

Chinese Bulletin of Botany ›› 2020, Vol. 55 ›› Issue (1): 76-82.DOI: 10.11983/CBB19208 cstr: 32102.14.CBB19208
• TECHNIQUES AND METHODS • Previous Articles Next Articles
Dan Zhu,Hanwei Cao,Yuan Li,Dongtao Ren( )
)
Received:2019-10-24
Accepted:2019-12-31
Online:2020-01-01
Published:2020-01-03
Contact:
Dongtao Ren
Dan Zhu,Hanwei Cao,Yuan Li,Dongtao Ren. Protocols for Analyzing Plant Phospho-proteins[J]. Chinese Bulletin of Botany, 2020, 55(1): 76-82.
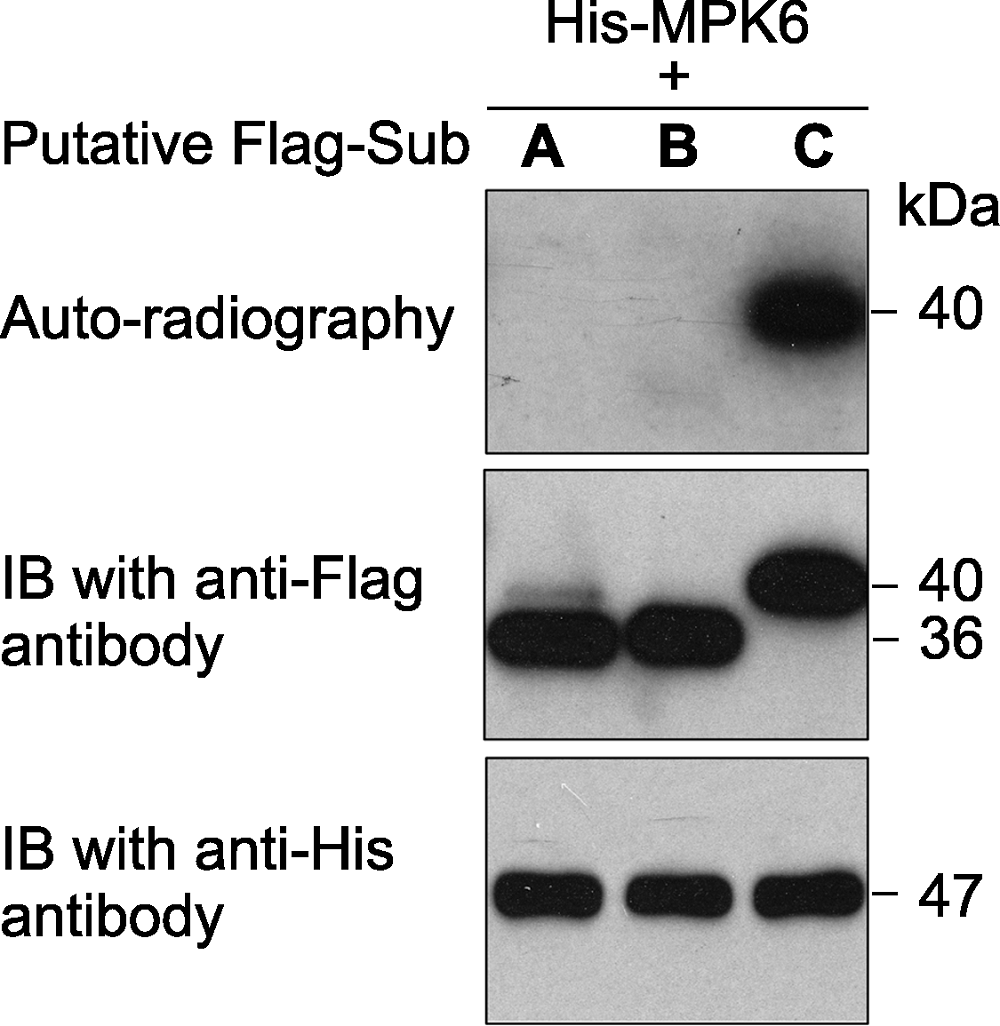
Figure 1 An in vitro phosphorylation assay of 3 putative substrate proteins by MPK6 in Arabidopsis thaliana MPK6 was fused with the 6×His tag and the 3 putative substrates were fused with the Flag tag. After phosphorylation reaction, the proteins in the mixture were separated on a SDS- PAGE gel. The gel was dried and exposed to X-ray film (top). Immunoblotting with anti-His and anti-Flag antibodies were used to show the levels of the substrate (middle) and kinase (bottom) proteins in the reactions.
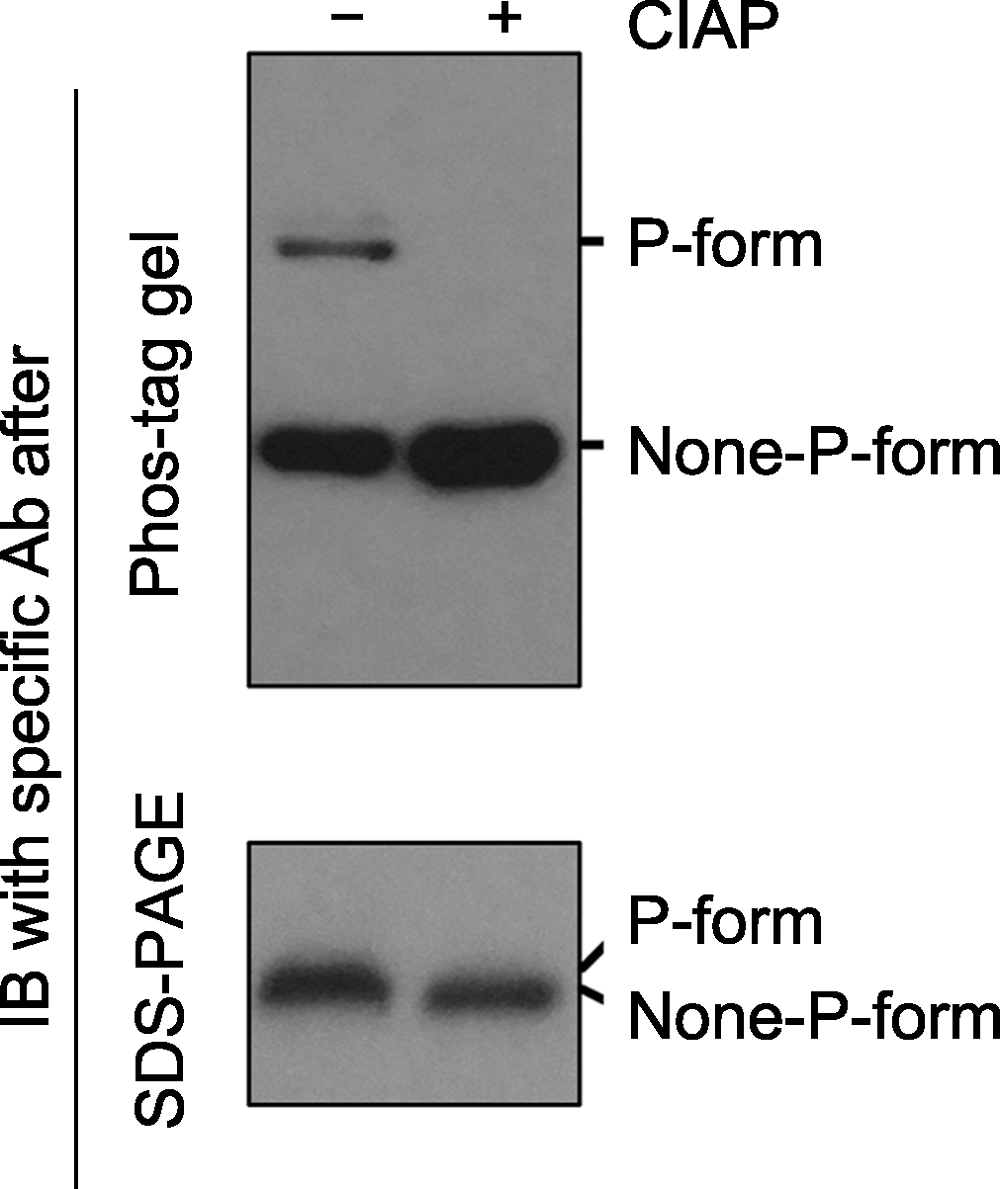
Figure 2 Immunoblotting detection of phospho-proteins in samples after Phos-tag gel and SDS-PAGE gel separation Phospho-proteins can be dephosphorylated by phosphatases (e.g. Calf intestine alkaline phosphatase, CIAP). Protein samples treated with (+) or without (-) CIAP were separated by Phos-tag (top) and SDS-PAGE (bottom) gels, and the specific protein was further detected by immunoblotting. The P-form and None-P-form of the protein were separated clearly by a Phos-tag gel (top), while the two forms were not separated by a SDS-PAGE gel (bottom). The missing of the upper band and the increasing of the bottom band after Phos-tag gel separation indicated that P-form protein was completely dephosphorylated by CIAP treatment.
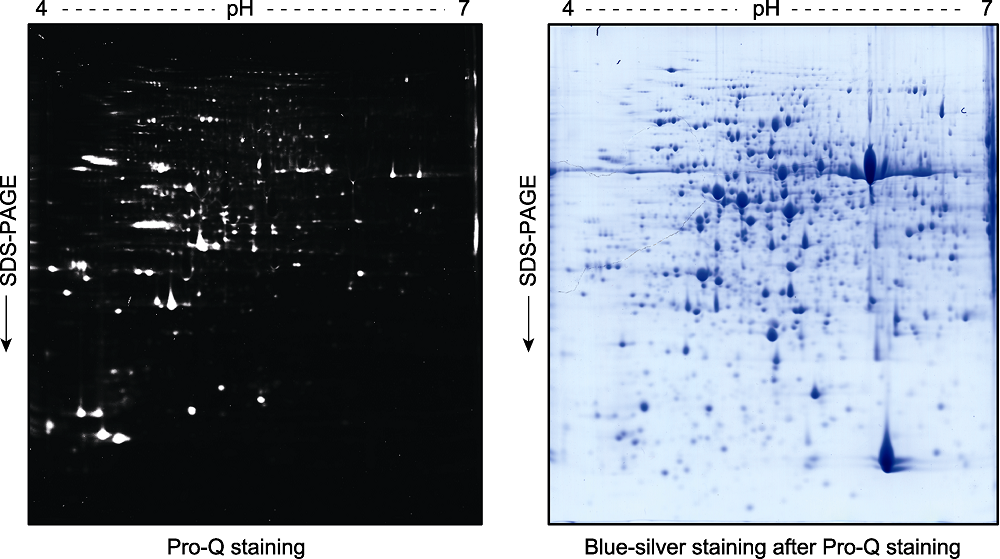
Figure 3 Pro-Q staining assay of the phospho-proteins in total proteins extracted from Arabidopsis thaliana seedlings and separated by a 2D gel Two-week-old Arabidopsis thaliana seedlings were used for total protein extraction. The total proteins were separated by a 2D gel and the phospho-proteins were stained by Pro-Q (left). The gel was scanned with a Typhoon 9410 fluorescence scanner, and then stained with blue-silver to visualize the proteins (right).
| [1] | Agrawal GK, Thelen JJ (2005). Development of a simplified, economical polyacrylamide gel staining protocol for phosphoproteins. Proteomics 5, 4684-4688. |
| [2] | Agrawal GK, Thelen JJ (2006). Large scale identification and quantitative profiling of phosphoproteins expressed during seed filling in oilseed rape. Mol Cell Proteomics 5, 2044-2059. |
| [3] | Blom N, Sicheritz-Pontén T, Gupta R, Gammeltoft S, Brunak S (2004). Prediction of post-translational glycosylation and phosphorylation of proteins from the amino acid sequence. Proteomics 4, 1633-1649. |
| [4] | Candiano G, Bruschi M, Musante L, Santucci L, Ghiggeri GM, Carnemolla B, Orecchia P, Zardi L, Righetti PG (2004). Blue silver: a very sensitive colloidal Coomassie G-250 staining for proteome analysis. Electrophoresis 25, 1327-1333. |
| [5] | Chao Q, Liu XY, Mei YC, Gao ZF, Chen YB, Qian CR, Hao YB, Wang BC (2014). Light-regulated phosphorylation of maize phosphoenolpyruvate carboxykinase plays a vital role in its activity. Plant Mol Biol 85, 95-105. |
| [6] | Chen MJ, Dixon JE, Manning G (2017). Genomics and evolution of protein phosphatases. Sci Signal 10, eaag1796. |
| [7] | de la Fuente van Bentem S, Hirt H (2007). Using phosphoproteomics to reveal signaling dynamics in plants. Trends Plant Sci 12, 404-411. |
| [8] | Ficarro SB, McCleland ML, Stukenberg PT, Burke DJ, Ross MM, Shabanowitz J, Hunt DF, White FM (2002). Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat Biotechnol 20, 301-305. |
| [9] | Fischer EH, Krebs EG (1955). Conversion of phosphorylase B to phosphorylase A in muscle extracts. J Biol Chem 216, 121-132. |
| [10] | Frost DC, Li LJ (2014). Recent advances in mass spectrometry-based glycoproteomics. Adv Protein Chem Struct Biol 95, 71-123. |
| [11] | Hubbard MJ, Cohen P (1993). On target with a new mechanism for the regulation of protein phosphorylation. Trends Biochem Sci 18, 172-177. |
| [12] | Ke YQ, Han GQ, He HQ, Li JX (2009). Differential regulation of proteins and phosphoproteins in rice under drought stress. Biochem Biophys Res Commun 379, 133-138. |
| [13] | Khan M, Takasaki H, Komatsu S (2005). Comprehensive phosphoproteome analysis in rice and identification of phosphoproteins responsive to different hormones/stresses. J Proteome Res 4, 1592-1599. |
| [14] | Kim HS, Fernandes G, Lee CW (2016). Protein phosphatases involved in regulating mitosis: facts and hypotheses. Mol Cell 39, 654-662. |
| [15] | Kinoshita E, Kinoshita-Kikuta E, Koike T (2007). Specific recognition and detection of phosphorylated proteins using characteristics of metal ion. Yakugaku Zasshi 127, 1897-1913. |
| [16] | Kosako H, Nagano K (2011). Quantitative phosphoproteomics strategies for understanding protein kinase-mediated signal transduction pathways. Expert Rev Proteomics 8, 81-94. |
| [17] | Krupa A, Preethi G, Srinivasan N (2004). Structural modes of stabilization of permissive phosphorylation sites in protein kinases: distinct strategies in Ser/Thr and Tyr kinases. J Mol Biol 339, 1025-1039. |
| [18] | Laemmli UK (1970). Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227, 680-685. |
| [19] | Laugesen S, Bergoin A, Rossignol M (2004). Deciphering the plant phosphoproteome: tools and strategies for a challenging task. Plant Physiol Biochem 42, 929-936. |
| [20] | Peck SC (2003). Early phosphorylation events in biotic stress. Curr Opin Plant Biol 6, 334-338. |
| [21] | Prak S, Hem S, Boudet J, Viennois G, Sommerer N, Rossignol M, Maurel C, Santoni V (2008). Multiple phosphorylations in the C-terminal tail of plant plasma membrane aquaporins: role in subcellular trafficking of AtPIP2;1 in response to salt stress. Mol Cell Proteomics 7, 1019-1030. |
| [22] | Wang PC, Zhao Y, Li ZP, Hsu CC, Liu X, Fu LW, Hou YJ, Du YY, Xie SJ, Zhang CG, Gao JH, Cao MJ, Huang XS, Zhu YF, Tang K, Wang XG, Tao WA, Xiong Y, Zhu JK (2018). Reciprocal regulation of the TOR kinase and ABA receptor balances plant growth and stress response. Mol Cell 69, 100-112. |
| [23] | Whiteman SA, Nühse TS, Ashford DA, Sanders D, Maathuis FJ (2008). A proteomic and phosphoproteomic analysis of Oryza sativa plasma membrane and vacuolar membrane. Plant J 56, 146-156. |
| [24] | Wu CF, Wang RN, Liang QJ, Liang JJ, Li WK, Jung SY, Qin J, Lin SH, Kuang J (2010). Dissecting the M phase- specific phosphorylation of serine-proline or threonine- proline motifs. Mol Biol Cell 21, 1470-1481. |
| [25] | Yang C, Wang ZG, Zhu PF (2004). Recent advances of protein phosphorylation in proteome. Prog Physiol Sci 35, 119-124. |
| [26] | Yin XJ, Wang X, Komatsu S (2018). Phosphoproteomics: protein phosphorylation in regulation of seed germination and plant growth. Curr Protein Pept Sci 19, 401-412. |
| [27] | Zhang X, Cui YN, Yu M, Su BD, Gong W, Baluška F, Komis G, Samaj J, Shan XY, Lin JX (2019). Phosphorylation-mediated dynamics of nitrate transceptor NRT1.1 regulate auxin flux and nitrate signaling in lateral root growth. Plant Physiol 181, 480-498. |
| [28] | Zhu WG (2017). Regulation of p53 acetylation. Sci China Life Sci 60, 321-323. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||