cas9. Nat Biotechnol 31, 688-691. |
| [44] |
Li T, Liu B, Spalding MH, Weeks DP, Yang B ( 2012). High-efficiency talen-based gene editing produces disease-resistant rice. Nat Biotechnol 30, 390-392.
DOI URL PMID |
| [45] |
Liang PP, Xu YW, Zhang XY, Ding CH, Huang R, Zhang Z, Lv J, Xie XW, Chen YX, Li YJ, Sun Y, Bai YF, Zhou SY, Ma WB, Zhou CQ, Huang JJ ( 2015). CRISPR/Cas9- mediated gene editing in human tripronuclear zygotes. Prot Cell 6, 363-372.
DOI URL PMID |
| [1] | Qiang Zhang, Zhenyu Zhao, Pinghua Li. Research Progress of Gene Editing Technology in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 978-998. |
| [2] | Ziyang Wang, Shengxue Liu, Zhirui Yang, Feng Qin. Genetic Dissection of Drought Resistance in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 883-902. |
| [3] | Danling Hu, Yongwei Sun. Advances in Virus-mediated Genome Editing Technology in Plants [J]. Chinese Bulletin of Botany, 2024, 59(3): 452-462. |
| [4] | Chunyan Miao, Mingming Li, Xin Zuo, Ning Ding, Jiafang Du, Juan Li, Zhongyi Zhang, Fengqing Wang. Establishment of CRISPR/Cas9 Gene Editing System in Rehmannia henryi [J]. Chinese Bulletin of Botany, 2023, 58(6): 905-916. |
| [5] | Yanjun Guo, Feng Chen, Jingwen Luo, Wei Zeng, Wenliang Xu. The Biosynthesis of Plant Cell Wall Xylan and Its Application [J]. Chinese Bulletin of Botany, 2023, 58(2): 316-334. |
| [6] | Na Wang, Teng Jiang, Binxi Wang, Lifang Niu, Hao Lin. Advances in Haploid Breeding Technology and Its Application in Alfalfa and Other Legume Forages [J]. Chinese Bulletin of Botany, 2022, 57(6): 756-763. |
| [7] | He Xiaoling, Liu Pengcheng, Ma Bojun, Chen Xifeng. Advance in Gene-editing Technology Based on CRISPR/Cas9 and Its Application in Plants [J]. Chinese Bulletin of Botany, 2022, 57(4): 508-531. |
| [8] | Lubin Tan, Chuanqing Sun. Rapid Domestication of Wild Allotetraploid Rice: Starting a New Era of Human Agricultural Civilization [J]. Chinese Bulletin of Botany, 2021, 56(2): 134-137. |
| [9] | Xianrong Xie, Dongchang Zeng, Jiantao Tan, Qinlong Zhu, Yaoguang Liu. CRISPR-based DNA Fragment Deletion in Plants [J]. Chinese Bulletin of Botany, 2021, 56(1): 44-49. |
| [10] | Huijin Fan, Kangming Jin, Renying Zhuo, Guirong Qiao. Cloning and Expression Analysis of Different Truncated U3 Promoters in Phyllostachys edulis [J]. Chinese Bulletin of Botany, 2020, 55(3): 299-307. |
| [11] | Lu Zhang,Xinhua He. Nitrogen Utilization Mechanism in C3 and C4 Plants [J]. Chinese Bulletin of Botany, 2020, 55(2): 228-239. |
| [12] | Wang Ying, Li Xianggan, Qiu Lijuan. Research Progress in Off-target in CRISPR/Cas9 Genome Editing [J]. Chinese Bulletin of Botany, 2018, 53(4): 528-541. |
| [13] | LI Xin-Lei CHEN Fa-Di. Advances of Genetic Improvement and Germplasm Resources for Chrysanthemum [J]. Chinese Bulletin of Botany, 2004, 21(04): 392-401. |
| [14] | LIANG Ji CHEN Xiao-YangLIN Shan-Zhi XIE Xiang-Ming. Advance of Studies on Agrobacterium rhizogenesRi Plasmid rol Genes and Their Applications for Forest Tree Genetic Improvement [J]. Chinese Bulletin of Botany, 2002, 19(06): 650-658. |
| [15] | TENG Nian-Jun CHEN Fa-Di. Advances of Genetic Improvement for Ornamental Plants through Genetic Engineering [J]. Chinese Bulletin of Botany, 2002, 19(05): 538-545. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||