

Chinese Bulletin of Botany ›› 2021, Vol. 56 ›› Issue (3): 330-338.DOI: 10.11983/CBB21057 cstr: 32102.14.CBB21057
Previous Articles Next Articles
Binbin Hu1,2, Zhihui Xue1, Cui Zhang1,2,*( )
)
Received:2020-04-02
Accepted:2021-05-07
Online:2021-05-01
Published:2021-05-07
Contact:
Cui Zhang
Binbin Hu, Zhihui Xue, Cui Zhang. Protocols for Small RNA FISH in Plants[J]. Chinese Bulletin of Botany, 2021, 56(3): 330-338.
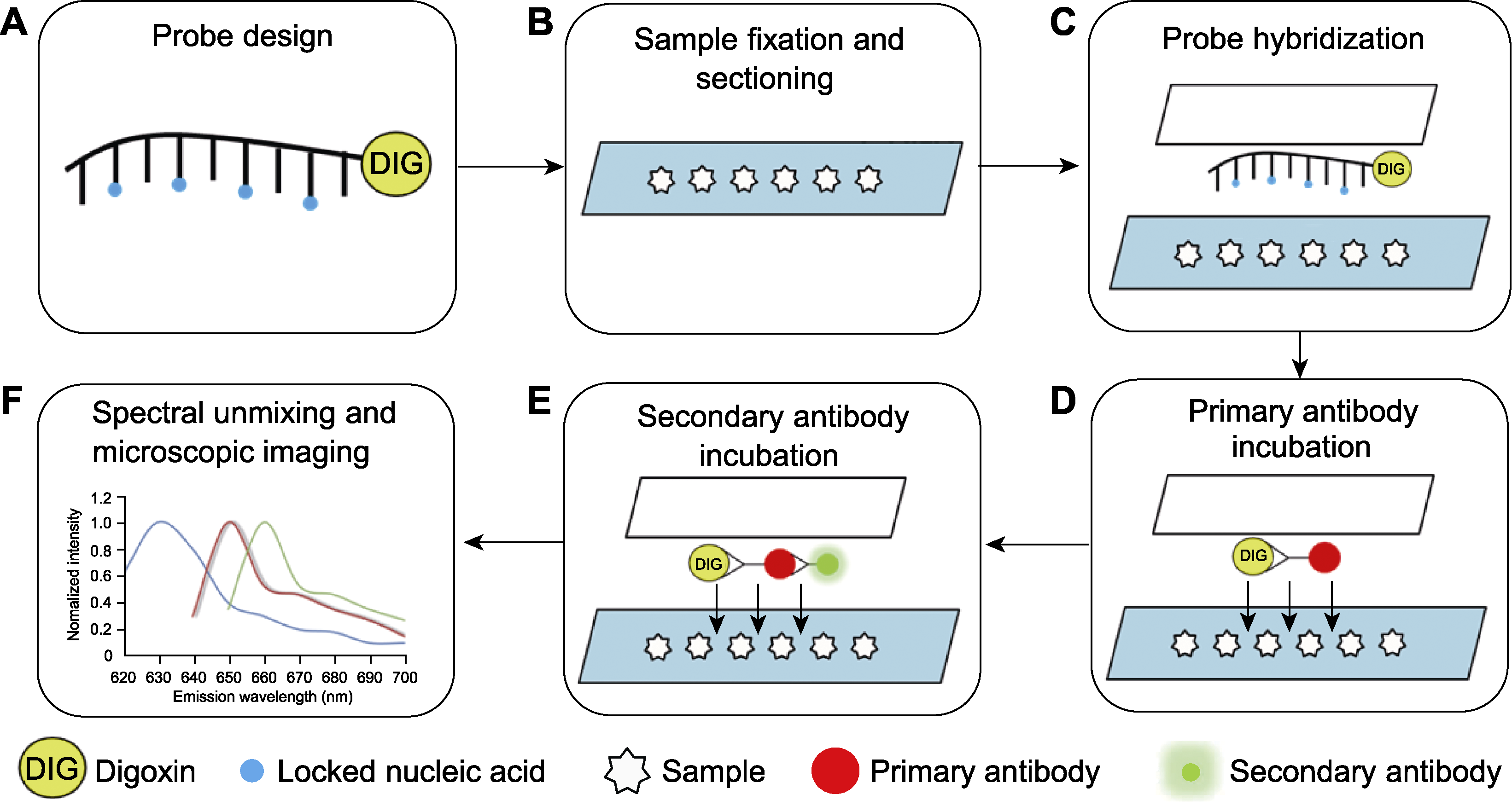
Figure 1 Outlines of small RNA fluorescent in situ hybridization (A) Probe design; (B) Paraffin sectioning; (C) Probe hybridization; (D) Primary antibody incubation; (E) Secondary antibody incubation; (F) Spectral unmixing and imaging by confocal microscope
| 1 |
Baumberger N, Baulcombe DC (2005). Arabidopsis ARGONAUTE1 is an RNA slicer that selectively recruits microRNAs and short interfering RNAs. Proc Natl Acad Sci USA 102, 11928-11933.
DOI URL |
| 2 |
Bhogale S, Mahajan AS, Natarajan B, Rajabhoj M, Thulasiram HV, Banerjee AK (2014). MicroRNA156: a potential graft-transmissible microRNA that modulates plant architecture and tuberization in Solanum tuberosum ssp. andigena. Plant Physiol 164, 1011-1027.
DOI URL |
| 3 |
Bleckmann A, Dresselhaus T (2016). Fluorescent whole- mount RNA in situ hybridization (F-WISH) in plant germ cells and the fertilized ovule. Methods 98, 66-73.
DOI PMID |
| 4 |
Bruno L, Muto A, Spadafora ND, Iaria D, Chiappetta A, Van Lijsebettens M, Bitonti MB (2011). Multi-probe in situ hybridization to whole mount Arabidopsis seedlings. Int J Dev Biol 55, 197-203.
DOI URL |
| 5 |
Bruno L, Ronchini M, Gagliardi O, Corinti T, Chiappetta A, Gerola P, Bitonti MB (2015). Analysis of ATGUS1 and ATGUS2 in Arabidopsis root apex by a highly sensitive TSA-MISH method. Int J Dev Biol 59, 221-228.
DOI URL |
| 6 |
Buhtz A, Springer F, Chappell L, Baulcombe DC, Kehr J (2008). Identification and characterization of small RNAs from the phloem of Brassica napus. Plant J 53, 739-749.
DOI URL |
| 7 |
Carlsbecker A, Lee JY, Roberts CJ, Dettmer J, Lehesranta S, Zhou J, Lindgren O, Moreno-Risueno MA, Vatén A, Thitamadee S, Campilho A, Sebastian J, Bowman JL, Helariutta Y, Benfey PN (2010). Cell signaling by microRNA165/6 directs gene dose-dependent root cell fate. Nature 465, 316-321.
DOI PMID |
| 8 |
Chen XM (2004). A microRNA as a translational repressor of APETALA2 in Arabidopsis flower development. Science 303, 2022-2025.
DOI URL |
| 9 |
Chen XM (2009). Small RNAs and their roles in plant development. Annu Rev Cell Dev Biol 25, 21-44.
DOI URL |
| 10 | Chitwood DH, Nogueira FT, Howell MD, Montogmery TA, Carrington JC, Timmermans MC (2008). Pattern formation in leaves via small RNA mobility. Dev Biol 319, 587-588. |
| 11 |
Chitwood DH, Nogueira FTS, Howell MD, Montgomery TA, Carrington JC, Timmermans MCP (2009). Pattern formation via small RNA mobility. Gene Dev 23, 549-554.
DOI URL |
| 12 | Fischer AH, Jacobson KA, Rose J, Zeller R (2008). Paraffin embedding tissue samples for sectioning. CSH Protoc 2008, pdb.prot4989. |
| 13 |
Himanen K, Woloszynska M, Boccardi TM, De Groeve S, Nelissen H, Bruno L, Vuylsteke M, Van Lijsebettens M (2012). Histone H2b monoubiquitination is required to reach maximal transcript levels of circadian clock genes in Arabidopsis. Plant J 72, 249-260.
DOI URL |
| 14 |
Howell MD, Fahlgren N, Chapman EJ, Cumbie JS, Sullivan CM, Givan SA, Kasschau KD, Carrington JC (2007). Genome-wide analysis of the RNA-DEPENDENT RNA POLYMERASE6/DICER-LIKE4 pathway in Arabidop- sis reveals dependency on miRNA- and tasiRNA-directed targeting. Plant Cell 19, 926-942.
PMID |
| 15 |
Huang K, Baldrich P, Meyers BC, Caplan JL (2019). sRNA-FISH: versatile fluorescent in situ detection of small RNAs in plants. Plant J 98, 359-369.
DOI |
| 16 |
Huen AK, Rodriguez-Medina C, Ho AYY, Atkins CA, Smith PMC (2017). Long-distance movement of phospha- te starvation-responsive microRNAs in Arabidopsis. Plant Biol 19, 643-649.
DOI URL |
| 17 |
Javelle M, Timmermans MCP (2012). In situ localization of small RNAs in plants by using LNA probes. Nat Protoc 7, 533-541.
DOI PMID |
| 18 | Jiang JM (2019). Fluorescence in situ hybridization in plants: recent developments and future applications. Chromoso- me Res 27, 153-165. |
| 19 |
Jones-Rhoades MW, Bartel DP, Bartel B (2006). Micro- RNAs and their regulatory roles in plants. Annu Rev Plant Biol 57, 19-53.
PMID |
| 20 |
Jung JH, Park CM (2007). MIR166/ 165 genes exhibit dynamic expression patterns in regulating shoot apical meris- tem and floral development in Arabidopsis. Planta 225, 1327-1338.
DOI URL |
| 21 | Kidner C, Timmermans M (2006). In situ hybridization as a tool to study the role of microRNAs in plant development. Methods Mol Biol 342, 159-179. |
| 22 | Kim VN, Han JJ, Siomi MC (2009). Biogenesis of small RNAs in animals. Nat Rev Mol Cell Biol 10, 126-139. |
| 23 |
Knauer S, Holt AL, Rubio-Somoza I, Tucker EJ, Hinze A, Pisch M, Javelle M, Timmermans MC, Tucker MR, Laux T (2013). A protodermal miR394 signal defines a region of stem cell competence in the Arabidopsis shoot meristem. Dev Cell 24, 125-132.
DOI URL |
| 24 |
Kurihara Y, Watanabe Y (2004). Arabidopsis micro-RNA biogenesis through dicer-like 1 protein functions. Proc Natl Acad Sci USA 101, 12753-12758.
DOI URL |
| 25 |
Lamb JC, Danilova T, Bauer MJ, Meyer JM, Holland JJ, Jensen MD, Birchler JA (2007). Single-gene detection and karyotyping using small-target fluorescence in situ hybridization on maize somatic chromosomes. Genetics 175, 1047-1058.
DOI URL |
| 26 |
Lee Y, Kim M, Han JJ, Yeom KH, Lee S, Baek SH, Kim VN (2004). MicroRNA genes are transcribed by RNA polymerase II. EMBO J 23, 4051-4060.
DOI URL |
| 27 |
Levsky JM, Singer RH (2003). Fluorescence in situ hybridization: past, present and future. J Cell Sci 116, 2833-2838.
DOI URL |
| 28 |
Li JJ, Yang ZY, Yu B, Liu J, Chen XM (2005). Methylation protects miRNAs and siRNAs from a 3'-end uridylation activity in Arabidopsis. Curr Biol 15, 1501-1507.
DOI URL |
| 29 |
Li S, Wang XT, Xu WY, Liu T, Cai CM, Chen LY, Clark CB, Ma JX (2021). Unidirectional movement of small RNAs from shoots to roots in interspecific heterografts. Nat Plants 7, 50-59.
DOI URL |
| 30 |
Lin SI, Chiang SF, Lin WY, Chen JW, Tseng CY, Wu PC, Chiou TJ (2008). Regulatory network of microRNA399 and PHO2 by systemic signaling. Plant Physiol 147, 732-746.
DOI URL |
| 31 |
Liu QL, Yao XZ, Pi LM, Wang H, Cui XF, Huang H (2009). The ARGONAUTE10 gene modulates shoot apical meris- tem maintenance and establishment of leaf polarity by repressing miR165/166 in Arabidopsis. Plant J 58, 27-40.
DOI URL |
| 32 |
Lu J, Tsourkas A (2009). Imaging individual microRNAs in single mammalian cells in situ. Nucleic Acids Res 37, e100.
DOI URL |
| 33 |
Moissiard G, Parizotto EA, Himber C, Voinnet O (2007). Transitivity in Arabidopsis can be primed, requires the redundant action of the antiviral Dicer-like 4 and Dicer-like 2, and is compromised by viral-encoded suppressor proteins. RNA 13, 1268-1278.
DOI URL |
| 34 |
Nodine MD, Bartel DP (2010). MicroRNAs prevent precocious gene expression and enable pattern formation during plant embryogenesis. Gene Dev 24, 2678-2692.
DOI URL |
| 35 |
Nogueira FTS, Chitwood DH, Madi S, Ohtsu K, Schnable PS, Scanlon MJ, Timmermans MCP (2009). Regulation of small RNA accumulation in the maize shoot apex. PLoS Genet 5, e1000320.
DOI URL |
| 36 |
Nogueira FTS, Madi S, Chitwood DH, Juarez MT, Timmer- mans MCP (2007). Two small regulatory RNAs establish opposing fates of a developmental axis. Gene Dev 21, 750-755.
PMID |
| 37 |
Ori N, Cohen AR, Etzioni A, Brand A, Yanai O, Shleizer S, Menda N, Amsellem Z, Efroni I, Pekker I, Alvarez JP, Blum E, Zamir D, Eshed Y (2007). Regulation of LANCEOLATE by miR319 is required for compound-leaf development in tomato. Nat Genet 39, 787-791.
DOI URL |
| 38 |
Pagliarani C, Gambino G (2019). Small RNA mobility: spread of RNA silencing effectors and its effect on developmental processes and stress adaptation in plants. Int J Mol Sci 20, 4306.
DOI URL |
| 39 |
Pant BD, Buhtz A, Kehr J, Scheible WR (2008). MicroRNA399 is a long-distance signal for the regulation of plant phosphate homeostasis. Plant J 53, 731-738.
DOI URL |
| 40 |
Pant BD, Musialak-Lange M, Nuc P, May P, Buhtz A, Kehr J, Walther D, Scheible WR (2009). Identification of nutrient-responsive Arabidopsis and rapeseed microRNAs by comprehensive real-time polymerase chain reaction profiling and small RNA sequencing. Plant Physiol 150, 1541-1555.
DOI URL |
| 41 |
Parizotto EA, Dunoyer P, Rahm N, Himber C, Voinnet O (2004). In vivo investigation of the transcription, processing, endonucleolytic activity, and functional relevance of the spatial distribution of a plant miRNA. Gene Dev 18, 2237-2242.
PMID |
| 42 |
Park MY, Wu G, Gonzalez-Sulser A, Vaucheret H, Poethig RS (2005). Nuclear processing and export of microRNAs in Arabidopsis. Proc Natl Acad Sci USA 102, 3691-3696.
DOI URL |
| 43 |
Raman S, Greb T, Peaucelle A, Blein T, Laufs P, Theres K (2008). Interplay of miR164, CUP-SHAPED COTYLEDON genes and LATERAL SUPPRESSOR controls axillary meristem formation in Arabidopsis thaliana. Plant J 55, 65-76.
DOI URL |
| 44 |
Rozier F, Mirabet V, Vernoux T, Das P (2014). Analysis of 3D gene expression patterns in plants using whole-mount RNA in situ hybridization. Nat Protoc 9, 2464-2475.
DOI URL |
| 45 |
Skopelitis DS, Hill K, Klesen S, Marco CF, von Born P, Chitwood DH, Timmermans MCP (2018). Gating of miRNA movement at defined cell-cell interfaces governs their impact as positional signals. Nat Commun 9, 3107.
DOI PMID |
| 46 |
Tirichine L, Andrey P, Biot E, Maurin Y, Gaudin V (2009). 3D fluorescent in situ hybridization using Arabidopsis leaf cryosections and isolated nuclei. Plant Methods 5, 11.
DOI PMID |
| 47 |
Vaucheret H, Vazquez F, Crété P, Bartel DP (2004). The action of ARGONAUTE1 in the miRNA pathway and its regulation by the miRNA pathway are crucial for plant development. Gene Dev 18, 1187-1197.
PMID |
| 48 |
Wollmann H, Mica E, Todesco M, Long JA, Weigel D (2010). On reconciling the interactions between APETA- LA2, miR172 and AGAMOUS with the ABC model of flower development. Development 137, 3633-3642.
DOI PMID |
| 49 |
Woloszynska M, Le Gall S, Neyt P, Boccardi TM, Grasser M, Längst G, Aesaert S, Coussens G, Dhondt S, Van De Slijke E, Bruno L, Fung-Uceda J, Mas P, Van Montagu M, Inzé D, Himanen K, De Jaeger G, Grasser KD, Van Lijsebettens M (2019). Histone 2B monoubiquitination complex integrates transcript elongation with RNA processing at circadian clock and flowering regulators. Proc Natl Acad Sci USA 116, 8060-8069.
DOI URL |
| 50 |
Xue ZH, Liu LY, Zhang C (2020). Regulation of shoot apical meristem and axillary meristem development in plants. Int J Mol Sci 21, 2917.
DOI URL |
| 51 |
Yang WB, Schuster C, Prunet N, Dong QK, Landrein B, Wightman R, Meyerowitz EM (2020). Visualization of protein coding, long noncoding, and nuclear RNAs by fluorescence in situ hybridization in sections of shoot apical meristems and developing flowers. Plant Physiol 182, 147-158.
DOI URL |
| 52 |
Yang WB, Wightman R, Meyerowitz EM (2017). Cell cycle control by nuclear sequestration of CDC20 and CDH1 mRNA in plant stem cells. Mol Cell 68, 1108-1119.
DOI URL |
| 53 |
Yu Y, Zhang YC, Chen XM, Chen YQ (2019). Plant noncoding RNAs: hidden players in development and stress responses. Annu Rev Cell Dev Biol 35, 407-431.
DOI |
| 54 |
Zhang C, Fan LS, Le BH, Ye PY, Mo BX, Chen XM (2020). Regulation of ARGONAUTE10 expression enables temporal and spatial precision in axillary meristem initiation in Arabidopsis. Dev Cell 55, 603-616.
DOI URL |
| 55 | Zhang C, Wang J, Wenkel S, Chandler JW, Werr W, Jiao YL (2018). Spatiotemporal control of axillary meristem formation by interacting transcriptional regulators. Development 145, dev158352. |
| [1] | Lu Zi-Jia, Wang Tian-Rui, Zheng Si-Si, Cao Jian-Guo, Kozlowski Gregor, Song Yi-Gang. Environmental adaptive genetic variation and genetic vulnerability of relict plant Pterocarya hupehensis [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [2] | ZHANG Zi-Rui, Zhou Jing, HU Yan-Ping, Liang Shuang, MA Yong-Peng, CHEN Wei-Le. Root-associated Fungal Communities of the Critically Endangered Plant Pinus Squamata [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [3] | JIANG Xiao-Yu, YU Xin-Miao, LIAO Qin, ZHANG Jin-Wei, WU Xue-Feng, WANG Xu, PAN Jun-Tong, WANG Jun-Feng, MU Chun-Sheng, SHI Yu-Jie. Studies on the emission of nitrous oxide from terrestrial plants [J]. Chin J Plant Ecol, 2025, 49(4): 513-525. |
| [4] | Tao Xie, Yifan Zhang, Yunhui Liu, Huiyu You, Jibenben Xia, Rong Ma, Chunni Zhang, Xuejun Hua. Research progress of iron-sulfur cluster synthesis system and regulation in plant mitochondria [J]. Chinese Bulletin of Botany, 2025, 60(4): 1-0. |
| [5] | Gan Xie, Jing Xuan, Qidi Fu, Ze Wei, Kai Xue, Hairui Luo, Jixi Gao, Min Li. Establishing an intelligent identification model for unmanned aerial vehicle surveys of grassland plant diversity [J]. Biodiv Sci, 2025, 33(4): 24236-. |
| [6] | Xiong Lianglin, Liang Guolu, Guo Qigao, Jing Danlong. Advances in the Regulation of Alternative Splicing of Genes in Plants in Response to Abiotic Stress [J]. Chinese Bulletin of Botany, 2025, 60(3): 435-448. |
| [7] | Zhang Chan, Zhao Suya, Zhang Xinran, Wang Yifan, Wang Linlin. Impacts of alien pollinators on native plant‒pollinator interactions [J]. Biodiv Sci, 2025, 33(2): 24443-. |
| [8] | Jing Xuan, Qidi Fu, Gan Xie, Kai Xue, Hairui Luo, Ze Wei, Mingyue Zhao, Liang Zhi, Huawei Wan, Jixi Gao, Min Li. An Artificial Intelligence Model for Identifying Grassland Plants in Northern China [J]. Chinese Bulletin of Botany, 2025, 60(1): 74-80. |
| [9] | Yan Deng, Limin Lu, Qiang Zhang, Zhiduan Chen, Haihua Hu. A Comprehensive Evaluation of the Plastid DNA Data Gaps of Vascular Plants in Species and Geographic Area [J]. Chinese Bulletin of Botany, 2025, 60(1): 1-16. |
| [10] | Ruoyue Li, Xiaochao Yang, Zhanqing Hao, Shihong Jia. The intensity of heat waves and insect herbivory on campus plants and their relationship with leaf functional traits [J]. Biodiv Sci, 2025, 33(1): 24283-. |
| [11] | Xiangtan Yao, Xinyi Zhang, Yang Chen, Ye Yuan, Wangda Cheng, Tianrui Wang, Yingxiong Qiu. Genomic resequencing reveals the genetic diversity of the cultivated water caltrop, and the origin and domestication of ‘Nanhuling’ [J]. Biodiv Sci, 2024, 32(9): 24212-. |
| [12] | WANG Yi-Tong, Yeerjiang BAIKETUERHAN, LIAO Dan, WANG Juan. Correlation between elemental biometric characteristics and sexual dimorphism in leaves of dioecious Acer barbinerve at different growth stages [J]. Chin J Plant Ecol, 2024, 48(6): 760-769. |
| [13] | Chunjiao Xia, Yunguang Li, Shu Xia, Wei Pang, Chunli Chen. Flow Cytometric Analysis and Sorting in Plant Genomics [J]. Chinese Bulletin of Botany, 2024, 59(5): 774-782. |
| [14] | QU Ze-Kun, ZHU Li-Qin, JIANG Qi, WANG Xiao-Hong, YAO Xiao-Dong, CAI Shi-Feng, LUO Su-Zhen, sCHEN Guang-Shui. Nutrient foraging strategies of arbuscular mycorrhizal tree species in a subtropical evergreen broadleaf forest and their relationship with fine root morphology [J]. Chin J Plant Ecol, 2024, 48(4): 416-427. |
| [15] | Lansha Luo, Wenpei Song, Qingzhu Hua, Dawei Li, Hong Liang, Xianzhi Zhang. Research Progress on Plant Sex-determination Genes and Their Epigenetic Regulation [J]. Chinese Bulletin of Botany, 2024, 59(2): 278-290. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||