

Chinese Bulletin of Botany ›› 2024, Vol. 59 ›› Issue (6): 950-962.DOI: 10.11983/CBB24075 cstr: 32102.14.CBB24075
Special Issue: 大食物观; 粮食安全; 玉米生物学与分子设计(2024年59卷6期)
• INVITED REVIEWS • Previous Articles Next Articles
Received:2024-05-13
Accepted:2024-07-23
Online:2024-11-10
Published:2024-08-08
Contact:
*E-mail: liwenxue@caas.cn
Qingguo Du, Wenxue Li. Research Progress in the Regulation of Development and Stress Responses by Long Non-coding RNAs in Maize[J]. Chinese Bulletin of Botany, 2024, 59(6): 950-962.

Figure 1 The classification of long non-coding RNA (lncRNA) Blue boxes indicate the exon of mRNA, red lines represent lncRNA, black arrows indicate the direction of transcription.
| 数据库 | 物种及主要功能 | 网址 | 参考文献 |
|---|---|---|---|
| PlantNATsDB | 玉米、水稻、小麦和拟南芥等69个物种; NAT数据库, NAT调控网络 | Chen et al., | |
| PLncDB V2.0 | 玉米、水稻、小麦和拟南芥等80个物种; lncRNA基因组位置、序列、结构、表达模式、表观遗传及调控网络 | Jin et al., | |
| PNRD | 玉米、水稻和拟南芥等150个物种; lncRNA和小分子非编码RNA (sRNA)序列、结构、表达模式和功能 | Yi et al., | |
| LncPheDB | 玉米、小麦、水稻和高粱等9个物种; lncRNA序列、农艺性状表型和单核苷酸多态性 | Lou et al., | |
| GreeNC 2.0 | 玉米、水稻和拟南芥等96个物种; lncRNA序列及种间和种内lncRNA序列聚类分析 | Di Marsico et al., | |
| wLNCdb | 小麦; lncRNA序列、表达模式、共表达基因和单核苷酸多态性 | Zhang et al., | |
| RiceLncPedia | 水稻; lncRNA表达谱、基因组变异、lncRNA- miRNA相互作用、转座因子相关lncRNA以及lncRNA与农艺性状的关联 | Zhang et al., | |
| scPLAD | 拟南芥; 单细胞水平lncRNA图谱浏览器 | He et al., |
Table 1 Summary of database depositing plant long non-coding RNA (lncRNA)
| 数据库 | 物种及主要功能 | 网址 | 参考文献 |
|---|---|---|---|
| PlantNATsDB | 玉米、水稻、小麦和拟南芥等69个物种; NAT数据库, NAT调控网络 | Chen et al., | |
| PLncDB V2.0 | 玉米、水稻、小麦和拟南芥等80个物种; lncRNA基因组位置、序列、结构、表达模式、表观遗传及调控网络 | Jin et al., | |
| PNRD | 玉米、水稻和拟南芥等150个物种; lncRNA和小分子非编码RNA (sRNA)序列、结构、表达模式和功能 | Yi et al., | |
| LncPheDB | 玉米、小麦、水稻和高粱等9个物种; lncRNA序列、农艺性状表型和单核苷酸多态性 | Lou et al., | |
| GreeNC 2.0 | 玉米、水稻和拟南芥等96个物种; lncRNA序列及种间和种内lncRNA序列聚类分析 | Di Marsico et al., | |
| wLNCdb | 小麦; lncRNA序列、表达模式、共表达基因和单核苷酸多态性 | Zhang et al., | |
| RiceLncPedia | 水稻; lncRNA表达谱、基因组变异、lncRNA- miRNA相互作用、转座因子相关lncRNA以及lncRNA与农艺性状的关联 | Zhang et al., | |
| scPLAD | 拟南芥; 单细胞水平lncRNA图谱浏览器 | He et al., |
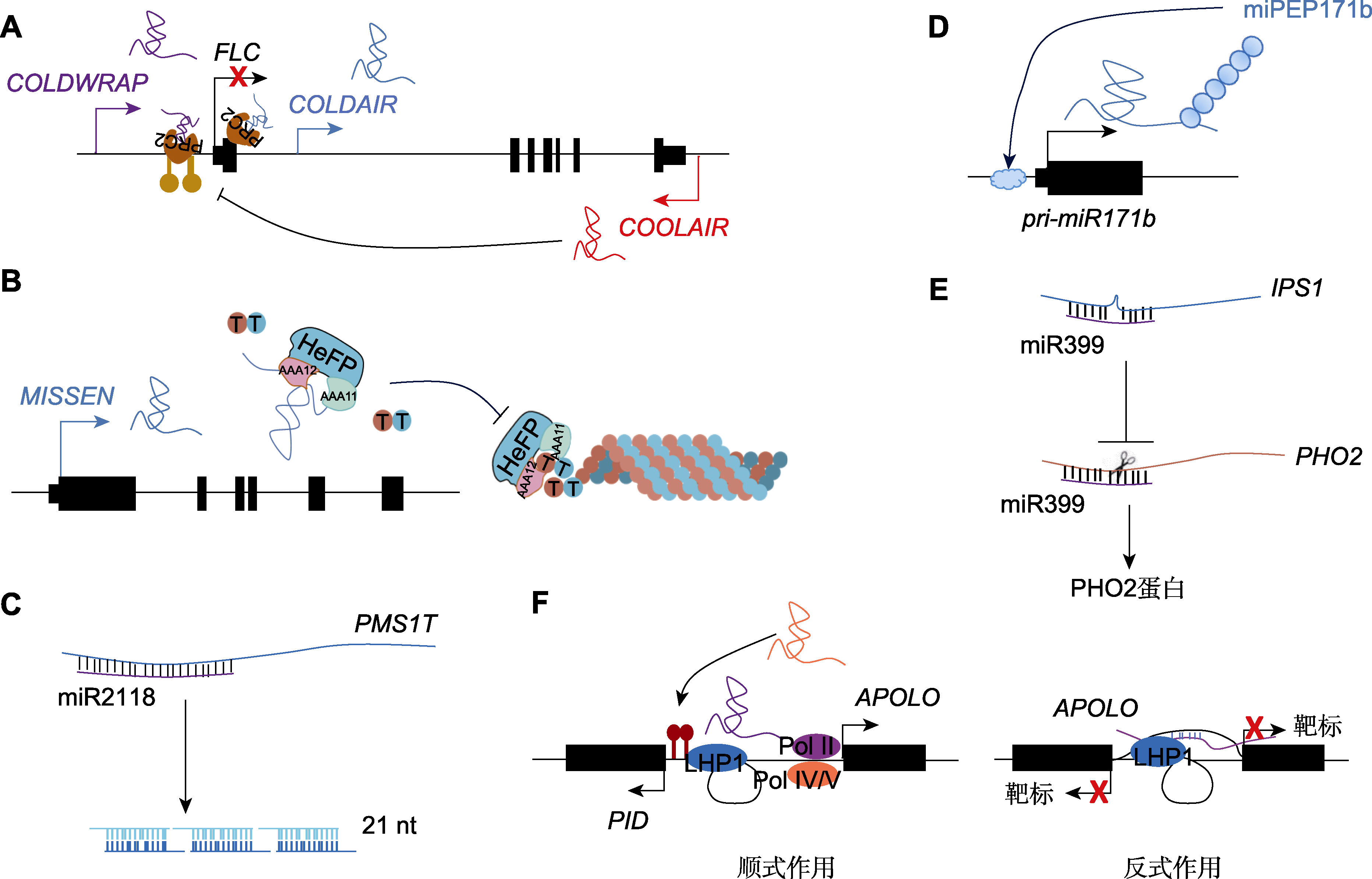
Figure 2 Representative models for the roles of plant long non-coding RNA (lncRNA) (A) lncRNAs mediate the coordinated switching of chromatin states; (B) lncRNAs act as a bait by binding to transcriptional regulators; (C) lncRNAs act as precursors of sRNA; (D) lncRNAs can translated into small peptides to regulate target genes in cis or in trans; (E) lncRNAs act as a target mimic for miRNA; (F) lncRNAs regulate the formation of chromatin loops
| lncRNA ID | lncRNA分子调控机制 | 功能 | 参考文献 |
|---|---|---|---|
| ZM401 | 调控花粉发育关键基因ZmADS2、MZm3-3和ZmC5的表达 | 花粉发育 | Ma et al., |
| GARR2 | 调控赤霉素和生长素通路中相关基因的表达 | 响应赤霉素, 调控株高 | Li et al., |
| cis-NATZmNAC48 | 负调控ZmNAC48的表达, 影响叶片气孔关闭 | 干旱 | Mao et al., |
| MSTRG8888.1 | 通过靶向bHLH转录因子, 影响下游耐盐相关基因的表达 | 盐胁迫 | Liu et al., |
| PILNCR1 | 作为竞争性内源RNA分子结合ZmmiRNA399, 并抑制ZmmiRNA399对靶基因ZmPHO2的切割 | 磷酸盐胁迫 | Du et al., |
| PILNCR2 | 与ZmPHT1s形成双链结构, 保护ZmPHT1s不被ZmmiR399切割 | 磷酸盐胁迫 | Wang et al., |
Table 2 Function of long non-coding RNA (lncRNA) in maize
| lncRNA ID | lncRNA分子调控机制 | 功能 | 参考文献 |
|---|---|---|---|
| ZM401 | 调控花粉发育关键基因ZmADS2、MZm3-3和ZmC5的表达 | 花粉发育 | Ma et al., |
| GARR2 | 调控赤霉素和生长素通路中相关基因的表达 | 响应赤霉素, 调控株高 | Li et al., |
| cis-NATZmNAC48 | 负调控ZmNAC48的表达, 影响叶片气孔关闭 | 干旱 | Mao et al., |
| MSTRG8888.1 | 通过靶向bHLH转录因子, 影响下游耐盐相关基因的表达 | 盐胁迫 | Liu et al., |
| PILNCR1 | 作为竞争性内源RNA分子结合ZmmiRNA399, 并抑制ZmmiRNA399对靶基因ZmPHO2的切割 | 磷酸盐胁迫 | Du et al., |
| PILNCR2 | 与ZmPHT1s形成双链结构, 保护ZmPHT1s不被ZmmiR399切割 | 磷酸盐胁迫 | Wang et al., |
| [1] |
Ariel F, Jegu T, Latrasse D, Romero-Barrios N, Christ A, Benhamed M, Crespi M (2014). Noncoding transcription by alternative RNA polymerases dynamically regulates an auxin-driven chromatin loop. Mol Cell 55, 383-396.
DOI PMID |
| [2] | Ariel F, Lucero L, Christ A, Mammarella MF, Jegu T, Veluchamy A, Mariappan K, Latrasse D, Blein T, Liu C, Benhamed M, Crespi M (2020). R-Loop mediated trans action of the APOLO long noncoding RNA. Mol Cell 77, 1055-1065. |
| [3] |
Ariel F, Romero-Barrios N, Jégu T, Benhamed M, Crespi M (2015). Battles and hijacks: noncoding transcription in plants. Trends Plant Sci 20, 362-371.
DOI PMID |
| [4] |
Aung K, Lin SI, Wu CC, Huang YT, Su CL, Chiou TJ (2006). pho2, a phosphate overaccumulator, is caused by a nonsense mutation in a microRNA399 target gene. Plant Physiol 141, 1000-1011.
DOI PMID |
| [5] |
Beló A, Beatty MK, Hondred D, Fengler KA, Li BL, Rafalski A (2010). Allelic genome structural variations in maize detected by array comparative genome hybridization. Theor Appl Genet 120, 355-367.
DOI PMID |
| [6] | Boerner S, McGinnis KM (2012). Computational identification and functional predictions of long noncoding RNA in Zea mays. PLoS One 7, e43047. |
| [7] |
Brannan CI, Dees EC, Ingram RS, Tilghman SM (1990). The product of the H19gene may function as an RNA. Mol Cell Biol 10, 28-36.
DOI PMID |
| [8] | Chen DJ, Yuan CH, Zhang J, Zhang Z, Bai L, Meng YJ, Chen LL, Chen M (2012). PlantNATsDB: a comprehensive database of plant natural antisense transcripts. Nucleic Acids Res 40, D1187-D1193. |
| [9] |
Chen Q, Zhang XD, Shi JC, Yan MH, Zhou T (2021). Origins and evolving functionalities of tRNA-derived small RNAs. Trends Biochem Sci 46, 790-804.
DOI PMID |
| [10] | Chen XM, Rechavi O (2022). Plant and animal small RNA communications between cells and organisms. Nat Rev Mol Cell Biol 23, 185-203. |
| [11] | Chiou TJ, Aung K, Lin SI, Wu CC, Chiang SF, Su CL (2006). Regulation of phosphate homeostasis by microRNA in Arabidopsis. Plant Cell 18, 412-421. |
| [12] |
Csorba T, Questa JI, Sun QW, Dean C (2014). Antisense COOLAIR mediates the coordinated switching of chromatin states at FLC during vernalization. Proc Natl Acad Sci USA 111, 16160-16165.
DOI PMID |
| [13] | Di Marsico M, Paytuvi-Gallart A, Sanseverino W, Aiese Cigliano R (2022). GreeNC 2.0: a comprehensive database of plant long non-coding RNAs. Nucleic Acids Res 50, D1442-D1447. |
| [14] |
Ding JH, Lu Q, Ouyang YD, Mao HL, Zhang PB, Yao JL, Xu CG, Li XH, Xiao JH, Zhang QF (2012). A long noncoding RNA regulates photoperiod-sensitive male sterility, an essential component of hybrid rice. Proc Natl Acad Sci USA 109, 2654-2659.
DOI PMID |
| [15] | Djebali S, Davis CA, Merkel A, Dobin A, Lassmann T, Mortazavi A, Tanzer A, Lagarde J, Lin W, Schlesinger F, Xue CH, Marinov GK, Khatun J, Williams BA, Zaleski C, Rozowsky J, Röder M, Kokocinski F, Abdelhamid RF, Alioto T, Antoshechkin I, Baer MT, Bar NS, Batut P, Bell K, Bell I, Chakrabortty S, Chen X, Chrast J, Curado J, Derrien T, Drenkow J, Dumais E, Dumais J, Duttagupta R, Falconnet E, Fastuca M, Fejes-Toth K, Ferreira P, Foissac S, Fullwood MJ, Gao H, Gonzalez D, Gordon A, Gunawardena H, Howald C, Jha S, Johnson R, Kapranov P, King B, Kingswood C, Luo OJ, Park E, Persaud K, Preall JB, Ribeca P, Risk B, Robyr D, Sammeth M, Schaffer L, See LH, Shahab A, Skancke J, Suzuki AM, Takahashi H, Tilgner H, Trout D, Walters N, Wang HE, Wrobel J, Yu YB, Ruan XA, Hayashizaki Y, Harrow J, Gerstein M, Hubbard T, Reymond A, Antonarakis SE, Hannon G, Giddings MC, Ruan YJ, Wold B, Carninci P, Guigó R, Gingeras TR (2012). Landscape of transcription in human cells. Nature 489, 101-108. |
| [16] |
Downes BP, Stupar RM, Gingerich DJ, Vierstra RD (2003). The HECT ubiquitin-protein ligase (UPL) family in Arabidopsis: UPL3 has a specific role in trichome development. Plant J 35, 729-742.
PMID |
| [17] | Du QG, Wang K, Zou C, Xu C, Li WX (2018). The PILNCR1-miR399 regulatory module is important for low phosphate tolerance in maize. Plant Physiol 177, 1743-1753. |
| [18] | Du Toit A (2013). Non-coding RNA: RNA stability control by Pol II. Nat Rev Mol Cell Biol 14, 128-129. |
| [19] | Fan YR, Yang JY, Mathioni SM, Yu JS, Shen JQ, Yang XF, Wang L, Zhang QH, Cai ZX, Xu CG, Li XH, Xiao JH, Meyers BC, Zhang QF (2016). PMS1T, producing phased small-interfering RNAs, regulates photoperiod- sensitive male sterility in rice. Proc Natl Acad Sci USA 113, 15144-15149. |
| [20] |
Fang J, Zhang FT, Wang HR, Wang W, Zhao F, Li ZJ, Sun CH, Chen FM, Xu F, Chang SQ, Wu L, Bu QY, Wang PR, Xie JK, Chen F, Huang XH, Zhang YJ, Zhu XG, Han B, Deng XJ, Chu CC (2019). Ef-cd locus shortens rice maturity duration without yield penalty. Proc Natl Acad Sci USA 116, 18717-18722.
DOI PMID |
| [21] |
Fang XF, Wu Z, Raitskin O, Webb K, Voigt P, Lu TC, Howard M, Dean C (2020). The 3' processing of antisense RNAs physically links to chromatin-based transcriptional control. Proc Natl Acad Sci USA 117, 15316-15321.
DOI PMID |
| [22] |
Franco-Zorrilla JM, Valli A, Todesco M, Mateos I, Puga MI, Rubio-Somoza I, Leyva A, Weigel D, García JA, Paz-Ares J (2007). Target mimicry provides a new mechanism for regulation of microRNA activity. Nat Genet 39, 1033-1037.
DOI PMID |
| [23] | Fujii H, Chiou TJ, Lin SI, Aung K, Zhu JK (2005). A miRNA involved in phosphate-starvation response in Arabidopsis. Curr Biol 15, 2038-2043. |
| [24] | Guo GH, Liu XY, Sun FL, Cao J, Huo N, Wuda B, Xin MM, Hu ZR, Du JK, Xia R, Rossi V, Peng HR, Ni ZF, Sun QX, Yao YY (2018). Wheat miR9678 affects seed germination by generating phased siRNAs and modulating abscisic acid/gibberellin signaling. Plant Cell 30, 796-814. |
| [25] |
Han LQ, Mu ZN, Luo Z, Pan QC, Li L (2019). New lncRNA annotation reveals extensive functional divergence of the transcriptome in maize. J Integr Plant Biol 61, 394-405.
DOI |
| [26] | He ZH, Lan YM, Zhou XK, Yu BJ, Zhu T, Yang F, Fu LY, Chao HY, Wang JH, Feng RX, Zuo SM, Lan WZ, Chen CL, Chen M, Zhao X, Hu KM, Chen DJ (2024). Single- cell transcriptome analysis dissects lncRNA-associated gene networks in Arabidopsis. Plant Commun 5, 100717. |
| [27] |
Heo JB, Sung S (2011). Vernalization-mediated epigenetic silencing by a long intronic noncoding RNA. Science 331, 76-79.
DOI PMID |
| [28] |
Hermans-Beijnsberger S, van Bilsen M, Schroen B (2018). Long non-coding RNAs in the failing heart and vasculature. Noncoding RNA Res 3, 118-130.
DOI PMID |
| [29] | Hong YH, Zhang YY, Cui J, Meng J, Chen YH, Zhang CW, Yang JX, Luan YS (2022). The lncRNA39896-miR166b- HDZs module affects tomato resistance to Phytophthora infestans. J Integr Plant Biol 64, 1979-1993. |
| [30] | Hu B, Zhu CG, Li F, Tang JY, Wang YQ, Lin AH, Liu LC, Che RH, Chu CC (2011). LEAF TIP NECROSIS1 plays a pivotal role in the regulation of multiple phosphate starvation responses in rice. Plant Physiol 156, 1101-1115. |
| [31] |
Hu XL, Wei QY, Wu HY, Huang YX, Peng XJ, Han GM, Ma Q, Zhao Y (2022). Identification and characterization of heat-responsive lncRNAs in maize inbred line CM1. BMC Genomics 23, 208.
DOI PMID |
| [32] | Huang DQ, Feurtado JA, Smith MA, Flatman LK, Koh C, Cutler AJ (2017). Long noncoding miRNA gene represses wheat β-diketone waxes. Proc Natl Acad Sci USA 114, E3149-E3158. |
| [33] | Huang TK, Han CL, Lin SI, Chen YJ, Tsai YC, Chen YR, Chen JW, Lin WY, Chen PM, Liu TY, Chen YS, Sun CM, Chiou TJ (2013). Identification of downstream components of ubiquitin-conjugating enzyme PHOSPHATE2 by quantitative membrane proteomics in Arabidopsis roots. Plant Cell 25, 4044-4060. |
| [34] | Jabnoune M, Secco D, Lecampion C, Robaglia C, Shu QY, Poirier Y (2013). A rice cis-natural antisense RNA acts as a translational enhancer for its cognate mRNA and contributes to phosphate homeostasis and plant fitness. Plant Cell 25, 4166-4182. |
| [35] |
Jin JJ, Lu P, Xu YL, Li ZF, Yu SZ, Liu J, Wang H, Chua NH, Cao PJ (2021). PLncDB V2.0: a comprehensive encyclopedia of plant long noncoding RNAs. Nucleic Acids Res 49, D1489-D1495.
DOI PMID |
| [36] | Kim DH, Sung S (2017). Vernalization-triggered intragenic chromatin loop formation by long noncoding RNAs. Dev Cell 40, 302-312. |
| [37] | Lai JS, Li RQ, Xu X, Jin WW, Xu ML, Zhao HN, Xiang ZK, Song WB, Ying K, Zhang M, Jiao YP, Ni PX, Zhang JG, Li D, Guo XS, Ye KX, Jian M, Wang B, Zheng HS, Liang HQ, Zhang XQ, Wang SC, Chen SJ, Li JS, Fu Y, Springer NM, Yang HM, Wang J, Dai JR, Schnable PS, Wang J (2010). Genome-wide patterns of genetic variation among elite maize inbred lines. Nat Genet 42, 1027-1030. |
| [38] | Lauressergues D, Couzigou JM, Clemente HS, Martinez Y, Dunand C, Bécard G, Combier JP (2015). Primary transcripts of microRNAs encode regulatory peptides. Nature 520, 90-93. |
| [39] | Li L, Eichten SR, Shimizu R, Petsch K, Yeh CT, Wu W, Chettoor AM, Givan SA, Cole RA, Fowler JE, Evans MMS, Scanlon MJ, Yu JM, Schnable PS, Timmermans MCP, Springer NM, Muehlbauer GJ (2014). Genome- wide discovery and characterization of maize long non- coding RNAs. Genome Biol 15, R40. |
| [40] | Li W, Chen YD, Wang YL, Zhao J, Wang YJ (2022a). Gypsy retrotransposon-derived maize lncRNA GARR2 modulates gibberellin response. Plant J 110, 1433-1446. |
| [41] | Li XH, Chen WW, Lu SQ, Fang JT, Zhu H, Zhang XB, Qi YW (2022b). Full-length transcriptome analysis of maize root tips reveals the molecular mechanism of cold stress during the seedling stage. BMC Plant Biol 22, 398. |
| [42] | Lin SI, Chiang SF, Lin WY, Chen JW, Tseng CY, Wu PC, Chiou TJ (2008). Regulatory network of microRNA399 and PHO2 by systemic signaling. Plant Physiol 147, 732-746. |
| [43] |
Liu P, Zhang YC, Zou CY, Yang C, Pan GT, Ma LL, Shen YO (2022). Integrated analysis of long non-coding RNAs and mRNAs reveals the regulatory network of maize seedling root responding to salt stress. BMC Genomics 23, 50.
DOI PMID |
| [44] | Lou DJ, Li F, Ge JY, Fan WY, Liu ZR, Wang YY, Huang JF, Xing M, Guo WL, Wang SZ, Qiao WH, Han ZY, Qian Q, Yang QW, Zheng XM (2022). LncPheDB: a genome- wide lncRNAs regulated phenotypes database in plants. aBIOTECH 3, 169-177. |
| [45] | Lu J, Zhen S, Zhang J, Xie Y, He C, Wang X, Wang Z, Zhang S, Li Y, Cui Y, Wang G, Wang J, Liu J, Li L, Gu R, Zheng X, Fu J (2023). Combined population transcriptomic and genomic analysis reveals cis-regulatory differentiation of non-coding RNAs in maize. Theor Appl Genet 136, 16. |
| [46] | Lv YD, Liang ZK, Ge M, Qi WC, Zhang TF, Lin F, Peng ZH, Zhao H (2016). Genome-wide identification and functional prediction of nitrogen-responsive intergenic and intronic long non-coding RNAs in maize (Zea mays L.). BMC Genomics 17, 350. |
| [47] |
Ma JX, Yan BX, Qu YY, Qin FF, Yang YT, Hao XJ, Yu JJ, Zhao Q, Zhu DY, Ao GM (2008). Zm401, a short-open reading-frame mRNA or noncoding RNA, is essential for tapetum and microspore development and can regulate the floret formation in maize. J Cell Biochem 105, 136-146.
DOI PMID |
| [48] | Mao Y, Xu J, Wang Q, Li GB, Tang X, Liu TH, Feng XJ, Wu FK, Li ML, Xie WB, Lu YL (2021). A natural antisense transcript acts as a negative regulator for the maize drought stress response gene ZmNAC48. J Exp Bot 72, 2790-2806. |
| [49] |
Melé M, Mattioli K, Mallard W, Shechner DM, Gerhardinger C, Rinn JL (2017). Chromatin environment, transcriptional regulation, and splicing distinguish lincRNAs and mRNAs. Genome Res 27, 27-37.
DOI PMID |
| [50] | Moison M, Pacheco JM, Lucero L, Fonouni-Farde C, Rodríguez-Melo J, Mansilla N, Christ A, Bazin J, Benhamed M, Ibañez F, Crespi M, Estevez JM, Ariel F (2021). The lncRNA APOLO interacts with the transcription factor WRKY42 to trigger root hair cell expansion in response to cold. Mol Plant 14, 937-948. |
| [51] | Ng SY, Lin L, Soh BS, Stanton LW (2013). Long noncoding RNAs in development and disease of the central nervous system. Trends Genet 29, 461-468. |
| [52] |
Palazzo AF, Koonin EV (2020). Functional long non-coding RNAs evolve from junk transcripts. Cell 183, 1151-1161.
DOI PMID |
| [53] |
Ponting CP, Oliver PL, Reik W (2009). Evolution and functions of long noncoding RNAs. Cell 136, 629-641.
DOI PMID |
| [54] |
Qin T, Zhao HY, Cui P, Albesher N, Xiong LM (2017). A nucleus-localized long non-coding RNA enhances drought and salt stress tolerance. Plant Physiol 175, 1321-1336.
DOI PMID |
| [55] |
Ratcliffe OJ, Kumimoto RW, Wong BJ, Riechmann JL (2003). Analysis of the Arabidopsis MADS AFFECTING FLOWERING gene family: MAF2 prevents vernalization by short periods of cold. Plant Cell 15, 1159-1169.
DOI PMID |
| [56] | Röhrig H, Schmidt J, Miklashevichs E, Schell J, John M (2002). Soybean ENOD40 encodes two peptides that bind to sucrose synthase. Proc Natl Acad Sci USA 99, 1915-1920. |
| [57] | Sharma A, Badola PK, Bhatia C, Sharma D, Trivedi PK (2020). Primary transcript of miR858 encodes regulatory peptide and controls flavonoid biosynthesis and development in Arabidopsis. Nat Plants 6, 1262-1274. |
| [58] |
Shi XF, Sun M, Liu HB, Yao YW, Song Y (2013). Long non-coding RNAs: a new frontier in the study of human diseases. Cancer Lett 339, 159-166.
DOI PMID |
| [59] | Song YP, Xuan AR, Bu CH, Ci D, Tian M, Zhang DQ (2019). Osmotic stress-responsive promoter upstream transcripts (PROMPTs) act as carriers of MYB transcription factors to induce the expression of target genes in Populus simonii. Plant Biotechnol J 17, 164-177. |
| [60] | Statello L, Guo CJ, Chen LL, Huarte M (2020). Gene regulation by long non-coding RNAs and its biological functions. Nat Rev Mol Cell Biol 22, 96-118. |
| [61] |
Sun Q, Liu XG, Yang J, Liu WW, Du QG, Wang HQ, Fu CX, Li WX (2018). MicroRNA528 affects lodging resistance of maize by regulating lignin biosynthesis under nitrogen-luxury conditions. Mol Plant 11, 806-814.
DOI PMID |
| [62] | Tadege M, Sheldon CC, Helliwell CA, Upadhyaya NM, Dennis ES, Peacock WJ (2003). Reciprocal control of flowering time by OsSOC1 in transgenic Arabidopsis and by FLC in transgenic rice. Plant Biotechnol J 1, 361-369. |
| [63] | Tang X, Li QM, Feng XJ, Yang B, Zhong X, Zhou Y, Wang Q, Mao Y, Xie WB, Liu TH, Tang Q, Guo W, Wu FK, Feng XJ, Wang QJ, Lu YL, Xu J (2023). Identification and functional analysis of drought-responsive long noncoding RNAs in maize roots. Int J Mol Sci 24, 15039. |
| [64] | Tian YK, Zheng H, Zhang F, Wang SL, Ji XR, Xu C, He YH, Ding Y (2019). PRC2 recruitment and H3K27me3 deposition at FLC require FCA binding of COOLAIR. Sci Adv 5, eaau7246. |
| [65] | Todesco M, Rubio-Somoza I, Paz-Ares J, Weigel D (2010). A collection of target mimics for comprehensive analysis of microRNA function in Arabidopsis thaliana. PLoS Genet 6, e1001031. |
| [66] | Wang H, Niu QW, Wu HW, Liu J, Ye J, Yu N, Chua NH (2015). Analysis of non-coding transcriptome in rice and maize uncovers roles of conserved lncRNAs associated with agriculture traits. Plant J 84, 404-416. |
| [67] |
Wang KC, Chang HY (2011). Molecular mechanisms of long noncoding RNAs. Mol Cell 43, 904-914.
DOI PMID |
| [68] | Wang TZ, Zhao MG, Zhang XX, Liu M, Yang CG, Chen YH, Chen RJ, Wen JQ, Mysore KS, Zhang WH (2017). Novel phosphate deficiency-responsive long non-coding RNAs in the legume model plant Medicago truncatula. J Exp Bot 68, 5937-5948. |
| [69] | Wang Y, Luo XJ, Sun F, Hu JH, Zha X, Su W, Yang JS (2018). Overexpressing lncRNA LAIR increases grain yield and regulates neighbouring gene cluster expression in rice. Nat Commun 9, 3516. |
| [70] | Wang YF, Wang ZH, Du QG, Wang K, Zou CQ, Li WX (2023). The long non-coding RNA PILNCR2 increases low phosphate tolerance in maize by interfering with miRNA399-guided cleavage of ZmPHT1s. Mol Plant 16, 1146-1159. |
| [71] | Wang YQ, Fan XD, Lin F, He GM, Terzaghi W, Zhu DM, Deng XW (2014). Arabidopsis noncoding RNA mediates control of photomorphogenesis by red light. Proc Natl Acad Sci USA 111, 10359-10364. |
| [72] |
Wierzbicki AT, Haag JR, Pikaard CS (2008). Noncoding transcription by RNA polymerase Pol IVb/Pol V mediates transcriptional silencing of overlapping and adjacent genes. Cell 135, 635-648.
DOI PMID |
| [73] | Xing LJ, Zhu M, Luan MD, Zhang M, Jin L, Liu YP, Zou JJ, Wang L, Xu MY (2022). miR169q and NUCLEAR FACTOR YA8 enhance salt tolerance by activating PEROXIDASE1 expression in response to ROS. Plant Physiol 188, 608-623. |
| [74] | Yang J, Wei HB, Hou M, Chen LH, Zou T, Ding H, Jing YF, Zhang XM, Zhao YP, Liu Q, Heng YQ, Wu H, Wang BB, Kong DX, Wang HY (2023). ZmSPL13 and ZmSPL29 act together to promote vegetative and reproductive transition in maize. New Phytol 239, 1505-1520. |
| [75] | Yi X, Zhang ZH, Ling Y, Xu WY, Su Z (2015). PNRD: a plant non-coding RNA database. Nucleic Acids Res 43, D982-D989. |
| [76] | Zhan JP, Meyers BC (2023). Plant small RNAs: their biogenesis, regulatory roles, and functions. Annu Rev Plant Biol 74, 21-51. |
| [77] | Zhang N, Liu ZG, Sun SC, Liu SY, Lin JH, Peng YF, Zhang XX, Yang H, Cen X, Wu J (2020). Response of AtR8 lncRNA to salt stress and its regulation on seed germination in Arabidopsis. Chin Bull Bot 55, 421-429. (in Chinese) |
|
张楠, 刘自广, 孙世臣, 刘圣怡, 林建辉, 彭疑芳, 张晓旭, 杨贺, 岑曦, 吴娟 (2020). 拟南芥AtR8 IncRNA对盐胁迫响应及其对种子萌发的调节作用. 植物学报 55, 421-429.
DOI |
|
| [78] | Zhang S, Wu CY (2019). Long noncoding RNA Ef-cd promotes maturity without yield penalty in rice. Chin Bull Bot 54, 550-553. (in Chinese) |
|
张硕, 吴昌银 (2019). 长链非编码RNA基因Ef-cd调控水稻早熟与稳产. 植物学报 54, 550-553.
DOI |
|
| [79] | Zhang ZF, Xu Y, Yang F, Xiao BZ, Li GL (2021). RiceLncPedia: a comprehensive database of rice long non- coding RNAs. Plant Biotechnol J 19, 1492-1494. |
| [80] | Zhang ZH, Zhang RJ, Meng FF, Chen YM, Wang WX, Yang K, Gao YJ, Xin MM, Du JK, Hu ZR, Ni ZF, Sun QX, Guo WL, Yao YY, Zhang ZH, Zhang RJ, Meng FF, Chen YM, Wang WX, Yang K, Gao YJ, Xin MM, Du JK, Hu ZR, Ni ZF, Sun QX, Guo WL, Yao YY (2023). A comprehensive atlas of long non-coding RNAs provides insight into grain development in wheat. Seed Biol 2, 12. |
| [81] |
Zhao SR, Zhang Y, Gordon W, Quan J, Xi HL, Du S, von Schack D, Zhang BH (2015). Comparison of stranded and non-stranded RNA-seq transcriptome profiling and investigation of gene overlap. BMC Genomics 16, 675.
DOI PMID |
| [82] | Zhao XY, Li JR, Lian B, Gu HQ, Li Y, Qi YJ (2018). Global identification of Arabidopsis lncRNAs reveals the regulation of MAF4 by a natural antisense RNA. Nat Commun 9, 5056. |
| [83] | Zhao YS, Zhu P, Hepworth J, Bloomer R, Antoniou- Kourounioti RL, Doughty J, Heckmann A, Xu CY, Yang HC, Dean C (2021). Natural temperature fluctuations promote COOLAIR regulation of FLC. Genes Dev 35, 888-898. |
| [84] |
Zhou YF, Zhang YC, Sun YM, Yu Y, Lei MQ, Yang YW, Lian JP, Feng YZ, Zhang Z, Yang L, He RR, Huang JH, Cheng Y, Liu YW, Chen YQ (2021). The parent-of-origin lncRNA MISSEN regulates rice endosperm development. Nat Commun 12, 6525.
DOI PMID |
| [1] | Yue Ming, Peiyao Hao, Lingqian Tan, Xi Zheng. A study on urban biodiversity conservation and enhancement in china based on the concept of green and high-quality development of cities [J]. Biodiv Sci, 2025, 33(5): 24524-. |
| [2] | Xiong Lianglin, Liang Guolu, Guo Qigao, Jing Danlong. Advances in the Regulation of Alternative Splicing of Genes in Plants in Response to Abiotic Stress [J]. Chinese Bulletin of Botany, 2025, 60(3): 435-448. |
| [3] | Tan Xiangping, Liang Xiaodong, Luo Shixiao, Wei Dan, Yang Min, Liu Guofeng, Qu Chao, Wang Hongfeng, Hu Yuhua, Jiang Jun, Zeng Youpai, Wang Jun, Yan Yuehong, Wang Ruijiang, Cao Honglin, Liao Jingping. Establishing a regional plant ex situ conservation system centered around the national botanical garden: A detailed exploration from Guangdong Province [J]. Biodiv Sci, 2025, 33(2): 24326-. |
| [4] | Qingyang Li, Cui Liu, Li He, Shan Peng, Jiayin Ma, Ziyi Hu, Hongbo Liu. Cloning and Functional Analysis of the BnaA02.CPSF6 Gene from Brassica napus [J]. Chinese Bulletin of Botany, 2025, 60(1): 62-73. |
| [5] | Yaping Wang, Wenquan Bao, Yu’e Bai. Advances in the Application of Single-cell Transcriptomics in Plant Growth, Development and Stress Response [J]. Chinese Bulletin of Botany, 2025, 60(1): 101-113. |
| [6] | Tao Wang, Jinglei Feng, Cui Zhang. Research Progress on Molecular Mechanisms of Heat Stress Affecting the Growth and Development of Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 963-977. |
| [7] | Suowei Wu, Xueli An, Xiangyuan Wan. Molecular Mechanisms of Male Sterility and their Applications in Biotechnology-based Male-sterility Hybrid Seed Production in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 932-949. |
| [8] | Mingmin Zheng, Qiang Huang, Peng Zhang, Xiaowei Liu, Zhuofan Zhao, Hongyang Yi, Tingzhao Rong, Moju Cao. Research Progress on Cytoplasmic Male Sterility and Fertility Restoration in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 999-1006. |
| [9] | Qiaoxia Li, Youlong Li, Jigang Li, Chenlong Chen, Kun Sun. Effects of photoperiods on the development of chasmogamous and cleistogamous flowers in Viola monbeigii and V. dissecta [J]. Biodiv Sci, 2024, 32(6): 23484-. |
| [10] | Yuan Li, Kaijian Fan, Tai An, Cong Li, Junxia Jiang, Hao Niu, Weiwei Zeng, Yanfang Heng, Hu Li, Junjie Fu, Huihui Li, Liang Li. Study on Multi-environment Genome-wide Prediction of Inbred Agronomic Traits in Maize Natural Populations [J]. Chinese Bulletin of Botany, 2024, 59(6): 1041-1053. |
| [11] | Ziyang Wang, Shengxue Liu, Zhirui Yang, Feng Qin. Genetic Dissection of Drought Resistance in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 883-902. |
| [12] | Qiang Zhang, Zhenyu Zhao, Pinghua Li. Research Progress of Gene Editing Technology in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 978-998. |
| [13] | Wenli Yang, Zhao Li, Zhiming Liu, Zhihua Zhang, Jinsheng Yang, Yanjie Lü, Yongjun Wang. Senescence Characteristics of Maize Leaves at Different Maturity Stages and Their Effect on Phyllosphere Bacteria [J]. Chinese Bulletin of Botany, 2024, 59(6): 1024-1040. |
| [14] | Juan Yang, Yuelei Zhao, Xiaoyuan Chen, Baobao Wang, Haiyang Wang. Regulation Mechanism and Breeding Application of Flowering Time in Maize [J]. Chinese Bulletin of Botany, 2024, 59(6): 912-931. |
| [15] | Hengyu Yan, Zhaoxia Li, Yubin Li. Research Progress on Heat Stress Impact on Maize Growth and Heat-Tolerant Maize Screening in China [J]. Chinese Bulletin of Botany, 2024, 59(6): 1007-1023. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||
