

植物学报 ›› 2023, Vol. 58 ›› Issue (5): 687-700.DOI: 10.11983/CBB22204 cstr: 32102.14.CBB22204
裘天航, 王安安, 李利, 王迎春, 崔继鹏, 王紫瑶, 王蕊, 崔素霞( )
)
收稿日期:2022-08-31
接受日期:2022-10-24
出版日期:2023-09-01
发布日期:2023-09-21
通讯作者:
*E-mail: sxcui@cnu.edu.cn
基金资助:
Qiu Tianhang, Wang An’an, Li Li, Wang Yingchun, Cui Jipeng, Wang Ziyao, Wang Rui, Cui Suxia( )
)
Received:2022-08-31
Accepted:2022-10-24
Online:2023-09-01
Published:2023-09-21
Contact:
*E-mail: sxcui@cnu.edu.cn
摘要: 芦苇(Phragmites australis)分布范围广, 生物量巨大, 环境适应性强。受芦苇多倍体基因组的限制, 在基因水平上解释其独特的物种特质非常困难。利用三代测序及多种方法研究2种生态型芦苇(2n=8x)中RCA基因(编码Rubisco活化酶)的类型、结构、表达和定位模式。结果表明, 芦苇基因组中有4类RCA基因, 均属于RCA2β。转录水平和蛋白质水平检测以及免疫胶体金定位结果表明, 芦苇RCA的基因表达模式和蛋白质定位具有明显的生态型特异性。与沼泽芦苇(SR)相比, 沙丘芦苇(DR)中RCA的表达水平较低, 且更倾向于产生RCA2-β2产物。通过分相的蛋白双向电泳和质谱方法, 鉴定到6个高表达的RCA同型物。在DR中这些同型物高比例地分布于膜相。研究表明当芦苇处于极度恶劣的沙漠环境中, RCA转移到膜相并参与叶绿体的膜保护机制。
裘天航, 王安安, 李利, 王迎春, 崔继鹏, 王紫瑶, 王蕊, 崔素霞. 2种生态型芦苇RCA基因特征及表达特异性. 植物学报, 2023, 58(5): 687-700.
Qiu Tianhang, Wang An’an, Li Li, Wang Yingchun, Cui Jipeng, Wang Ziyao, Wang Rui, Cui Suxia. Characteristics and Expression Specificity of RCA Genes in Two Ecotypes of Phragmites australis. Chinese Bulletin of Botany, 2023, 58(5): 687-700.
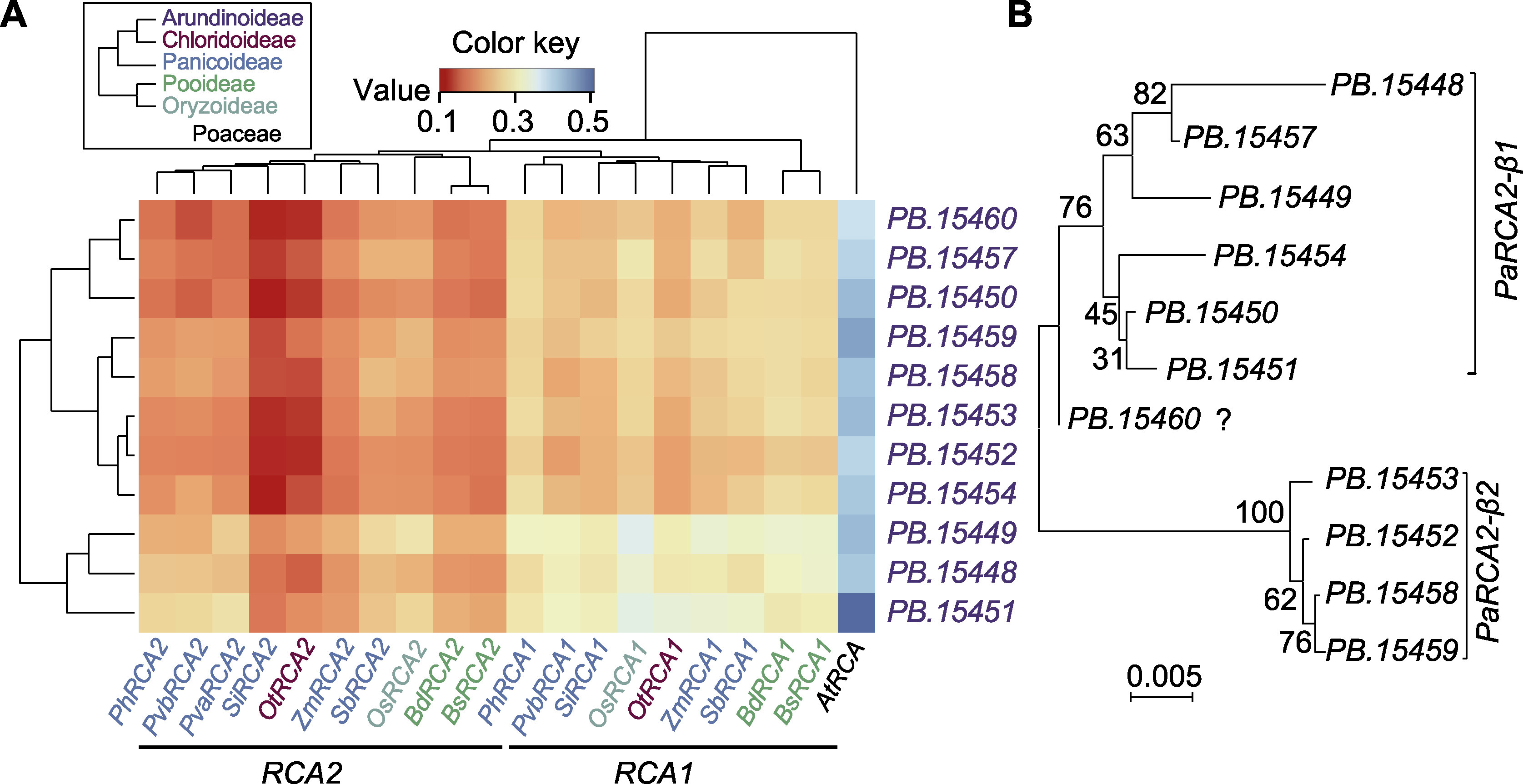
图1 芦苇预测RCA基因的分类 (A) 利用芦苇11个预测RCA基因和禾本科植物19个已知RCA基因序列生成的成对遗传距离矩阵图(颜色键显示从近(红色)到远(蓝色)的遗传距离; 图例中的系统发育树代表这些植物所属亚科的分类关系; 不同亚科以不同颜色突出显示, 外类群拟南芥则以黑色显示; 左侧是基于数据的泊松聚类树, 顶部是系统发育树); (B) 芦苇11个预测RCA基因的进化分析(所有系统发育树均使用RCA基因组序列(包括内含子和非翻译区)通过MEGA7的MUSCLE比对和邻接法构建)
Figure 1 Classification of predicted RCA genes in Phragmites australis (A) The pairwise genetic distance matrix was generated by using 11 predicted RCA genes of P. australis and 19 known RCA gene sequences of Poaceae (the color key shows genetic distance from close (red) to far (blue); the phylogeny tree in the legend shows the taxonomic relationships of Poaceae subfamilies used in this study; the different subfamilies are highlighted by different colors, while out-group Arabidopsis thaliana shown in black; the data based Poisson clustering tree is on the left, and the phylogeny tree is on the top); (B) Evolutionary analysis of 11 predicted RCA genes of P. australis (all phylogeny trees in this figure were constructed using RCA genomic sequences (including introns and UTR) by MUSCLE alignment of MEGA7 with Neighbor-joining method)
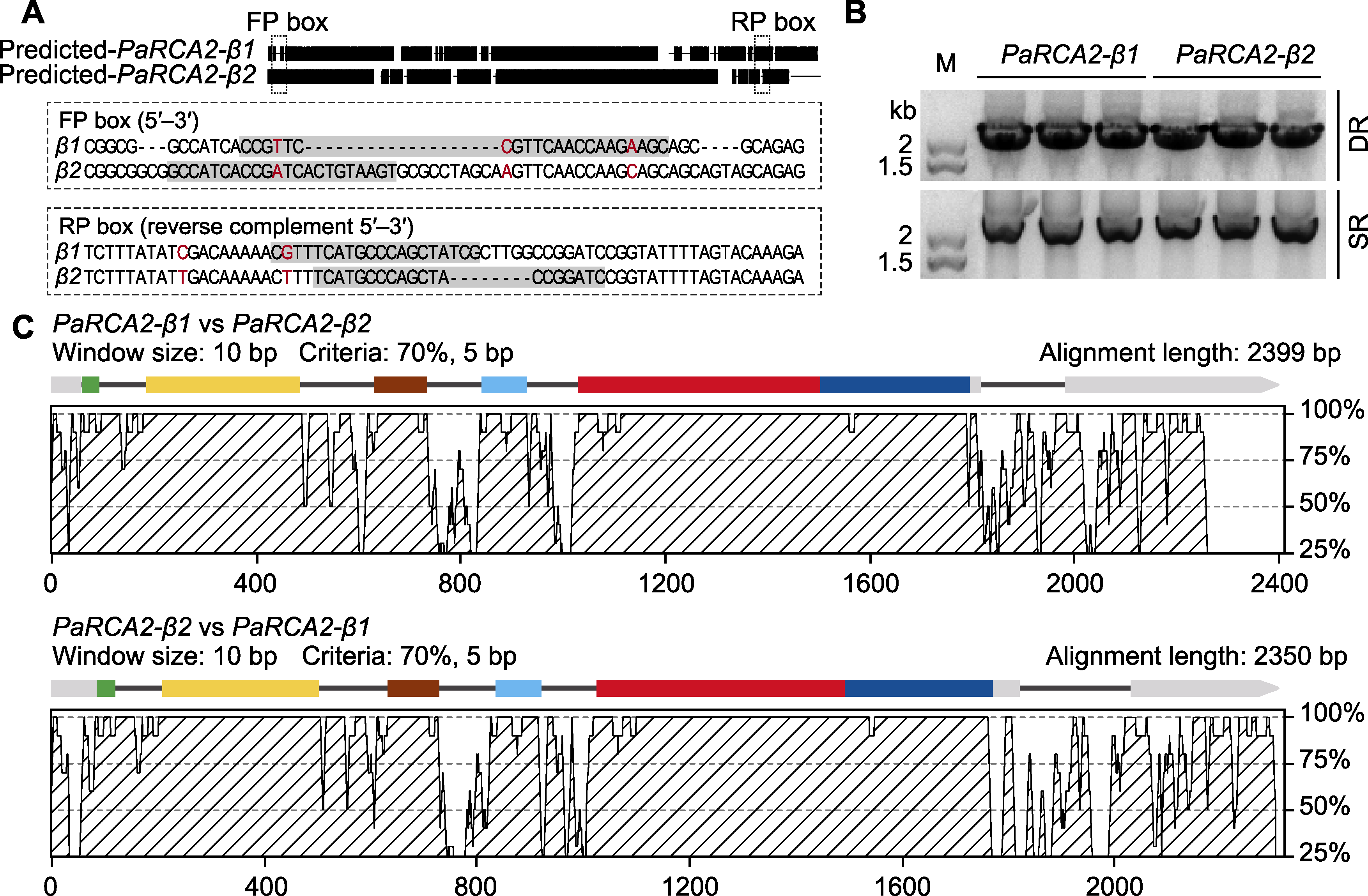
图2 芦苇中2个主要RCA基因的鉴定和差异分析 (A) RCA2基因特异性引物的设计(方框表示引物位置和序列; 差异核苷酸残基以红色字体突出显示); (B) 以基因组DNA为模板, 通过PCR扩增分别获得沼泽芦苇(SR)和沙丘芦苇(DR) RCA条带; (C) PaRCA2-β1和PaRCA2-β2序列和基因结构分析(用mVISTA的AVID程序进行对比; 外显子片段中的颜色显示与拟南芥RCA基因外显子的同源部分)
Figure 2 Identification and differential analysis of two RCA genes in Phragmites australis (A) The design of specific primers for the RCA2 genes (the boxes show the primer positions and sequences, the differential nucleotide residues are highlighted in red); (B) The RCA bands from swamp reed (SR) and desert-dune reed (DR) obtained by PCR amplification with genomic DNA as template; (C) Analysis of PaRCA2-β1 and PaRCA2-β2 in sequence and gene structure (the AVID program of mVISTA was used for comparative analysis; the color-coded exon segments were homologous to exons of Arabidopsis thaliana RCA gene)
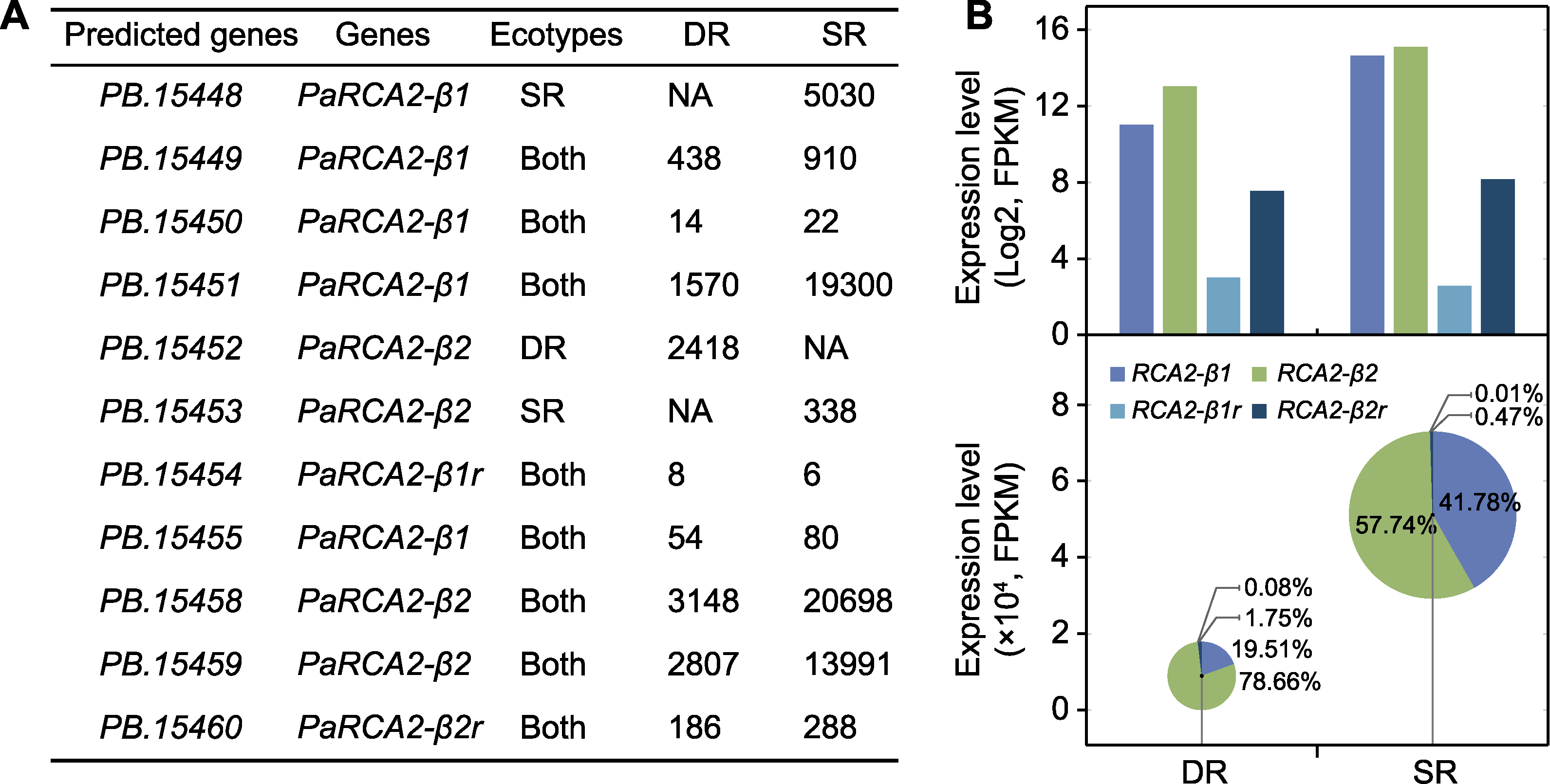
图3 2种生态型芦苇中RCA基因的表达模式 (A) PaRCA2基因的表达水平; (B) 沼泽芦苇(SR)和沙丘芦苇(DR)中RCA基因转录本差异的对数直方图(顶部)和气泡饼图(底部)。各类转录本占比以百分数标注。
Figure 3 Expression pattern of RCA in two ecotypes of Phragmites australis (A) Expression level of PaRCA2 genes; (B) Logarithmic histogram (top) and bubble-pie graph (bottom) of transcriptional differences of RCA genes in swamp reed (SR) and desert-dune reed (DR). The proportion of each transcript is annotated with percentage.
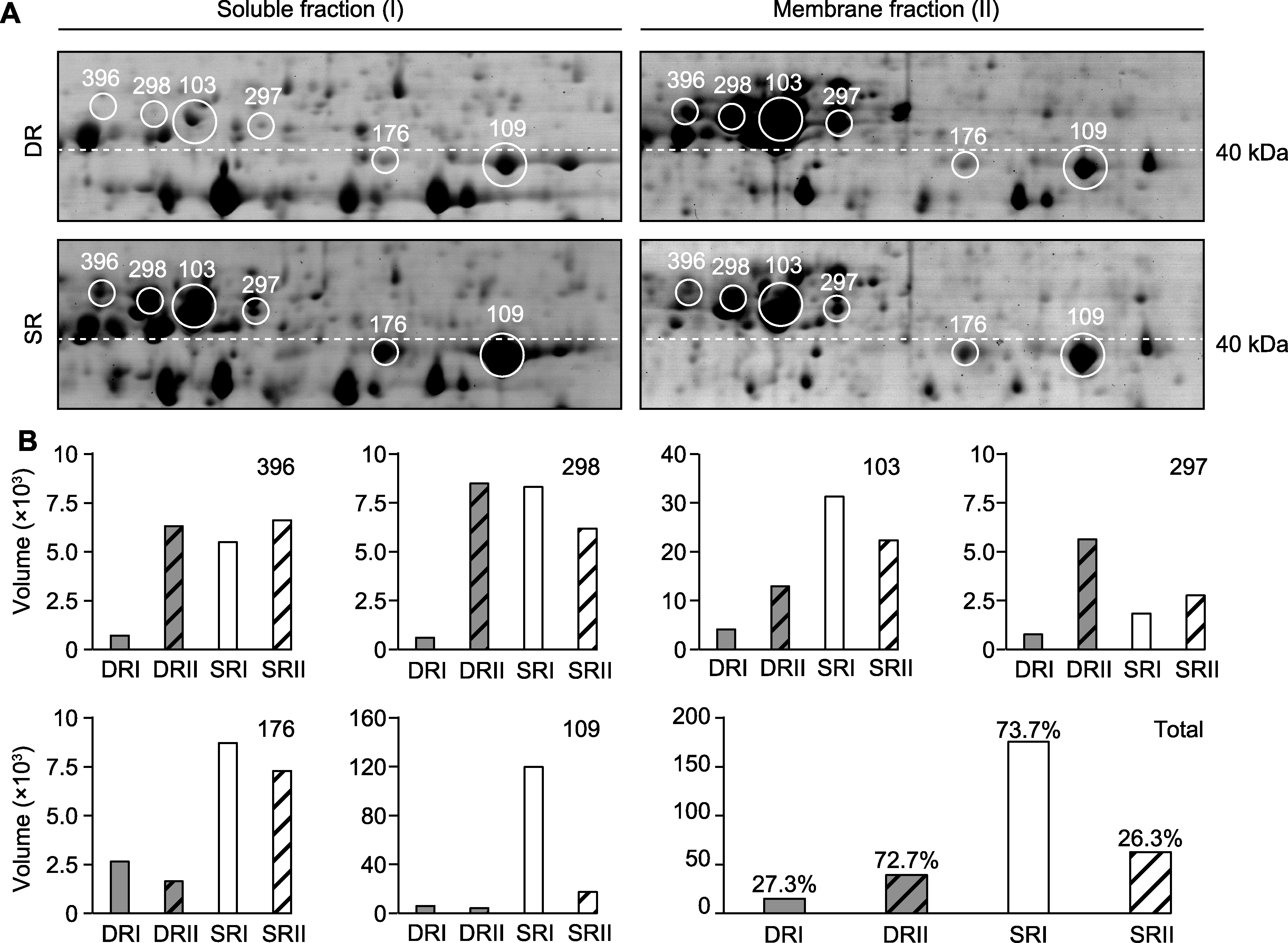
图4 2种生态型芦苇中RCA蛋白的双向电泳分析 (A) 可溶性组分(I)和膜组分(II)的蛋白质谱(用白色圆圈和数字标记PaRCA的6个同型物; 白色虚线表示40 kDa蛋白质的标准分子量位置); (B) PaRCA的定量分析, 显示6个RCA同型物和总RCA含量。SR: 沼泽芦苇; DR: 沙丘芦苇
Figure 4 Two-dimensional electrophoresis analysis of RCA proteins in two ecotypes of Phragmites australis (A) The protein profiles of the soluble fraction (I) and membrane fraction (II) (six isoforms of PaRCA are marked with white circles and numbers; the white dashed lines indicate the standard molecular weight position of 40 kDa protein); (B) Quantitative analysis of PaRCA, showing 6 RCA isoforms and total RCA content. SR: Swamp reed; DR: Desert-dune reed
| Ecotypes | Soluble fraction (FI) (%) | Membrane fraction (FII) (%) | FII/FI | |||
|---|---|---|---|---|---|---|
| L-RCA | S-RCA | L-RCA | S-RCA | L-RCA | S-RCA | |
| Swamp reed (SR) | 19.7 | 54.0 | 15.9 | 10.4 | 0.81 | 0.19 |
| Desert-dune reed (DR) | 11.4 | 15.9 | 61.7 | 11.0 | 5.41 | 0.69 |
表1 2种生态型芦苇中RCA同型物的分布差异
Table 1 Localization difference of RCA isoforms in two ecotypes of Phragmites australis
| Ecotypes | Soluble fraction (FI) (%) | Membrane fraction (FII) (%) | FII/FI | |||
|---|---|---|---|---|---|---|
| L-RCA | S-RCA | L-RCA | S-RCA | L-RCA | S-RCA | |
| Swamp reed (SR) | 19.7 | 54.0 | 15.9 | 10.4 | 0.81 | 0.19 |
| Desert-dune reed (DR) | 11.4 | 15.9 | 61.7 | 11.0 | 5.41 | 0.69 |

图5 RCA免疫金颗粒在2种生态型芦苇中的亚细胞分布 SR: 沼泽芦苇; DR: 沙丘芦苇; Mt: 线粒体; V: 液泡; Chl: 叶绿体; CW: 细胞壁; S: 细胞质基质; IS: 细胞间质; CS: 叶绿体基质; Gr: 基粒。Bars=1 μm
Figure 5 Subcellular distribution of RCA immune gold particles in two ecotypes of Phragmites australis SR: Swamp reed; DR: Desert-dune reed; Mt: Mitochondria; V: Vacuoles; Chl: Chloroplast; CW: Cell wall; S: Cytoplasmic stroma; IS: Intercellular stroma; CS: Chloroplast stroma; Gr: Granum. Bars=1 μm
| Ecotypes | Membrane fraction | Soluble fraction | |||
|---|---|---|---|---|---|
| Chl envelope | Thy membrane | Chl stroma | Thy lumen | Others | |
| Swamp reed (SR) | 17.5±7.0 | 29.1±7.5 | 19.1±6.3 | 15.7±5.2 | 18.7±7.3 |
| Desert-dune reed (DR) | 26.8±4.2 | 32.4±5.7 | 17.1±5.1 | 11.6±3.3 | 12.1±4.6 |
| P-value (t-test) | 9.95E-08 | 0.056 | 0.173 | 6.85E-04 | 1.26E-04 |
| DR/SR | 1.27 | 0.76 | |||
表2 2种生态型芦苇叶绿体中RCA免疫金颗粒的分布(数值为平均值±标准误, n=30)
Table 2 The distribution of RCA immune gold particles in the chloroplasts of two Phragmites australis ecotypes (values were means±SD, n=30)
| Ecotypes | Membrane fraction | Soluble fraction | |||
|---|---|---|---|---|---|
| Chl envelope | Thy membrane | Chl stroma | Thy lumen | Others | |
| Swamp reed (SR) | 17.5±7.0 | 29.1±7.5 | 19.1±6.3 | 15.7±5.2 | 18.7±7.3 |
| Desert-dune reed (DR) | 26.8±4.2 | 32.4±5.7 | 17.1±5.1 | 11.6±3.3 | 12.1±4.6 |
| P-value (t-test) | 9.95E-08 | 0.056 | 0.173 | 6.85E-04 | 1.26E-04 |
| DR/SR | 1.27 | 0.76 | |||
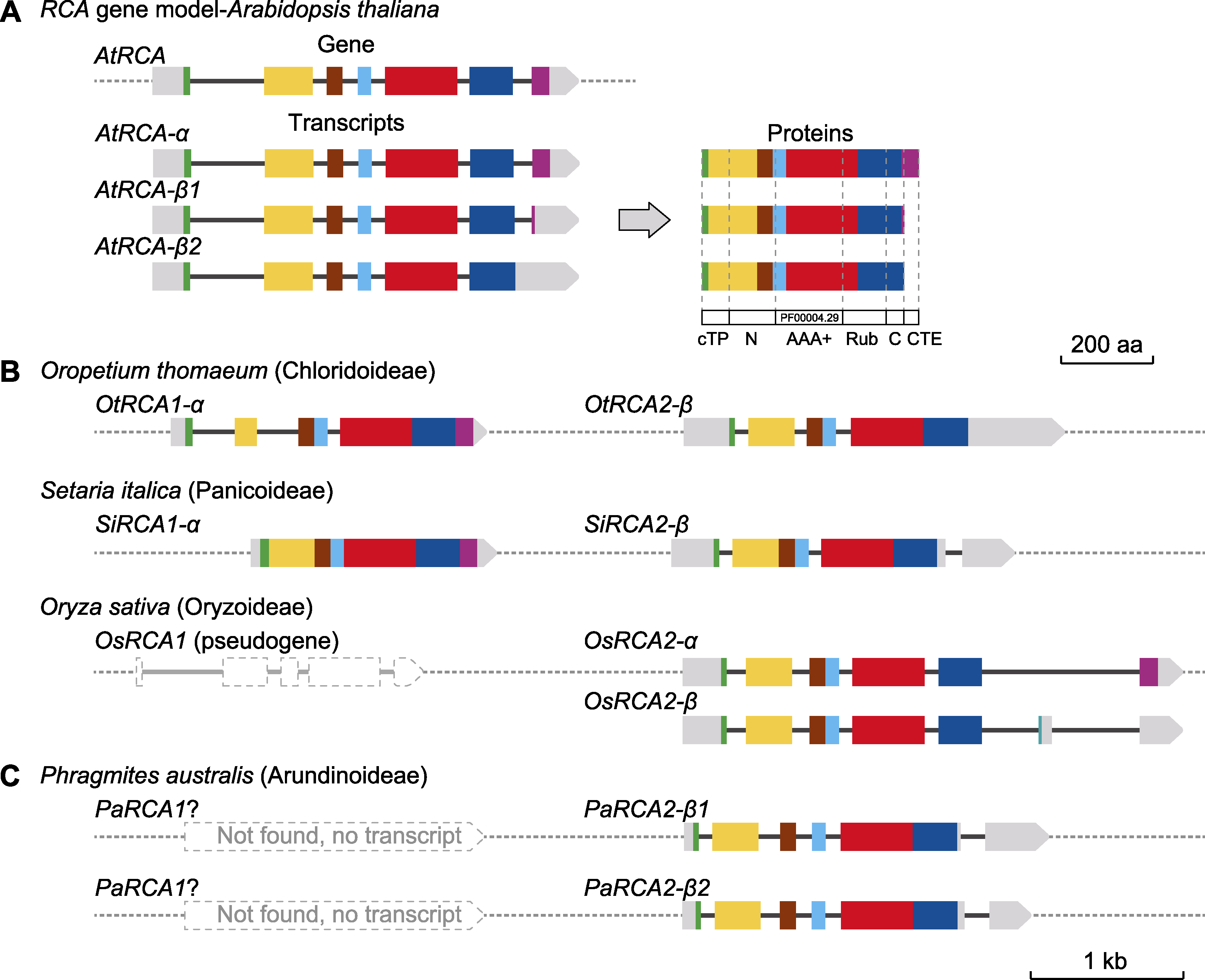
图6 芦苇和禾本科模式植物的RCA基因/转录本结构模型 (A) 拟南芥RCA基因结构及表达产物(7个外显子用不同颜色表示; 非翻译区为灰色, 深灰色细线代表内含子; 基因间序列用虚线表示); (B) 3种禾本科植物的RCA基因模型(复活草和谷子的基因特征相似; 水稻RCA1功能已丧失, RCA2同时具有2种表达产物); (C) 芦苇RCA基因模型。(B)和(C)中的颜色标注和定义通过序列同源性与拟南芥比对。cTP: 叶绿体转运肽; N: N端结构域; AAA+: ATP酶结构域; Rub: Rubisco大亚基结合结构域; C: C端结构域; CTE: 氧化还原相关C端延伸区
Figure 6 RCA gene/transcript models of Phragmites australis and Poaceae model plants (A) Gene structure and expression products of AtRCA (the seven exons were shown in different colors, gray represented untranslated region, thin dark gray lines represented introns, and dotted lines represented intergenic sequences); (B) RCA gene model in three Poaceae species (the gene characters were similar between Oropetium thomaeum and Setaria italica; pseudo OsRCA1 had lost its function, while OsRCA2 had two expression products simultaneously); (C) PaRCA gene model. Color labeling and definition in (B) and (C) were aligned with AtRCA by sequence similarity. cTP: Chloroplast transport peptide; N: N-terminal domain; AAA+: ATPase domain; Rub: Rubisco large subunit binding domain; C: C-terminal domain; CTE: Redox related C-terminal extension
| [1] | 张承烈, 陈国仓 (1991). 河西走廊不同生态类型芦苇的气体交换特点的研究. 生态学报 11, 250-255. |
| [2] | 张茜, 裘天航, 王安安, 周华健, 袁敏, 李利, 白素兰, 崔素霞 (2020). 北京地区芦苇资源状态及其多样性. 植物学报 55, 693-704. |
| [3] |
Bayramov S, Guliyev N (2014). Changes in Rubisco activase gene expression and polypeptide content in Brachypodium distachyon. Plant Physiol Biochem 81, 61-66.
DOI URL |
| [4] |
Boex-Fontvieille E, Daventure M, Jossier M, Hodges M, Zivy M, Tcherkez G (2014). Phosphorylation pattern of Rubisco activase in Arabidopsis leaves. Plant Biol 16, 550-557.
DOI URL |
| [5] |
Carmo-Silva E, Scales JC, Madgwick PJ, Martin AJP (2015). Optimizing Rubisco and its regulation for greater resource use efficiency. Plant Cell Environ 38, 1817-1832.
DOI URL |
| [6] |
Chen J, Wang P, Mi HL, Chen GY, Xu DQ (2010). Reversible association of Ribulose-1,5-bisphosphate carboxylase/oxygenase activase with the thylakoid membrane depends upon the ATP level and pH in rice without heat stress. J Exp Bot 61, 2939-2950.
DOI URL |
| [7] | Chen YX, Chen YS, Shi CM, Huang ZB, Zhang Y, Li SK, Li Y, Ye J, Yu C, Li Z, Zhang XQ, Wang J, Yang HM, Lin F, Chen Q (2018). SOAPnuke: a MapReduce acceleration- supported software for integrated quality control and preprocessing of high-throughput sequencing data. GigaScience 7, gix120. |
| [8] | Clevering OA, Lissner J (1999). Taxonomy, chromosome numbers, clonal diversity and population dynamics of Phragmites australis. Aquat Bot 66, 185-208. |
| [9] |
Cui S, Wang W, Zhang C (2002). Plant regeneration from callus cultures in two ecotypes of reed (Phragmites communis Trinius). In Vitro Cell Dev Biol Plant 38, 325-329.
DOI URL |
| [10] |
Cui SX, Hu J, Yang B, Shi L, Huang F, Tsai SN, Ngai SM, He YK, Zhang JH (2009). Proteomic characterization of Phragmites communis in ecotypes of swamp and desert dune. Proteomics 9, 3950-3967.
DOI URL |
| [11] |
Demirevska-kepova K, Hölzer R, Simova-stoilova L, Feller U (2005). Heat stress effects on Ribulose-1,5-bisphosphate carboxylase/oxygenase, Rubisco binding protein and Rubisco activase in wheat leaves. Biol Plant 49, 521-525.
DOI URL |
| [12] |
Deridder BP, Shybut ME, Dyle MC, Kremling KAG, Shapiro MB (2012). Changes at the 3′-untranslated region stabilize Rubisco activase transcript levels during heat stress in Arabidopsis. Planta 236, 463-476.
DOI URL |
| [13] |
Flecken M, Wang H, Popilka L, Hartl FU, Bracher A, Hayer-Hartl M (2020). Dual functions of a Rubisco activase in metabolic repair and recruitment to carboxysomes. Cell 183, 457-473.
DOI PMID |
| [14] |
Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I (2004). VISTA: computational tools for comparative genomics. Nucleic Acids Res 32 (Web Server issue),W273-9.
DOI PMID |
| [15] | Hall T (2011). BioEdit: an important software for molecular biology. GERF Bull Biosci 2, 60-61. |
| [16] | Kim D, Langmead B, Salzberg SL (2015). HISAT: a fast spliced aligner with low memory requirements. Nature Me- thods 12, 357-360. |
| [17] |
Kim SY, Stessman DJ, Wright DA, Spalding MH, Huber SC, Ort DR (2020). Arabidopsis plants expressing only the redox-regulated RCA-α isoform have constrained photosynthesis and plant growth. Plant J 103, 2250-2262.
DOI URL |
| [18] | Kumar RR, Goswami S, Singh K, Dubey K, Singh S, Sharma R, Verma N, Kala YK, Rai GK, Grover M, Mishra DC, Singh B, Pathak H, Chinnusamy V, Rai A, Praveen S (2016). Identification of putative Rubisco activase (TaRca1)—the catalytic chaperone regulating carbon assimilatory pathway in wheat (Triticum aestivum) un- der the heat stress. Front Plant Sci 7, 986. |
| [19] |
Kurek I, Chang TK, Bertain SM, Madrigal A, Liu L, Lassner MW, Zhu GH (2007). Enhanced thermostability of Arabidopsis Rubisco activase improves photosynthesis and growth rates under moderate heat stress. Plant Cell 19, 3230-3241.
DOI URL |
| [20] |
Li B, Dewey CN (2011). RSEM: accurate transcript quantification from RNA-seq data with or without a reference genome. BMC Bioinformatics 12, 323.
DOI PMID |
| [21] |
Li F, Fan G, Lu C, Xiao G, Zou C, Kohel RJ, Ma Z, Shang H, Ma X, Wu J, Liang X, Huang G, Percy RG, Liu K, Yang W, Chen W, Du X, Shi C, Yuan Y, Ye W, Liu X, Zhang X, Liu W, Wei H, Wei S, Huang G, Zhang X, Zhu S, Zhang H, Sun F, Wang X, Liang J, Wang J, He Q, Huang L, Wang J, Cui J, Song G, Wang K, Xu X, Yu JZ, Zhu Y, Yu S (2015). Genome sequence of cultivated upland cotton (Gossypium hirsutum TM-1) provides insights into genome evolution. Nat Biotechnol 33, 524-530.
DOI |
| [22] | Li H, Lin WF, Shen ZJ, Peng H, Zhou JJ, Zhu XY (2021). Physiological and proteomic analyses of different ecotypes of reed (Phragmites communis) in adaption to natural drought and salinity. Front Plant Sci 12, 1-15. |
| [23] |
Li L, Chen X, Shi L, Wang CJ, Fu B, Qiu TH, Cui SX (2017). A proteome translocation response to complex desert stress environments in perennial Phragmites sympatric ecotypes with contrasting water availability. Front Plant Sci 8, 511.
DOI PMID |
| [24] |
Love MI, Huber W, Anders S (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15, 550.
DOI URL |
| [25] | Merchant SS, Prochnik SE, Vallon O, Harris EH, Karpowicz SJ, Witman GB, Terry A, Salamov A, Fritz- Laylin LK, Maréchal-Drouard L, Marshall WF, Qu LH, Nelson DR, Sanderfoot AA, Spalding MH, Kapitonov VV, Ren QH, Ferris P, Lindquist E, Shapiro H, Lucas SM, Grimwood J, Schmutz J, Cardol P, Cerutti H, Chanfreau G, Chen CL, Cognat V, Croft MT, Dent R, Dutcher S, Fernández E, Fukuzawa H, González- Ballester D, González-Halphen D, Hallmann A, Hanikenne M, Hippler M, Inwood W, Jabbari K, Kalanon M, Kuras R, Lefebvre PA, Lemaire SD, Lobanov AV, Lohr M, Manuell A, Meier I, Mets L, Mittag M, Mittelmeier T, Moroney JV, Moseley J, Napoli C, Nedelcu AM, Niyogi K, Novoselov SV, Paulsen IT, Pazour G, Purton S, Ral JP, Riaño-Pachón DM, Riekhof W, Rymarquis L, Schroda M, Stern D, Umen J, Willows R, Wilson N, Zimmer SL, Allmer J, Balk J, Bisova K, Chen CJ, Elias M, Gendler K, Hauser C, Lamb MR, Ledford H, Long JC, Minagawa J, Page MD, Pan JM, Pootakham W, Roje S, Rose A, Stahlberg E, Terauchi AM, Yang PF, Ball S, Bowler C, Dieckmann CL, Gladyshev VN, Green P, Jorgensen R, Mayfield S, Mueller-Roeber B, Rajamani S, Sayre RT, Brokstein P, Dubchak I, Goodstein D, Hornick L, Huang YW, Jhaveri J, Luo YG, Martínez D, Ngau WCA, Otillar B, Poliakov A, Porter A, Szajkowski L, Werner G, Zhou KM, Grigoriev IV, Rokhsar DS, Grossman AR (2007). The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science 318, 245-250. |
| [26] |
Nagarajan R, Gill KS (2018). Evolution of Rubisco activase gene in plants. Plant Mol Biol 96, 69-87.
DOI PMID |
| [27] |
Oh DH, Kowalski KP, Quach QN, Wijesinghege C, Tanford P, Dassanayake M, Clay K (2022). Novel genome characteristics contribute to the invasiveness of Phragmites australis (common reed). Mol Ecol 31, 1142-1159.
DOI URL |
| [28] |
Porebski S, Bailey LG, Baum BR (1997). Modification of a CTAB DNA extraction protocol for plants containing high polysaccharide and polyphenol components. Plant Mol Biol Rep 15, 8-15.
DOI URL |
| [29] |
Portis AR Jr, Li C, Wang D, Salvucci ME (2008). Regulation of Rubisco activase and its interaction with Rubisco. J Exp Bot 59, 1597-1604.
DOI PMID |
| [30] |
Qian J, Rodermel SR (1993). Ribulose-1,5-bisphosphate carboxylase oxygenase activase cDNAs from Nicotiana tabacum. Plant Physiol 102, 683-684.
PMID |
| [31] |
Qiu TH, Cui SX (2021). Evolutionary analysis for Phragmites ecotypes based on full-length plastomes. Aquat Bot 170, 103349.
DOI URL |
| [32] |
Rozas J, Ferrer-Mata A, Sánchez-DelBarrio JC, Guirao- Rico S, Librado P, Ramos-Onsins SE, Sánchez-Gracia A (2017). DnaSP 6: DNA sequence polymorphism analysis of large datasets. Mol Biol Evol 34, 3299-3302.
DOI URL |
| [33] |
Salse J, Bolot S, Throude M, Jouffffe V, Piegu B, Quraishi UM, Calcagno T, Cooke R, Delseny M, Feuillet C (2008). Identification and characterization of shared duplications between rice and wheat provide new insight into grass genome evolution. Plant Cell 20, 11-24.
DOI PMID |
| [34] |
Salvucci ME, Crafts-Brandner SJ (2004). Relationship between the heat tolerance of photosynthesis and the thermal stability of Rubisco activase in plants from contrasting thermal environments. Plant Physiol 134, 1460-1470.
DOI PMID |
| [35] |
Salvucci ME, van de Loo FJ, Stecher D (2003). Two isoforms of Rubisco activase in cotton, the products of separate genes not alternative splicing. Planta 216, 736-744.
DOI PMID |
| [36] |
Salvucci ME, Werneke JM, Ogren WL, Portis AR Jr (1987). Purification and species distribution of Rubisco activase. Plant Physiol 84, 930-936.
DOI PMID |
| [37] |
Shen JB, Orozco EM, Ogren WL (1991). Expression of the two isoforms of spinach Ribulose-1,5-bisphosphate carboxylase activase and essentiality of the conserved lysine in the consensus nucleotide-binding domain. J Biol Chem 266, 8963-8968.
PMID |
| [38] |
Sheng X, Dong X, Zhang S, Jiang LP, Tan LL, Li X (2011). Unequal distribution of ubiquitinated proteins during Pinus bungeana pollen development. Trees 25, 407-414.
DOI URL |
| [39] |
To KY, Suen DF, Chen SCG (1999). Molecular characterization of Ribulose-1,5-bisphosphate carboxylase/oxygenase activase in rice leaves. Planta 209, 66-76.
DOI PMID |
| [40] | Tseng HHE (2016). Cogent: reconstructing the coding genome using full-length transcriptome sequences without a reference genome. Plant & Animal Genome Conference XXIV. San Diego, CA. |
| [41] |
Wang D, Li XF, Zhou ZJ, Feng XP, Yang WJ, Jiang DA (2010). Two Rubisco activase isoforms may play different roles in photosynthetic heat acclimation in the rice plant. Physiol Plant 139, 55-67.
DOI PMID |
| [42] |
Werneke JM, Zielinski RE, Ogren WL (1988). Structure and expression of spinach leaf cDNA encoding Ribulosebisphosphate carboxylase/oxygenase activase. Proc Natl Acad Sci USA 85, 787-791.
PMID |
| [43] | Wickham H (2016). Ggplot2: Elegant Graphics for Data Analysis (2nd edition). New York: Springer-Verlag. |
| [44] |
Yin Z, Meng F, Song H, Wang X, Xu XM, Yu DY (2010). Expression quantitative trait loci analysis of two genes encoding Rubisco activase in soybean. Plant Physiol 152, 1625-1637.
DOI PMID |
| [45] |
Yin Z, Zhang Z, Deng D, Chao MN, Gao QS, Wang YJ, Yang ZF, Bian YL, Hao DR, Xu CW (2014). Characterization of Rubisco activase genes in maize: an α-isoform gene functions alongside a β-isoform gene. Plant Physiol 164, 2096-2106.
DOI PMID |
| [46] |
Zhang JH, Wang LJ, Pan QH, Wang YZ, Zhan JC, Huang WD (2008). Accumulation and subcellular localization of heat shock proteins in young grape leaves during cross- adaptation to temperature stresses. Sci Hortic 117, 231-240.
DOI URL |
| [47] |
Zhang N, Kallis RP, Ewy RG, Portis AR (2002). Light modulation of Rubisco in Arabidopsis requires a capacity for redox regulation of the larger Rubisco activase isoform. Proc Natl Acad Sci USA 99, 3330-3334.
DOI PMID |
| [48] |
Zhang N, Portis AR (1999). Mechanism of light regulation of Rubisco: a specific role for the larger Rubisco activase isoform involving reductive activation by thioredoxin-f. Proc Natl Acad Sci USA 96, 9438-9443.
DOI PMID |
| [1] | 顾磊, 张棋, 张霞, 杨冰冰, 王芳岚, 刘文, 陈发菊. 盐肤木APETALA3/DEFICIENS同源基因的克隆与功能分析[J]. 植物学报, 2024, 59(4): 533-543. |
| [2] | 孟敬慈, 王国栋, 曹光兰, 胡楠林, 赵美玲, 赵延彤, 薛振山, 刘波, 朴文华, 姜明. 中国芦苇沼泽植物物种丰富度分布格局及其驱动因素[J]. 生物多样性, 2024, 32(2): 23194-. |
| [3] | 史欢欢, 雪穷, 于振林, 汪承焕. 密度、物种比例对盐沼植物种子萌发阶段种内、种间相互作用的影响[J]. 植物生态学报, 2023, 47(1): 77-87. |
| [4] | 张茜, 裘天航, 王安安, 周华健, 袁敏, 李利, 白素兰, 崔素霞. 北京地区芦苇资源状态及其多样性[J]. 植物学报, 2020, 55(6): 693-704. |
| [5] | 赵睿, 卜红梅, 宋献方, 高融瑾. 再生水补给河道内芦苇的光谱特征及其对水体氮和磷含量的响应[J]. 植物学报, 2020, 55(6): 666-676. |
| [6] | 郭瑞, 周际, 刘琪, 顾峰雪. 松嫩退化草地芦苇不同叶位叶片营养元素代谢特征[J]. 植物生态学报, 2018, 42(7): 734-740. |
| [7] | 李群, 赵成章, 赵连春, 王建良, 张伟涛, 姚文秀. 秦王川盐沼湿地芦苇比叶面积与叶片热耗散的关联性分析[J]. 植物生态学报, 2017, 41(9): 985-994. |
| [8] | 刘波, 吕宪国, 姜明, 张文广, 武海涛. 光照、水深交互作用对松嫩湿地芦苇种子萌发的影响[J]. 植物生态学报, 2015, 39(6): 616-620. |
| [9] | 陈清, 王义东, 郭长城, 王中良. 天津沼泽湿地芦苇叶片的碳稳定同位素比值分布特征及其环境影响因素[J]. 植物生态学报, 2015, 39(11): 1044-1052. |
| [10] | 胡楚琦, 刘金珂, 王天弘, 王文琳, 卢山, 周长芳. 三种盐胁迫对互花米草和芦苇光合作用的影响[J]. 植物生态学报, 2015, 39(1): 92-103. |
| [11] | 刘利丹, 介冬梅, 刘洪妍, 郭梅娥, 李楠楠. 东北地区芦苇植硅体的变化特征[J]. 植物生态学报, 2013, 37(9): 861-871. |
| [12] | 朱敏,张振华,于君宝,吴立新,韩广轩,杨利琼,邢庆会,谢宝华,毛培利,王光美. 氮沉降对黄河三角洲芦苇湿地土壤呼吸的影响[J]. 植物生态学报, 2013, 37(6): 517-529. |
| [13] | 杨利琼,韩广轩,于君宝,吴立新,朱敏,邢庆会,王光美,毛培利. 开垦对黄河三角洲湿地净生态系统CO2交换的影响[J]. 植物生态学报, 2013, 37(6): 503-516. |
| [14] | 贾庆宇, 周广胜, 周莉, 谢艳兵. 湿地芦苇植株氮素分布动态特征分析[J]. 植物生态学报, 2008, 32(4): 858-864. |
| [15] | 赵聪蛟, 邓自发, 周长芳, 关保华, 安树青, 陈琳, 陆霞梅. 氮水平和竞争对互花米草与芦苇叶特征的影响[J]. 植物生态学报, 2008, 32(2): 392-401. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||