

Chinese Bulletin of Botany ›› 2018, Vol. 53 ›› Issue (4): 487-501.DOI: 10.11983/CBB17082 cstr: 32102.14.CBB17082
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Guo Shulei1,2, Zhang Jun1, Qi Jianshuang1, Yue Runqing1, Han Xiaohua1, Yan Shufeng1, Lu Caixia1, Fu Xiaolei1, Chen Nana1, Ku Lixia2,*( ), Tie Shuanggui1,*(
), Tie Shuanggui1,*( )
)
Received:2017-04-13
Accepted:2017-10-25
Online:2018-07-01
Published:2018-09-11
Contact:
Ku Lixia,Tie Shuanggui
About author:† These authors contributed equally to this paper
Guo Shulei, Zhang Jun, Qi Jianshuang, Yue Runqing, Han Xiaohua, Yan Shufeng, Lu Caixia, Fu Xiaolei, Chen Nana, Ku Lixia, Tie Shuanggui. Analysis of Meta-quantitative Trait Loci and Their Candidate Genes Related to Leaf Shape in Maize[J]. Chinese Bulletin of Botany, 2018, 53(4): 487-501.
| Parents | Type of Pop. | Pop. size | No. of QTL for LL | No. of QTL for LW | No. of QTL for LAr | No. of QTL for LA | No./type of marker | Analysis method | Reference |
|---|---|---|---|---|---|---|---|---|---|
| B73×G79 | F2:3 | 214 | — | — | 7 | — | 185/RFLP | IM | Agrama et al., 1999 |
| B73×Mo17 | RIL | 180 | — | — | — | 9 | 192/SSR | CIM | Mickelson et al., 2002 |
| H21×Mo17 | F2:3 | 120 | — | — | — | 7 | 102/SSR | CIM | 于永涛等, 2006 |
| Zi330×K36 | F2:3 | 114 | — | — | — | 2 | 90/SSR | CIM | 于永涛等, 2006 |
| Ye478×Dan340 | F2:3 | 397 | — | — | — | 6 | 138/SSR | CIM | 路明等, 2007 |
| Mo17×Huangzao4 | RIL | 239 | 4 | 2 | 5 | — | 98/SSR | CIM | 郑祖平等, 2007 |
| Ye478×Wu312 | RIL | 218 | — | — | 7 | — | 184/SSR | ICIM | 刘建超等, 2010 |
| Yu82×Shen137 | F2:3 | 229 | 3 | 4 | 13 | 3 | 222/SSR | CIM | Ku et al., 2010, 2012a |
| Yu82×Yu87-1 | F2:3 | 256 | 5 | 5 | — | 5 | 216/SSR | CIM | Ku et al., 2012b |
| B73/Mo17 etc. | NAM | 4892 | 36 | 34 | — | 30 | 203000/SNP | GWAS | Tian et al., 2011 |
| N6×BT-1 | RIL | 250 | — | — | 6 | — | 207/SSR | CIM | 李贤唐等, 2011 |
| Y105×Y106 | F2 | 189 | 5 | 7 | 6 | 8 | 215/SSR | ICIM | 郭莹, 2012 |
| Y114×Y115 | F2 | 189 | 1 | 1 | 2 | 3 | 204/SSR | ICIM | 郭莹, 2012 |
| Ye478×Wu312 | RIL | 218 | 14 | 7 | 9 | — | 184/SSR | CIM | Cai et al., 2012 |
| Yu82×D132 | F2:3 | 245 | — | — | 18 | — | 204/SSR | CIM | Ku et al., 2012 |
| B73×1212 | RIL | 325 | 67 | 62 | — | — | 208/SSR | ICIM | 唐登国, 2013 |
| B73×Mo17 | RIL | 93 | 1 | 2 | — | 3 | IBM2 map | CIM | Wassom, 2013 |
| CY5×YL106 | F2:3 | 144 | — | — | — | 8 | 212/SSR | CIM | 刘正等, 2014 |
| Z58/87-1// PH6WC/Zi330 | CP | 228 | — | — | — | 13 | 225/SSR | IM | 张姿丽等, 2014b |
| T4×T19 | F2:3 | 232 | — | — | 4 | — | 81/SSR | CIM | 张姿丽等, 2014a |
| Yu82×Yu87-1 | RIL | 208 | — | 18 | — | — | 1370/SNP | CIM | Guo et al., 2015 |
| Yu82×Shen137 | RIL | 197 | — | 9 | — | — | 1411/SNP | CIM | Guo et al., 2015 |
| Zong3×Yu87-1 | RIL | 223 | — | 10 | — | — | 1479/SNP | CIM | Guo et al., 2015 |
| Yu537A×Shen137 | RIL | 212 | — | 9 | — | — | 1371/SNP | CIM | Guo et al., 2015 |
| B73×Mo17 | DH | 221 | 9 | 17 | 16 | 12 | 5935/bin markers | ICIM | 张志腾, 2015 |
| Xu178×K12 | RIL | 150 | — | — | — | 34 | 191/SSR | CIM | 常立国等, 2016 |
| M1-7×SYF | F2 | 259 | — | — | 36 | — | 218/SSR | CIM | 安允权等, 2016 |
| Yu82×D132 | RIL | 234 | 5 | 7 | 4 | — | 1226/SNP | CIM | Wei et al., 2016 |
Table 1 Summary of the QTL of leaf shape in maize reported previously
| Parents | Type of Pop. | Pop. size | No. of QTL for LL | No. of QTL for LW | No. of QTL for LAr | No. of QTL for LA | No./type of marker | Analysis method | Reference |
|---|---|---|---|---|---|---|---|---|---|
| B73×G79 | F2:3 | 214 | — | — | 7 | — | 185/RFLP | IM | Agrama et al., 1999 |
| B73×Mo17 | RIL | 180 | — | — | — | 9 | 192/SSR | CIM | Mickelson et al., 2002 |
| H21×Mo17 | F2:3 | 120 | — | — | — | 7 | 102/SSR | CIM | 于永涛等, 2006 |
| Zi330×K36 | F2:3 | 114 | — | — | — | 2 | 90/SSR | CIM | 于永涛等, 2006 |
| Ye478×Dan340 | F2:3 | 397 | — | — | — | 6 | 138/SSR | CIM | 路明等, 2007 |
| Mo17×Huangzao4 | RIL | 239 | 4 | 2 | 5 | — | 98/SSR | CIM | 郑祖平等, 2007 |
| Ye478×Wu312 | RIL | 218 | — | — | 7 | — | 184/SSR | ICIM | 刘建超等, 2010 |
| Yu82×Shen137 | F2:3 | 229 | 3 | 4 | 13 | 3 | 222/SSR | CIM | Ku et al., 2010, 2012a |
| Yu82×Yu87-1 | F2:3 | 256 | 5 | 5 | — | 5 | 216/SSR | CIM | Ku et al., 2012b |
| B73/Mo17 etc. | NAM | 4892 | 36 | 34 | — | 30 | 203000/SNP | GWAS | Tian et al., 2011 |
| N6×BT-1 | RIL | 250 | — | — | 6 | — | 207/SSR | CIM | 李贤唐等, 2011 |
| Y105×Y106 | F2 | 189 | 5 | 7 | 6 | 8 | 215/SSR | ICIM | 郭莹, 2012 |
| Y114×Y115 | F2 | 189 | 1 | 1 | 2 | 3 | 204/SSR | ICIM | 郭莹, 2012 |
| Ye478×Wu312 | RIL | 218 | 14 | 7 | 9 | — | 184/SSR | CIM | Cai et al., 2012 |
| Yu82×D132 | F2:3 | 245 | — | — | 18 | — | 204/SSR | CIM | Ku et al., 2012 |
| B73×1212 | RIL | 325 | 67 | 62 | — | — | 208/SSR | ICIM | 唐登国, 2013 |
| B73×Mo17 | RIL | 93 | 1 | 2 | — | 3 | IBM2 map | CIM | Wassom, 2013 |
| CY5×YL106 | F2:3 | 144 | — | — | — | 8 | 212/SSR | CIM | 刘正等, 2014 |
| Z58/87-1// PH6WC/Zi330 | CP | 228 | — | — | — | 13 | 225/SSR | IM | 张姿丽等, 2014b |
| T4×T19 | F2:3 | 232 | — | — | 4 | — | 81/SSR | CIM | 张姿丽等, 2014a |
| Yu82×Yu87-1 | RIL | 208 | — | 18 | — | — | 1370/SNP | CIM | Guo et al., 2015 |
| Yu82×Shen137 | RIL | 197 | — | 9 | — | — | 1411/SNP | CIM | Guo et al., 2015 |
| Zong3×Yu87-1 | RIL | 223 | — | 10 | — | — | 1479/SNP | CIM | Guo et al., 2015 |
| Yu537A×Shen137 | RIL | 212 | — | 9 | — | — | 1371/SNP | CIM | Guo et al., 2015 |
| B73×Mo17 | DH | 221 | 9 | 17 | 16 | 12 | 5935/bin markers | ICIM | 张志腾, 2015 |
| Xu178×K12 | RIL | 150 | — | — | — | 34 | 191/SSR | CIM | 常立国等, 2016 |
| M1-7×SYF | F2 | 259 | — | — | 36 | — | 218/SSR | CIM | 安允权等, 2016 |
| Yu82×D132 | RIL | 234 | 5 | 7 | 4 | — | 1226/SNP | CIM | Wei et al., 2016 |
| Bin | mQTL/QTL | CI (cM) | Candidate gene | Annotation | Homologous gene | E-value | Reference |
|---|---|---|---|---|---|---|---|
| 1.02 | mQTLW1-1 | 153.8-158.3 | GRMZM2G480386 | NAL7-like | Os03g0162000/YUCCA7 | 0 | Fujino et al., 2008 |
| 1.04 | q12SevLW1 | 270.6-286.2 | GRMZM2G011483 | SRL2-like | Os03g19520/SRL2 | 0 | Liu et al., 2016 |
| 1.05 | mQTLW1-2 | 485.9-505.6 | GRMZM2G141955 | YABBY6-like | Os12g42610/YAB6 | 3E-88 | Toriba et al., 2007; Zhang et al., 2009 |
| GRMZM2G003509 | PHB-like | AT2G34710/AtHB14 | 0 | Kim et al., 2003; Mallory et al., 2004 | |||
| 1.09 | mQTLW1-3 | 820.6-845.5 | GRMZM2G018414 | GRF8 | AT4G37740/AtGRF2 | 4E-37 | Debernardi et al., 2014 |
| 1.10 | q4LWm139 | 873.7-902.1 | GRMZM2G017087 | KNOTTED1 | AT4G08150/KN1 | 2E-114 | Ramirez et al., 2009 |
| GRMZM2G178261 | GRF1-like | AT2G22840/AtGRF1 | 2E-66 | Kim et al., 2003 | |||
| GRMZM2G180246 | AN3/GIF1 | AT5G28640/AtAN3/GIF1 | 1E-36 | Nelissen et al., 2015 | |||
| 1.11 | q4LLm155 | 941.6-1022.3 | GRMZM2G135447 | OSH43-like | Os03g0771500/OsH43 | 4E-99 | Sentoku et al., 2000 |
| 2.02 | q3LAr2a | 67.5-79.7 | GRMZM2G174784 | AP2-like | AT4G36920/AP2 | 7E-102 | Würschum et al., 2006; Mlotshwa et al., 2006 |
| 2.02 | mQTLW2-1 | 118.3-147.8 | GRMZM2G102346 | NAL1-like | Os04g52479/NAL1 | 0 | Kubo et al., 2017 |
| GRMZM2G041223 | GRF6 | AT3G13960/AtGRF5 | 1E-36 | Horiguchi et al., 2006; Debernardi et al., 2014 | |||
| 2.06 | q4RLAr2 | 355.7-374.4 | GRMZM2G444808 | HYL1-like | AT1G09700/HYL1 | 6E-56 | Liu et al., 2011 |
| 2.06 | q8SecLW2 | 370.0-381.6 | GRMZM2G083972 | LNG2-like | AT3G02170/LNG2 | 6E-57 | Lee et al., 2006 |
| GRMZM2G161382 | CYCD3;3-like | AT3G50070/CYCD3;3 | 5E-56 | Dewitte et al., 2007 | |||
| 2.07 | q7LL2b | 450.0-461.9 | GRMZM2G004619 | GRF4 | AT4G37740/AtGRF2 | 5E-28 | Kim et al., 2003 |
| 2.10 | q12LAr2 | 713.1-732.4 | GRMZM2G146688 | ANT-like | AT4G37750/ANT | 1E-131 | Mizukami and Fis- cher, 2000 |
| 3.02 | q4LWm309 | 69.6-90.3 | GRMZM2G143235 | ROT3-like | AT4G36380/ROT3 | 1E-176 | Kim et al., 1998 |
| 3.06 | mQTLW3-2 | 368.5-390.3 | GRMZM2G118250 | AS2-like | AT1G65620/AS2 | 1E-66 | Iwakawa et al., 2007 |
| 3.08 | q4LWm410 | 613.6-653.7 | GRMZM2G437460 | ARF3-like | AT2G33860/ARF3/ETT | 1E-169 | Kelley et al., 2012 |
| 4.04 | q13NLW4 | 220.5-230.6 | GRMZM2G402653 | OSH1-like | Os03g0727000/OsH1 | 1E-75 | Matsuoka et al., 1993 |
| 4.06 | mQTLW4 | 373.7-399.4 | GRMZM2G124566 | GRF9-like | AT2G36400/AtGRF3 | 1E-36 | Kim et al., 2003 |
| 4.08 | q7LL4 | 429.5-451.0 | GRMZM2G075117 | CYCD3;1-like | AT4G34160/CYCD3;1 | 4E-47 | Dewitte et al., 2007 |
| GRMZM2G105335 | GRF3 | AT3G13960/AtGRF5 | 8E-49 | Horiguchi et al., 2006; Debernardi et al., 2014 | |||
| 5.03 | q4LWm602 | 268.8-295.0 | GRMZM2G361518 | AGO10-like | AT5G43810/AGO10/ZLL | 0 | Zhu et al., 2011; Roodbarkelari et al., 2015 |
| 5.03 | q4LLm605 | 280.1-296.1 | GRMZM2G171349 | COW1-like | AT4G34580/COW1 | 3E-66 | Woo et al., 2007 |
| 5.06 | q4LLm661 | 500.7-521.1 | GRMZM2G034876 | GRF1 | AT3G13960/AtGRF5 | 1E-47 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 5.07 | mQTLW5-3 | 556.7-578.5 | GRMZM5G893117 | GRF9 | AT3G13960/AtGRF5 | 1E-27 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 6.01 | q8FirLW6-1 | 82.3-93.9 | GRMZM2G122537 | PRS1-like | AT2G28610/PRS1 | 6E-29 | Matsumoto and Ok- ada, 2001; Nakata et al., 2012 |
| GRMZM2G020187 | rgd1/lbl1 | AT5G23570/SGS3 | 1E-157 | Dotto et al., 2014 | |||
| mQTL/QTL | CI (cM) | Candidate gene | Annotation | Homologous gene | E-value | Reference | |
| 6.01 | mQTLAr6 | 92.8-108.9 | GRMZM2G098594 | GRF14 | AT3G13960/AtGRF5 | 1E-36 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 6.02 | q7PLL6 | 123.0-144.8 | GRMZM2G073671 | CYCB2;3-like | AT1G20610/CYCB2;3 | 1E-144 | Eloy et al., 2011 |
| GRMZM2G157820 | CLF-like | AT2G23380/CLF | 0 | Menges et al., 2005; Schatlowski et al., 2010 | |||
| 6.04 | q4LWm706 | 158.2-205.1 | GRMZM5G850129 | GRF7 | AT3G13960/AtGRF5 | 4E-41 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 7.00 | q4LWm762 | 8.0-93.0 | GRMZM2G028041 | OSH15-like | Os07g0129700/OsH15 | 1.E-168 | Nagasaki et al., 2001 |
| GRMZM2G019200 | DRL1-like | Os11g0312782/DRL1 | 0 | Jun et al., 2015 | |||
| 7.02 | q12EigLW7 | 200.7-217.9 | GRMZM2G107377 | CYCD3;2-like | AT5G67260/CYCD3;2 | 8E-49 | Dewitte et al., 2007 |
| 7.02 | q12FirLW7 | 229.3-246.3 | GRMZM2G082264 | mwp1 | Os09g23200/SLL1/Kan1 | 2E-170 | Candela et al., 2008 |
| 7.03 | q3LW7 | 380.6-498.2 | GRMZM2G096709 | GRF10 | AT4G37740/AtGRF2 | 4E-29 | Kim et al., 2003; Deb- ernardi et al., 2014 |
| 8.08 | q3LW8 | 599.1-603.8 | GRMZM5G874163 | ARF4-like | AT5G60450/ARF4 | 7E-120 | Pekker et al., 2005; Hunter et al., 2006 |
| 9.03 | mQTLW9 | 206.1-230.6 | GRMZM2G119359 | GRF12 | AT2G06200/AtGRF6 | 1E-30 | Kim et al., 2003; Liang et al., 2014 |
| GRMZM5G870176 | NRL-like | Os12g36890/CSLD4 | 0 | Hu et al., 2010 | |||
| 10.04 | mQTLW10 | 295.6-327.3 | GRMZM2G078641 | GRF2 | GRAT3G13960/AtGRF7 | 3E-156 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 10.06 | q1HNLAr10 | 367.0-383.9 | GRMZM2G061287 | CYCB2;4-like | AT1G76310/CYCB2;4 | 9E-148 | Menges et al., 2005; Eloy et al., 2011 |
Table 2 Candidate genes for maize leaf shape of mQTL/QTL
| Bin | mQTL/QTL | CI (cM) | Candidate gene | Annotation | Homologous gene | E-value | Reference |
|---|---|---|---|---|---|---|---|
| 1.02 | mQTLW1-1 | 153.8-158.3 | GRMZM2G480386 | NAL7-like | Os03g0162000/YUCCA7 | 0 | Fujino et al., 2008 |
| 1.04 | q12SevLW1 | 270.6-286.2 | GRMZM2G011483 | SRL2-like | Os03g19520/SRL2 | 0 | Liu et al., 2016 |
| 1.05 | mQTLW1-2 | 485.9-505.6 | GRMZM2G141955 | YABBY6-like | Os12g42610/YAB6 | 3E-88 | Toriba et al., 2007; Zhang et al., 2009 |
| GRMZM2G003509 | PHB-like | AT2G34710/AtHB14 | 0 | Kim et al., 2003; Mallory et al., 2004 | |||
| 1.09 | mQTLW1-3 | 820.6-845.5 | GRMZM2G018414 | GRF8 | AT4G37740/AtGRF2 | 4E-37 | Debernardi et al., 2014 |
| 1.10 | q4LWm139 | 873.7-902.1 | GRMZM2G017087 | KNOTTED1 | AT4G08150/KN1 | 2E-114 | Ramirez et al., 2009 |
| GRMZM2G178261 | GRF1-like | AT2G22840/AtGRF1 | 2E-66 | Kim et al., 2003 | |||
| GRMZM2G180246 | AN3/GIF1 | AT5G28640/AtAN3/GIF1 | 1E-36 | Nelissen et al., 2015 | |||
| 1.11 | q4LLm155 | 941.6-1022.3 | GRMZM2G135447 | OSH43-like | Os03g0771500/OsH43 | 4E-99 | Sentoku et al., 2000 |
| 2.02 | q3LAr2a | 67.5-79.7 | GRMZM2G174784 | AP2-like | AT4G36920/AP2 | 7E-102 | Würschum et al., 2006; Mlotshwa et al., 2006 |
| 2.02 | mQTLW2-1 | 118.3-147.8 | GRMZM2G102346 | NAL1-like | Os04g52479/NAL1 | 0 | Kubo et al., 2017 |
| GRMZM2G041223 | GRF6 | AT3G13960/AtGRF5 | 1E-36 | Horiguchi et al., 2006; Debernardi et al., 2014 | |||
| 2.06 | q4RLAr2 | 355.7-374.4 | GRMZM2G444808 | HYL1-like | AT1G09700/HYL1 | 6E-56 | Liu et al., 2011 |
| 2.06 | q8SecLW2 | 370.0-381.6 | GRMZM2G083972 | LNG2-like | AT3G02170/LNG2 | 6E-57 | Lee et al., 2006 |
| GRMZM2G161382 | CYCD3;3-like | AT3G50070/CYCD3;3 | 5E-56 | Dewitte et al., 2007 | |||
| 2.07 | q7LL2b | 450.0-461.9 | GRMZM2G004619 | GRF4 | AT4G37740/AtGRF2 | 5E-28 | Kim et al., 2003 |
| 2.10 | q12LAr2 | 713.1-732.4 | GRMZM2G146688 | ANT-like | AT4G37750/ANT | 1E-131 | Mizukami and Fis- cher, 2000 |
| 3.02 | q4LWm309 | 69.6-90.3 | GRMZM2G143235 | ROT3-like | AT4G36380/ROT3 | 1E-176 | Kim et al., 1998 |
| 3.06 | mQTLW3-2 | 368.5-390.3 | GRMZM2G118250 | AS2-like | AT1G65620/AS2 | 1E-66 | Iwakawa et al., 2007 |
| 3.08 | q4LWm410 | 613.6-653.7 | GRMZM2G437460 | ARF3-like | AT2G33860/ARF3/ETT | 1E-169 | Kelley et al., 2012 |
| 4.04 | q13NLW4 | 220.5-230.6 | GRMZM2G402653 | OSH1-like | Os03g0727000/OsH1 | 1E-75 | Matsuoka et al., 1993 |
| 4.06 | mQTLW4 | 373.7-399.4 | GRMZM2G124566 | GRF9-like | AT2G36400/AtGRF3 | 1E-36 | Kim et al., 2003 |
| 4.08 | q7LL4 | 429.5-451.0 | GRMZM2G075117 | CYCD3;1-like | AT4G34160/CYCD3;1 | 4E-47 | Dewitte et al., 2007 |
| GRMZM2G105335 | GRF3 | AT3G13960/AtGRF5 | 8E-49 | Horiguchi et al., 2006; Debernardi et al., 2014 | |||
| 5.03 | q4LWm602 | 268.8-295.0 | GRMZM2G361518 | AGO10-like | AT5G43810/AGO10/ZLL | 0 | Zhu et al., 2011; Roodbarkelari et al., 2015 |
| 5.03 | q4LLm605 | 280.1-296.1 | GRMZM2G171349 | COW1-like | AT4G34580/COW1 | 3E-66 | Woo et al., 2007 |
| 5.06 | q4LLm661 | 500.7-521.1 | GRMZM2G034876 | GRF1 | AT3G13960/AtGRF5 | 1E-47 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 5.07 | mQTLW5-3 | 556.7-578.5 | GRMZM5G893117 | GRF9 | AT3G13960/AtGRF5 | 1E-27 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 6.01 | q8FirLW6-1 | 82.3-93.9 | GRMZM2G122537 | PRS1-like | AT2G28610/PRS1 | 6E-29 | Matsumoto and Ok- ada, 2001; Nakata et al., 2012 |
| GRMZM2G020187 | rgd1/lbl1 | AT5G23570/SGS3 | 1E-157 | Dotto et al., 2014 | |||
| mQTL/QTL | CI (cM) | Candidate gene | Annotation | Homologous gene | E-value | Reference | |
| 6.01 | mQTLAr6 | 92.8-108.9 | GRMZM2G098594 | GRF14 | AT3G13960/AtGRF5 | 1E-36 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 6.02 | q7PLL6 | 123.0-144.8 | GRMZM2G073671 | CYCB2;3-like | AT1G20610/CYCB2;3 | 1E-144 | Eloy et al., 2011 |
| GRMZM2G157820 | CLF-like | AT2G23380/CLF | 0 | Menges et al., 2005; Schatlowski et al., 2010 | |||
| 6.04 | q4LWm706 | 158.2-205.1 | GRMZM5G850129 | GRF7 | AT3G13960/AtGRF5 | 4E-41 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 7.00 | q4LWm762 | 8.0-93.0 | GRMZM2G028041 | OSH15-like | Os07g0129700/OsH15 | 1.E-168 | Nagasaki et al., 2001 |
| GRMZM2G019200 | DRL1-like | Os11g0312782/DRL1 | 0 | Jun et al., 2015 | |||
| 7.02 | q12EigLW7 | 200.7-217.9 | GRMZM2G107377 | CYCD3;2-like | AT5G67260/CYCD3;2 | 8E-49 | Dewitte et al., 2007 |
| 7.02 | q12FirLW7 | 229.3-246.3 | GRMZM2G082264 | mwp1 | Os09g23200/SLL1/Kan1 | 2E-170 | Candela et al., 2008 |
| 7.03 | q3LW7 | 380.6-498.2 | GRMZM2G096709 | GRF10 | AT4G37740/AtGRF2 | 4E-29 | Kim et al., 2003; Deb- ernardi et al., 2014 |
| 8.08 | q3LW8 | 599.1-603.8 | GRMZM5G874163 | ARF4-like | AT5G60450/ARF4 | 7E-120 | Pekker et al., 2005; Hunter et al., 2006 |
| 9.03 | mQTLW9 | 206.1-230.6 | GRMZM2G119359 | GRF12 | AT2G06200/AtGRF6 | 1E-30 | Kim et al., 2003; Liang et al., 2014 |
| GRMZM5G870176 | NRL-like | Os12g36890/CSLD4 | 0 | Hu et al., 2010 | |||
| 10.04 | mQTLW10 | 295.6-327.3 | GRMZM2G078641 | GRF2 | GRAT3G13960/AtGRF7 | 3E-156 | Horiguchi et al., 2006; Debernardi et al., 2014 |
| 10.06 | q1HNLAr10 | 367.0-383.9 | GRMZM2G061287 | CYCB2;4-like | AT1G76310/CYCB2;4 | 9E-148 | Menges et al., 2005; Eloy et al., 2011 |

Figure 1 Distribution of leaf shape mQTL on maize chromosomes in the integrated mapChr: Chromosome. The red region is the identified location of the overlaps of mQTL on the chromosome; The position distribution and the name for mQTL are listed in the left of chromosome; Marker name and genetic distance (cM) are listed in the right of chromosome.
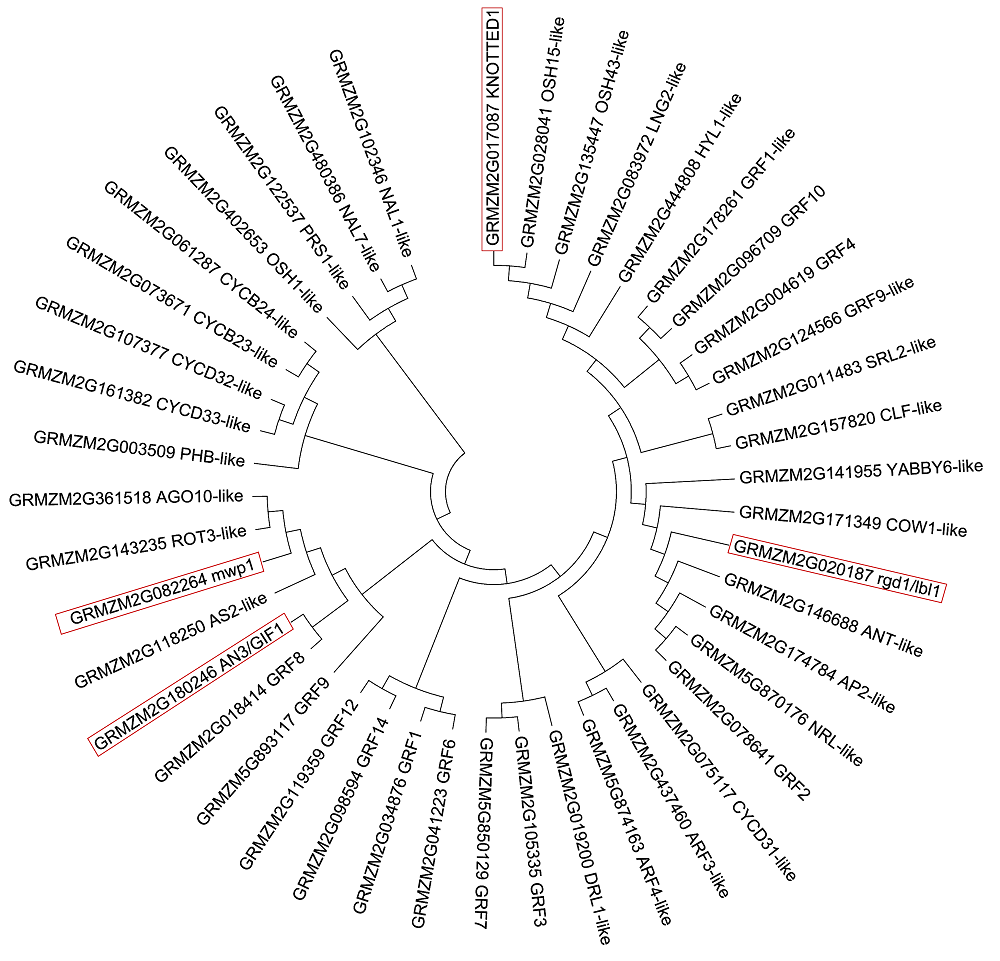
Figure 2 Phylogenetic trees of amino acid sequences of 44 candidate genes related to leaf shape of maizeFunction of candidate genes which have been known and cloned related to maize leaf shape are highlighted in the box
| 42 | Lee YK, Kim GT, Kim IJ, Park J, Kwak SS, Choi G, Chung WI (2006). LONGIFOLIA1 and LONGIFOLIA2, two homologous genes, regulate longitudinal cell elongation in Arabidopsis. Development 133, 4305-4314. |
| 43 | Liang G, He H, Li Y, Wang F, Yu DQ (2014). Molecular mechanism of microRNA396 mediating pistil development in Arabidopsis.Plant Physiol 164, 249-258. |
| 44 | Liu XF, Li M, Liu K, Tang D, Sun MF, Li YF, Shen Y, Du GJ, Cheng ZK (2016). Semi-Rolled Leaf 2 modulates rice leaf rolling by regulating abaxial side cell differentiation. J Exp Bot 67, 2139-2150. |
| 45 | Liu ZY, Jia LG, Wang H, He YK (2011). HYL1 regulates the balance between adaxial and abaxial identity for leaf flattening via miRNA-mediated pathways.J Exp Bot 62, 4367-6381. |
| 46 | Luan WJ, Liu YQ, Zhang FX, Song YL, Wang ZY, Peng YK, Sun ZX (2011). OsCD1 encodes a putative member of the cellulose synthase-like D sub-family and is essential for rice plant architecture and growth. Plant Biotechnol J 9, 513-524. |
| 47 | Mallory AC, Reinhart BJ, Jones-Rhoades MW, Tang GL, Zamore PD, Barton MK, Bartel DP (2004). MicroRNA control of PHABULOSA in leaf development: importance of pairing to the microRNA 5′ region.EMBO J 23, 3356-3364. |
| 48 | Matsumoto N, Okada K (2001). A homeobox gene, PRES- SED FLOWER, regulates lateral axis-dependent deve- lopment of Arabidopsis flowers. Genes Dev 15, 3355-3364. |
| 49 | Matsuoka M, Ichikawa H, Saito A, Tada Y, Fujimura T, Kano-Murakami Y (1993). Expression of a rice homeobox gene causes altered morphology of transgenic plants.Plant Cell 5, 1039-1048. |
| 50 | Menges M, de Jager SM, Gruissem W, Murray JAH (2005). Global analysis of the core cell cycle regulators of Arabidopsis identifies novel genes, reveals multiple and highly specific profiles of expression and provides a coherent model for plant cell cycle control.Plant J 41, 546-566. |
| 51 | Mickelson SM, Stuber CS, Senior L, Kaeppler SM (2002). Quantitative trait loci controlling leaf and tassel traits in a B73 × Mo17 population of maize.Crop Sci 42, 1902-1909. |
| 52 | Mizukami Y, Fischer RL (2000). Plant organ size control: AINTEGUMENTA regulates growth and cell numbers during organogenesis. Proc Natl Acad Sci USA 97, 942-947. |
| 53 | Mlotshwa S, Yang ZY, Kim YJ, Chen XM (2006). Floral patterning defects induced by Arabidopsis APETALA2 and microRNA172 expression in Nicotiana benthamiana.Plant Mol Biol 61, 781-793. |
| 54 | Nagasaki H, Sakamoto T, Sato Y, Matsuoka M (2001). Functional analysis of the conserved domains of a rice KNOX homeodomain protein, OSH15.Plant Cell 13, 2085-2098. |
| 55 | Nakata M, Matsumoto N, Tsugeki R, Rikirsch E, Laux T, Okada K (2012). Roles of the middle domain-specific WUSCHEL-RELATED HOMEOBOX genes in early development of leaves in Arabidopsis. Plant Cell 24, 519-535. |
| 56 | Nardmann J, Ji JB, Werr W, Scanlon MJ (2004). The maize duplicate genes narrow sheath 1 and narrow sheath 2 encode a conserved homeobox gene function in a lateral domain of shoot apical meristems. Development 131, 2827-2839. |
| 57 | Nelissen H, Eeckhout D, Demuynck K, Persiau G, Walton A, Van Bel M, Vervoort M, Candaele J, de Block J, Aes- aert S, Van Lijsebettens M, Goormachtig S, Vandepoele K, Van Leene J, Muszynski M, Gevaert K, Inzé D, De Jaeger G (2015). Dynamic changes in ANGUSTIFOLIA3 complex composition reveal a growth regulatory mechanism in the maize leaf.Plant Cell 27, 1605-1619. |
| 58 | Pekker I, Alvarez JP, Eshed Y (2005). Auxin response factors mediate Arabidopsis organ asymmetry via modulation of KANADI activity.Plant Cell 17, 2899-2910. |
| 59 | Qi J, Qian Q, Bu QY, Li SY, Chen Q, Sun JQ, Liang WX, Zhou YH, Chu CC, Li XG, Ren FG, Palme K, Zhao BR, Chen JF, Chen MS, Li CY (2008). Mutation of the rice Narrow leaf1 gene, which encodes a novel protein, affects vein patterning and polar auxin transport. Plant Physiol 147, 1947-1959. |
| 60 | Ramirez J, Bolduc N, Lisch D, Hake S (2009). Distal expression of knotted1 in maize leaves leads to reestablishment of proximal/distal patterning and leaf dissection.Plant Physiol 151, 1878-1888. |
| 61 | Reymond M, Muller B, Tardieu F (2004). Dealing with the genotype×environment interaction via a modelling approach: a comparison of QTLs of maize leaf length or width with QTLs of model parameters.J Exp Bot 55, 2461-2472. |
| 62 | Roodbarkelari F, Du F, Truernit E, Laux T (2015). ZLL/ AGO10 maintains shoot meristem stem cells during Arabidopsis embryogenesis by down-regulating ARF2- mediated auxin response.BMC Biol 13, 74. |
| 63 | Scanlon MJ, Chen KD, McKnight CC (2000). The narrow sheath duplicate genes: sectors of dual aneuploidy reveal ancestrally conserved gene functions during maize leaf development. Genetics 155, 1379-1389. |
| 64 | Schatlowski N, Stahl Y, Hohenstatt ML, Goodrich J, Schubert D (2010). The CURLY LEAF interacting protein BLISTER controls expression of polycomb-group target genes and cellular differentiation of Arabidopsis thaliana. Plant Cell 22, 2291-2305. |
| 65 | Sentoku N, Sato Y, Matsuoka M (2000). Overexpression of rice OSH genes induces ectopic shoots on leaf sheaths of transgenic rice plants. Dev Biol 220, 358-364. |
| 66 | Tian F, Bradbury PJ, Brown PJ, Hung H, Sun Q, Flint-Garcia S, Rocheford TR, McMullen MD, Holland JB, Buckler ES (2011). Genome-wide association study of leaf architecture in the maize nested association mapping population.Nat Genet 43, 159-162. |
| 67 | Timmermans MC, Schultes NP, Jankovsky JP, Nelson T (1998). Leaf bladeless 1 is required for dorsoventrality of lateral organs in maize. Development 125, 2813-2823. |
| 68 | Toriba T, Harada K, Takamura A, Nakamura H, Ichikawa H, Suzaki T, Hirano HY (2007). Molecular characterization the YABBY gene family in Oryza sativa and expression analysis of OsYABBY1. Mol Genet Genom 277, 457-468. |
| 69 | Vercruyssen L, Verkest A, Gonzalez N, Heyndrickx KS, Eeckhout D, Han SK, Jégu T, Archacki R, Van Leene J, Andriankaja M, De Bodt S, Abeel T, Coppens F, Dhondt S, De Milde L, Vermeersch M, Maleux K, Gevaert K, Jerzmanowski A, Benhamed M, Wagner D, Vandepoele K, De Jaeger G, Inzé D (2014). ANGUSTIFOLIA3 binds to SWI/SNF chromatin remodeling complexes to regulate transcription during Arabidopsis leaf development.Plant Cell 26, 210-229. |
| 70 | Wassom JJ (2013). Quantitative trait loci for leaf angle, leaf width, leaf length, and plant height in a maize (Zea mays L.) B73 × Mo17 population. Maydica 58, 318-321. |
| 71 | Wei XM, Wang XB, Guo SL, Zhou JL, Shi Y, Wang HT, Dou DD, Song XH, Li GH, Ku LX, Chen YH (2016). Epistatic and QTL × environment interaction effects on leaf area-associated traits in maize.Plant Breed 135, 671-676. |
| 72 | Woo YM, Park HJ, Su’udi M, Yang JI, Park JJ, Back K, Park YM, An G (2007). Constitutively wilted 1, a member of the rice YUCCA gene family, is required for maintaining water homeostasis and an appropriate root to shoot ratio. Plant Mol Biol 65, 125-136. |
| 73 | Wu L, Zhang DF, Xue M, Qian JJ, He Y, Wang SC (2014). Overexpression of the maize GRF10, an endogenous truncated growth-regulating factor protein, leads to reduction in leaf size and plant height.J Integr Plant Biol 56, 1053-1063. |
| 74 | Würschum T, Groß-Hardt R, Laux T (2006). APETALA2 regulates the stem cell niche in the Arabidopsis shoot meristem. Plant Cell 18, 295-307. |
| 75 | Yoshikawa T, Eiguchi M, Hibara KI, Ito JI, Nagato Y (2013). Rice SLENDER LEAF 1 gene encodes cellulose synthase-like D4 and is specifically expressed in M-phase cells to regulate cell proliferation. J Exp Bot 64, 2049-2061. |
| 76 | Zhang DF, Li B, Jia GQ, Zhang TF, Dai JR, Li JS, Wang SC (2008). Isolation and characterization of genes encoding GRF transcription factors and GIF transcriptional coactivators in Maize (Zea mays L.). Plant Sci 175, 809-817. |
| 77 | Zhang GH, Xu Q, Zhu XD, Qian Q, Xue HW (2009). SHALLOT-LIKE1 is a KANADI transcription factor that modulates rice leaf rolling by regulating leaf abaxial cell development.Plant Cell 21, 719-735. |
| 1 | 安允权, 张君, 席章营, 李明娜, 李沛, 王顺喜, 张莹莹, 陈彦惠, 吴连成 (2016). 玉米不同叶位叶面积的QTL定位. 分子植物育种 14, 2113-2120. |
| 2 | 常立国, 何坤辉, 刘建超, 薛吉全 (2016). 不同环境条件下玉米叶夹角的QTL定位. 玉米科学 24(4), 49-55. |
| 3 | 郭莹 (2012). 利用不同F2群体定位玉米株型性状的QTL. 硕士论文. 重庆: 西南大学. pp. 45-47. |
| 4 | 鞠培娜, 方云霞, 邹国兴, 彭友林, 孙川, 胡江, 董国军, 曾大力, 郭龙彪, 张光恒, 高振宇, 钱前 (2010). 一个新的水稻叶形突变体(tll1)的遗传分析与精细定位. 植物学报 45, 654-661. |
| 5 | 李贤唐, 丁俊强, 王瑞霞, 吴建宇 (2011). 玉米株型相关性状的QTL定位与分析. 江苏农业科学 39(2), 21-25. |
| 6 | 刘建超, 褚群, 蔡红光, 米国华, 陈范骏 (2010). 玉米SSR连锁图谱构建及叶面积的QTL定位. 遗传 32, 625-631. |
| 7 | 刘正, 余婷婷, 梅秀鹏, 陈淅宁, 王国强, 王久光, 刘朝显, 王旭, 蔡一林 (2014). 玉米穗上叶夹角和叶间距的QTL定位. 农业生物技术学报 22, 177-187. |
| 8 | 路明, 周芳, 谢传晓, 李明顺, 徐云碧, Warburton M, 张世煌 (2007). 玉米杂交种掖单13号的SSR连锁图谱构建与叶夹角和叶向值的QTL定位与分析. 遗传 29, 1131-1138. |
| 9 | 唐登国 (2013). 玉米叶宽和叶长性状的QTL定位与分析. 硕士论文. 雅安: 四川农业大学. pp. 29-43. |
| 10 | 于永涛, 张吉民, 石云素, 宋燕春, 王天宇, 黎裕 (2006). 利用不同群体对玉米株高和叶片夹角的QTL分析. 玉米科学 14(2), 88-92. |
| 11 | 袁园园, 王丽, 赵盼盼, 王林嵩 (2016). 棉花类结瘤素MtN21基因家族生物信息学分析. 植物学报 51, 515-524. |
| 12 | 张姿丽, 蒋锋, 刘鹏飞, 陈青春, 张媛, 王晓明 (2014a). 甜玉米穗位叶面积QTL定位. 湖北农业科学 53, 1502-1505. |
| 13 | 张姿丽, 刘鹏飞, 蒋锋, 陈青春, 张媛, 王晓明, 王汉宁 (2014b). 基于四交群体的玉米叶夹角和叶向值QTL定位分析. 中国农业大学学报 19(4), 7-16. |
| 14 | 张志腾 (2015). 玉米叶型相关性状QTL定位与分析. 硕士论文. 雅安: 四川农业大学. pp. 21-23. |
| 15 | 郑祖平, 黄玉碧, 田孟良, 谭振波 (2007). 不同供氮水平下玉米株型相关性状的QTLs定位和上位性效应分析. 玉米科学 15(2), 14-18. |
| 16 | Agrama HAS, Zakaria AG, Said FB, Tuinstra M (1999). Identification of quantitative trait loci for nitrogen use efficiency in maize.Mol Breed 5, 187-195. |
| 17 | Cai HG, Chu Q, Yuan LX, Liu JC, Chen XH, Chen FJ, Mi GH, Zhang FS (2012). Identification of quantitative trait loci for leaf area and chlorophyll content in maize ( Zea mays) under low nitrogen and low phosphorus supply. Mol Breed 30, 251-266. |
| 18 | Candaele J, Demuynck K, Mosoti D, Beemster GTS, Inzé D, Nelissen H (2014). Differential methylation during maize leaf growth targets developmentally regulated ge- nes.Plant Physiol 164, 1350-1364. |
| 19 | Candela H, Johnston R, Gerhold A, Foster T, Hake S (2008). The Milkweed pod1 gene encodes a KANADI protein that is required for abaxial/adaxial patterning in maize leaves. Plant Cell 20, 2073-2087. |
| 20 | Darvasi A, Soller M (1997). A simple method to calculate resolving power and confidence interval of QTL map location.Behav Genet 27, 125-132. |
| 21 | Debernardi JM, Mecchia MA, Vercruyssen L, Smaczniak C, Kaufmann K, Inze D, Rodriguez RE, Palatnik JF (2014). Post-transcriptional control of GRF transcription factors by microRNA miR396 and GIF co-activator affects leaf size and longevity.Plant J 79, 413-426. |
| 22 | Dewitte W, Scofield S, Alcasabas AA, Maughan SC, Menges M, Braun N, Collins C, Nieuwland J, Prinsen E, Sundaresan V, Murray JAH (2007). Arabidopsis CYCD3 D-type cyclins link cell proliferation and endocycles and are rate-limiting for cytokinin responses.Proc Natl Acad Sci USA 104, 14537-14542. |
| 23 | Ding ZQ, Lin ZF, Li Q, Wu H, Xiang CY, Wang JF (2015). DNL1, encodes cellulose synthase-like D4, is a major QTL for plant height and leaf width in rice(Oryza sativa L.). Biochem Biophys Res Commun 457, 133-140. |
| 24 | Dotto MC, Petsch KA, Aukerman MJ, Beatty M, Hammell M, Timmermans MC (2014). Genome-wide analysis of leaf bladeless1-regulated and phased small RNAs underscores the importance of the TAS3 ta-siRNA pathway to maize development. PLoS Genet 10, e1004826. |
| 25 | Eloy NB, de Freitas Lima M, Van Damme D, Vanhaeren H, Gonzalez N, de Milde L, Hemerly AS, Beemster GTS, Inzé D, Ferreira PCG (2011). The APC/C subunit 10 plays an essential role in cell proliferation during leaf development. Plant J 68, 351-363. |
| 26 | Facette MR, Shen ZX, Björnsdóttir FR, Briggs SP, Smith LG (2013). Parallel proteomic and phosphoproteomic ana- lyses of successive stages of maize leaf development.Plant Cell 25, 2798-2812. |
| 27 | Fujino K, Matsuda Y, Ozawa K, Nishimura T, Koshiba T, Fraaije MW, Sekiguchi H (2008). NARROW LEAF 7 controls leaf shape mediated by auxin in rice. Mol Genet Genom 279, 499-507. |
| 28 | Goffinet B, Gerber S (2000). Quantitative trait loci: a meta- analysis.Genetics 155, 463-473. |
| 29 | Guo SL, Ku LX, Qi JS, Tian ZQ, Han T, Zhang LK, Su HH, Ren ZZ, Chen YH (2015). Genetic analysis and major quantitative trait locus mapping of leaf widths at different positions in multiple populations.PLoS One 10, e0119095. |
| 30 | Horiguchi G, Ferjani A, Fujikura U, Tsukaya H (2006). Coordination of cell proliferation and cell expansion in the control of leaf size in Arabidopsis thaliana. J Plant Res 119, 37-42. |
| 31 | Hu J, Zhu L, Zeng DL, Gao ZY, Guo LB, Fang YX, Zhang GH, Dong GJ, Yan MX, Liu J, Qian Q (2010). Identification and characterization of NARROW AND ROLLED LEAF 1, a novel gene regulating leaf morphology and plant architecture in rice. Plant Mol Biol 73, 283-292. |
| 32 | Hunter C, Willmann MR, Wu G, Yoshikawa M, de la Luz Gutiérrez-Nava M, Poethig SR (2006). Trans-acting siRNA-mediated repression of ETTIN and ARF4 regulates heteroblasty in Arabidopsis.Development 133, 2973-2981. |
| 33 | Iwakawa H, Iwasaki M, Kojima S, Ueno Y, Soma T, Tanaka H, Semiarti E, Machida Y, Machida C (2007). Expression of the ASYMMETRIC LEAVES 2 gene in the adaxial domain of Arabidopsis leaves represses cell proliferation in this domain and is critical for the development of properly expanded leaves. Plant J 51, 173-184. |
| 34 | Jun SE, Cho KH, Hwang JY, Abdel-Fattah W, Hammermeister A, Schaffrath R, Bowman JL, Kim GT (2015). Comparative analysis of the conserved functions of Arabidopsis DRL1 and yeast KTI12.Mol Cells 38, 243-250. |
| 35 | Kelley DR, Arreola A, Gallagher TL, Gasser CS (2012). ETTIN (ARF3) physically interacts with KANADI proteins to form a functional complex essential for integument deve- lopment and polarity determination in Arabidopsis.Deve- lopment 139, 1105-1109. |
| 36 | Kim GT, Tsukaya H, Uchimiya H (1998). The ROTUNDIFOLIA3 gene of Arabidopsis thaliana encodes a new member of the cytochrome P450 family that is required for the regulated polar elongation of leaf cells. Genes Dev 12, 2381-2391. |
| 37 | Kim JH, Choi D, Kende H (2003). The AtGRF family of putative transcription factors is involved in leaf and cotyledon growth in Arabidopsis.Plant J 36, 94-104. |
| 38 | Ku LX, Zhang J, Guo SL, Liu HY, Zhao RF, Chen YH (2012a). Integrated multiple population analysis of leaf architecture traits in maize (Zea mays L.). J Exp Bot 63, 261-274. |
| 39 | Ku LX, Zhang J, Zhang JC, Guo SL, Liu HY, Zhao RF, Yan QX, Chen YH (2012b). Genetic dissection of leaf area by jointing two F2:3 populations in maize (Zea mays L.). Plant Breed 131, 591-599. |
| 78 | Zhu HL, Hu FQ, Wang RH, Zhou X, Sze SH, Liou LW, Barefoot A, Dickman M, Zhang XR (2011). Arabidopsis Argonaute10 specifically sequesters miR166/165 to regulate shoot apical meristem development.Cell 145, 242-256. |
| 40 | Ku LX, Zhao WM, Zhang J, Wu LC, Wang CL, Wang PA, Zhang WQ, Chen YH (2010). Quantitative trait loci mapping of leaf angle and leaf orientation value in maize (Zea mays L.). Theor Appl Genet 121, 951-959. |
| 41 | Kubo FC, Yasui Y, Kumamaru T, Sato Y, Hirano HY (2017). Genetic analysis of rice mutants responsible for narrow leaf phenotype and reduced vein number.Genes Genet Syst 91, 235-240. |
| [1] | DENG Bei, WANG Xiao-Feng, LIAO Jun. Ecophysiological responses of herbaceous and woody plants to environmental stresses in the riparian zone of Three Gorges Reservoir: a meta-analysis [J]. Chin J Plant Ecol, 2024, 48(5): 623-637. |
| [2] | GENG Xue-Qi, TANG Ya-Kun, WANG Li-Na, DENG Xu, ZHANG Ze-Ling, ZHOU Ying. Nitrogen addition increases biomass but reduces nitrogen use efficiency of terrestrial plants in China [J]. Chin J Plant Ecol, 2024, 48(2): 147-157. |
| [3] | Chaoyu Zhu, Chengxiang Hu, Zhenan Zhu, Zhining Zhang, Lihai Wang, Jun Chen, Sanfeng Li, Jinjin Lian, Luyao Tang, Qianqian Zhong, Wenjing Yin, Yuexing Wang, Yuchun Rao. Mapping of QTLs Associated with Rice Panicle Traits and Candidate Gene Analysis [J]. Chinese Bulletin of Botany, 2024, 59(2): 217-230. |
| [4] | CHEN Xiang-Lei, CUI Shu-Juan, ZHAO Chen-Jun, GU Hong-Liang, CHEN Xiao-Ping, LI Jin-Long, SUN Jun. Predictive model of single leaf area for woody plants based on leaf morphology classification [J]. Chin J Plant Ecol, 2024, 48(12): 1683-1691. |
| [5] | Bao Zhu, Jiangzhe Zhao, Kewei Zhang, Peng Huang. OsCKX9 is Involved in Regulating the Rice Lamina Joint Development and Leaf Angle [J]. Chinese Bulletin of Botany, 2024, 59(1): 10-21. |
| [6] | LI Na, TANG Shi-Ming, GUO Jian-Ying, TIAN Ru, WANG Shan, HU Bing, LUO Yong-Hong, XU Zhu-Wen. Meta-analysis of effects of grazing on plant community properties in Nei Mongol grassland [J]. Chin J Plant Ecol, 2023, 47(9): 1256-1269. |
| [7] | Qiwei Jia, Qianqian Zhong, Yujia Gu, Tianqi Lu, Wei Li, Shuai Yang, Chaoyu Zhu, Chengxiang Hu, Sanfeng Li, Yuexing Wang, Yuchun Rao. Mapping of QTL for Cell Wall Related Components in Rice Stem and Analysis of Candidate Genes [J]. Chinese Bulletin of Botany, 2023, 58(6): 882-892. |
| [8] | Jiayi Jin, Yiting Luo, Huimin Yang, Tao Lu, Hanfei Ye, Jiyi Xie, Kexin Wang, Qianyu Chen, Yuan Fang, Yuexing Wang, Yuchun Rao. QTL Mapping and Expression Analysis on Candidate Genes Related to Chlorophyll Content in Rice [J]. Chinese Bulletin of Botany, 2023, 58(3): 394-403. |
| [9] | Jiesheng Rao, Tao Yang, Xi Tian, Wencong Liu, Xiaofeng Wang, Hengjun Qian, Zehao Shen. Vertical structural characteristics of a semi-humid evergreen broad-leaved forest and common tree species based on a portable backpack LiDAR [J]. Biodiv Sci, 2023, 31(11): 23216-. |
| [10] | Yanyi Wang, Yimei Zhang, Canwei Xia, Anders Pape Møller. A meta-analysis of the effects in alpha acoustic indices [J]. Biodiv Sci, 2023, 31(1): 22369-. |
| [11] | WANG Guang-Ya, CHEN Bing-Hua, HUANG Yu-Chen, JIN Guang-Ze, LIU Zhi-Li. Effects of growing position on leaflet trait variations and its correlations in Fraxinus mandshurica [J]. Chin J Plant Ecol, 2022, 46(6): 712-721. |
| [12] | Wei Heping, Lu Tao, Jia Qiwei, Deng Fei, Zhu Hao, Qi Zehua, Wang Yuxi, Ye Hanfei, Yin Wenjing, Fang Yuan, Mu Dan, Rao Yuchun. QTL Mapping of Candidate Genes for Heading Date in Rice [J]. Chinese Bulletin of Botany, 2022, 57(5): 588-595. |
| [13] | Hanfei Ye, Wenjing Yin, Yian Guan, Kairu Yang, Qianyu Chen, Shuying Yu, Xudong Zhu, Dedong Xin, Wei Zhang, Yuexing Wang, Yuchun Rao. QTL Mapping and Candidate Gene Analysis of Vitamin E in Rice Grain [J]. Chinese Bulletin of Botany, 2022, 57(2): 157-170. |
| [14] | ZHENG Zhou-Tao, ZHANG Yang-Jian. Variation in ecosystem water use efficiency and its attribution analysis during 1982-2018 in Qingzang Plateau [J]. Chin J Plant Ecol, 2022, 46(12): 1486-1496. |
| [15] | LIU Bing-Bing, WEI Jian-Xin, HU Tian-Yu, YANG Qiu-Li, LIU Xiao-Qiang, WU Fa-Yun, SU Yan-Jun, GUO Qing-Hua. Validation and uncertainty analysis of satellite remote sensing products for monitoring China’s forest ecosystems—Based on massive UAV LiDAR data [J]. Chin J Plant Ecol, 2022, 46(10): 1305-1316. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||