

Chinese Bulletin of Botany ›› 2018, Vol. 53 ›› Issue (3): 313-321.DOI: 10.11983/CBB17193 cstr: 32102.14.CBB17193
Special Issue: 药用植物专辑 (2018年53卷3期)
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Ren Mengyun1,2, Du Leshan1, Chen Yanjun1,2, Zhang Dun2, Shen Qi2, Guan Xiao1,*( ), Zhang Yindong2,*(
), Zhang Yindong2,*( )
)
Received:2017-10-18
Accepted:2018-04-30
Online:2018-05-01
Published:2018-09-11
Contact:
Guan Xiao,Zhang Yindong
Ren Mengyun, Du Leshan, Chen Yanjun, Zhang Dun, Shen Qi, Guan Xiao, Zhang Yindong. Analysis on Genetic Diversity of Cynomorium songaricum by ITS Sequence[J]. Chinese Bulletin of Botany, 2018, 53(3): 313-321.
| Sample No. | Location | Longitude E (°) | Latitude N (°) | Height (m) | Number | |
|---|---|---|---|---|---|---|
| R1 | Zhangye, Gansu | 102.522 | 36.822 | 1803 | 11 | |
| R2 | Minqin, Gansu | 100.772 | 39.223 | 1370 | 17 | |
| R3 | Jinchang, Gansu | 102.54 | 38.44 | 1466 | 7 | |
| R4 | Sandan, Gansu | 101.118 | 38.897 | 2020 | 10 | |
| R5 | Gaotai, Gansu | 99.086 | 39.787 | 1339 | 11 | |
| R6 | Sunan, Gansu | 99.176 | 39.604 | 1395 | 10 | |
| R7 | Sunan, Gansu | 99.474 | 39.445 | 1391 | 12 | |
| R8 | Jiuquan, Gansu | 98.55 | 40.3 | 1232 | 4 | |
| R9 | Yumen, Gansu | 97.196 | 40.516 | 1395 | 10 | |
| R10 | Guazhou, Gansu | 95.585 | 40.212 | 1343 | 11 | |
| R11 | Dunhuang, Gansu | 94.583 | 40.368 | 1033 | 11 | |
| R12 | Subei, Gansu | 96.619 | 39.397 | 2355 | 10 | |
| R13 | Delingha, Qinghai | 97.334 | 37.206 | 2780 | 11 | |
| R14 | Geermu, Qinghai | 92.839 | 36.701 | 2790 | 7 | |
| R15 | Dulan, Qinghai | 98.164 | 36.476 | 2972 | 12 | |
| R16 | Wulan, Qinghai | 98.622 | 36.458 | 2917 | 12 | |
| R17 | Gonghe, Qinghai | 100.183 | 36.446 | 2904 | 11 | |
| R18 | Guide, Qinghai | 101.639 | 36.219 | 2389 | 11 |
Table 1 Location of 18 populations of Cynomorium songaricum
| Sample No. | Location | Longitude E (°) | Latitude N (°) | Height (m) | Number | |
|---|---|---|---|---|---|---|
| R1 | Zhangye, Gansu | 102.522 | 36.822 | 1803 | 11 | |
| R2 | Minqin, Gansu | 100.772 | 39.223 | 1370 | 17 | |
| R3 | Jinchang, Gansu | 102.54 | 38.44 | 1466 | 7 | |
| R4 | Sandan, Gansu | 101.118 | 38.897 | 2020 | 10 | |
| R5 | Gaotai, Gansu | 99.086 | 39.787 | 1339 | 11 | |
| R6 | Sunan, Gansu | 99.176 | 39.604 | 1395 | 10 | |
| R7 | Sunan, Gansu | 99.474 | 39.445 | 1391 | 12 | |
| R8 | Jiuquan, Gansu | 98.55 | 40.3 | 1232 | 4 | |
| R9 | Yumen, Gansu | 97.196 | 40.516 | 1395 | 10 | |
| R10 | Guazhou, Gansu | 95.585 | 40.212 | 1343 | 11 | |
| R11 | Dunhuang, Gansu | 94.583 | 40.368 | 1033 | 11 | |
| R12 | Subei, Gansu | 96.619 | 39.397 | 2355 | 10 | |
| R13 | Delingha, Qinghai | 97.334 | 37.206 | 2780 | 11 | |
| R14 | Geermu, Qinghai | 92.839 | 36.701 | 2790 | 7 | |
| R15 | Dulan, Qinghai | 98.164 | 36.476 | 2972 | 12 | |
| R16 | Wulan, Qinghai | 98.622 | 36.458 | 2917 | 12 | |
| R17 | Gonghe, Qinghai | 100.183 | 36.446 | 2904 | 11 | |
| R18 | Guide, Qinghai | 101.639 | 36.219 | 2389 | 11 |
| Haplotype | Variable sites | Abundance | ||||||
|---|---|---|---|---|---|---|---|---|
| 63 | 86 | 197 | 274 | 421 | 605 | 623 | ||
| H1 | G | G | G | T | G | C | G | 143 |
| H2 | A | G | G | T | G | C | T | 3 |
| H3 | G | A | G | T | G | C | G | 1 |
| H4 | G | A | G | T | G | G | G | 2 |
| H5 | A | G | G | T | G | C | G | 26 |
| H6 | G | G | A | T | G | C | G | 5 |
| H7 | G | G | A | G | G | C | G | 1 |
| H8 | G | G | G | T | G | G | G | 4 |
| H9 | G | G | G | T | A | C | G | 3 |
Table 2 Variable sites of ITS sequence haplotypes of Cynomorium songaricum
| Haplotype | Variable sites | Abundance | ||||||
|---|---|---|---|---|---|---|---|---|
| 63 | 86 | 197 | 274 | 421 | 605 | 623 | ||
| H1 | G | G | G | T | G | C | G | 143 |
| H2 | A | G | G | T | G | C | T | 3 |
| H3 | G | A | G | T | G | C | G | 1 |
| H4 | G | A | G | T | G | G | G | 2 |
| H5 | A | G | G | T | G | C | G | 26 |
| H6 | G | G | A | T | G | C | G | 5 |
| H7 | G | G | A | G | G | C | G | 1 |
| H8 | G | G | G | T | G | G | G | 4 |
| H9 | G | G | G | T | A | C | G | 3 |
| Population code | Samples | Haptotypes | Hd |
|---|---|---|---|
| R1 | 11 | H1, H2, H3, H4 | 0.25974 |
| R2 | 17 | H1, H5, H6 | 0.11586 |
| R3 | 7 | H1, H5 | 0.43956 |
| R4 | 10 | H1, H2, H5 | 0.42632 |
| R5 | 11 | H1, H6 | 0.24675 |
| R6 | 10 | H1, H7, H8 | 0.19474 |
| R7 | 12 | H1, H5 | 0.08333 |
| R8 | 4 | H1, H5, H9 | 0.60714 |
| R9 | 10 | H1, H8 | 0.10000 |
| R10 | 11 | H1, H5, H8 | 0.17749 |
| R11 | 11 | H1, H4, H6 | 0.17749 |
| R12 | 10 | H1 | 0 |
| R13 | 11 | H1 | 0 |
| R14 | 7 | H1, H6 | 0.14286 |
| R15 | 12 | H1, H9 | 0.08333 |
| R16 | 12 | H1, H9 | 0.08333 |
| R17 | 11 | H1, H8 | 0.09091 |
| R18 | 11 | H1 | 0 |
| Total | 188 | 0.29420 |
Table 3 Haplotype diversity and composition of Cynomorium songaricum from18 populations
| Population code | Samples | Haptotypes | Hd |
|---|---|---|---|
| R1 | 11 | H1, H2, H3, H4 | 0.25974 |
| R2 | 17 | H1, H5, H6 | 0.11586 |
| R3 | 7 | H1, H5 | 0.43956 |
| R4 | 10 | H1, H2, H5 | 0.42632 |
| R5 | 11 | H1, H6 | 0.24675 |
| R6 | 10 | H1, H7, H8 | 0.19474 |
| R7 | 12 | H1, H5 | 0.08333 |
| R8 | 4 | H1, H5, H9 | 0.60714 |
| R9 | 10 | H1, H8 | 0.10000 |
| R10 | 11 | H1, H5, H8 | 0.17749 |
| R11 | 11 | H1, H4, H6 | 0.17749 |
| R12 | 10 | H1 | 0 |
| R13 | 11 | H1 | 0 |
| R14 | 7 | H1, H6 | 0.14286 |
| R15 | 12 | H1, H9 | 0.08333 |
| R16 | 12 | H1, H9 | 0.08333 |
| R17 | 11 | H1, H8 | 0.09091 |
| R18 | 11 | H1 | 0 |
| Total | 188 | 0.29420 |
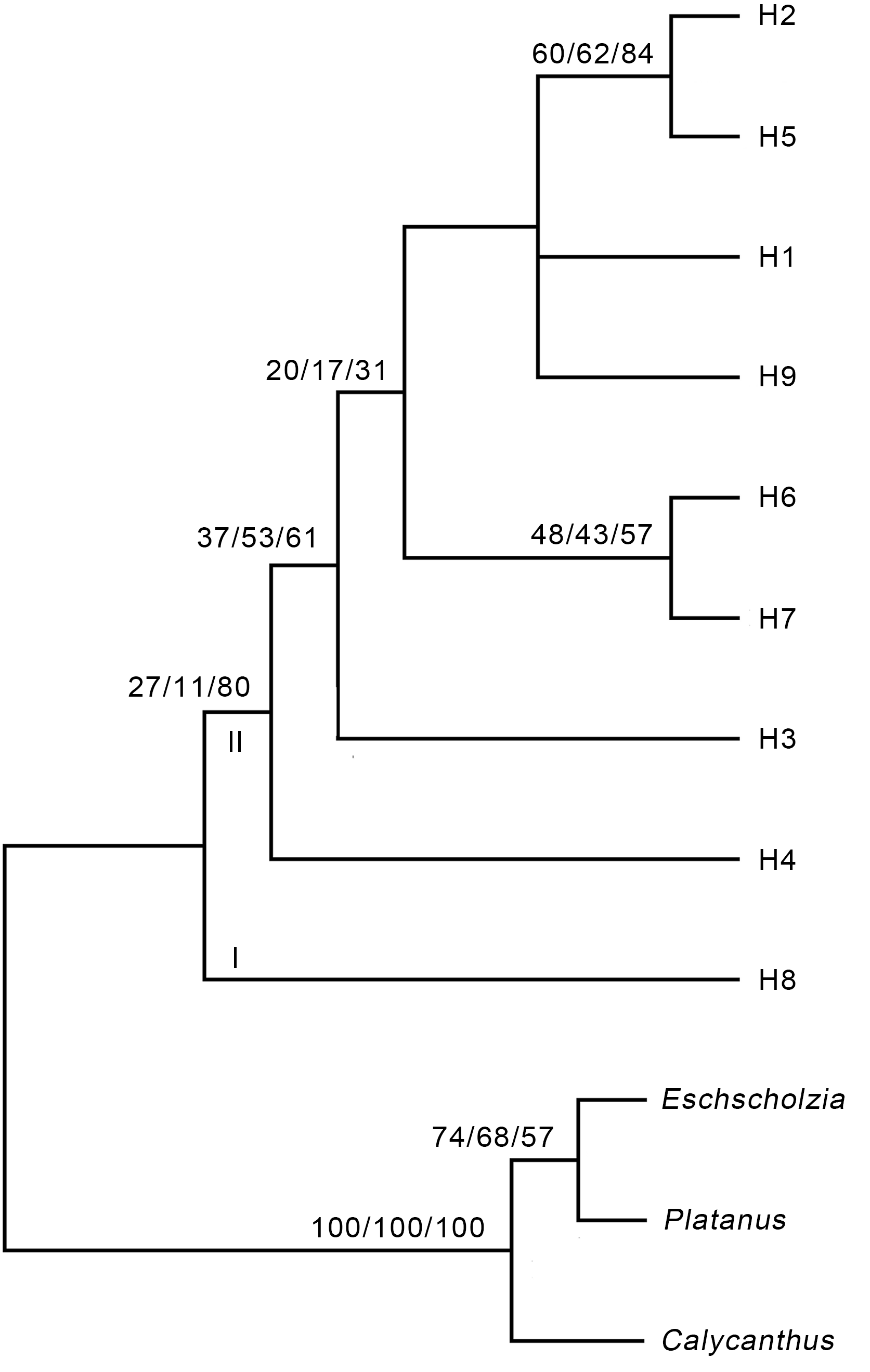
Figure 1 Strict consensus tree based on the ITS sequence of Cynomorium songaricumLength=784, CI=0.949 0, RI=0.911 7, RCI=0.865 2. The num- bers on the branch represent the support rate of MP/ML/BI, respectively.
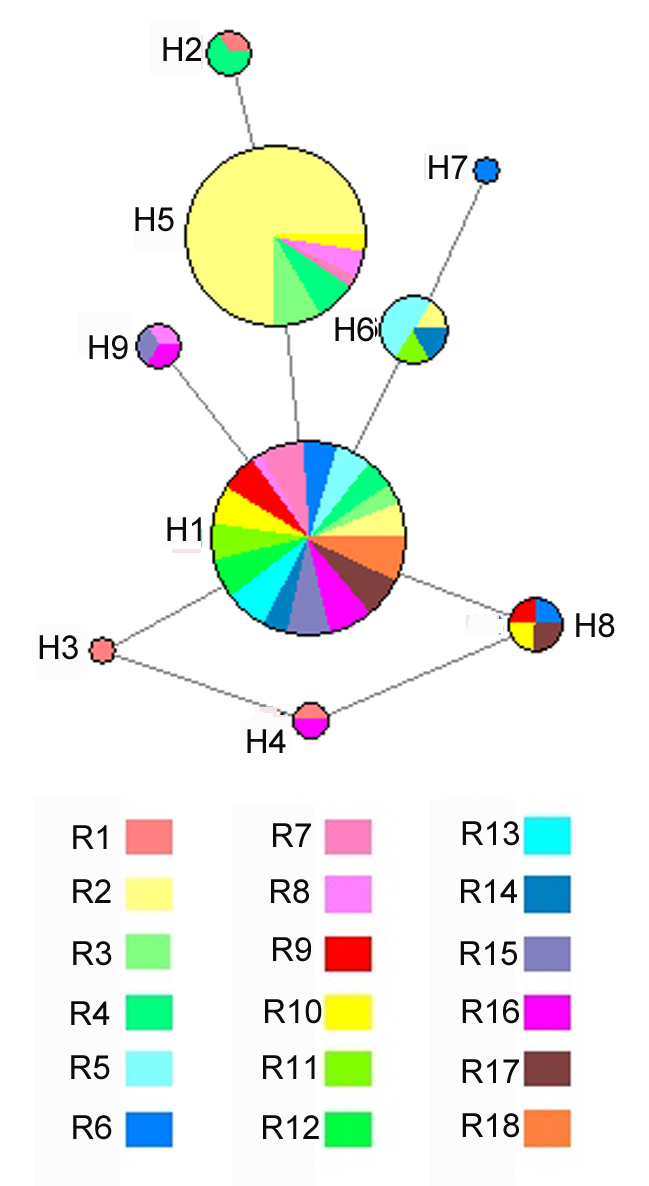
Figure 2 Haplotype network based on ITS sequence of Cynomorium songaricum from 18 populationsThe size of circles are proportional to the relative frequency of the haplotype; Different colors represent different populations of Cynomorium songaricum.
| Sequence | HS (SE) | HT (SE) | GST (SE) | NST (SE) |
|---|---|---|---|---|
| ITS | 0.179 (0.0390) | 0.277 (0.0820) | 0.353 (0.1914) | 0.408 (0.2038) |
Table 4 Genetic structural parameters of Cynomorium songaricum
| Sequence | HS (SE) | HT (SE) | GST (SE) | NST (SE) |
|---|---|---|---|---|
| ITS | 0.179 (0.0390) | 0.277 (0.0820) | 0.353 (0.1914) | 0.408 (0.2038) |
| Source of variation | df | Sum of squares | Variance components | Percentage | FST | P |
|---|---|---|---|---|---|---|
| Among populations | 17 | 28.833 | 0.07689 Va | 44.57 | 0.44566 | <0.001* |
| Within populations | 358 | 34.239 | 0.09564 Vb | 55.43 | ||
| Total | 375 | 63.072 | 0.17253 |
Table 5 Analysis of molecular variance for ribosome haplotypes of Cynomorium songaricum
| Source of variation | df | Sum of squares | Variance components | Percentage | FST | P |
|---|---|---|---|---|---|---|
| Among populations | 17 | 28.833 | 0.07689 Va | 44.57 | 0.44566 | <0.001* |
| Within populations | 358 | 34.239 | 0.09564 Vb | 55.43 | ||
| Total | 375 | 63.072 | 0.17253 |
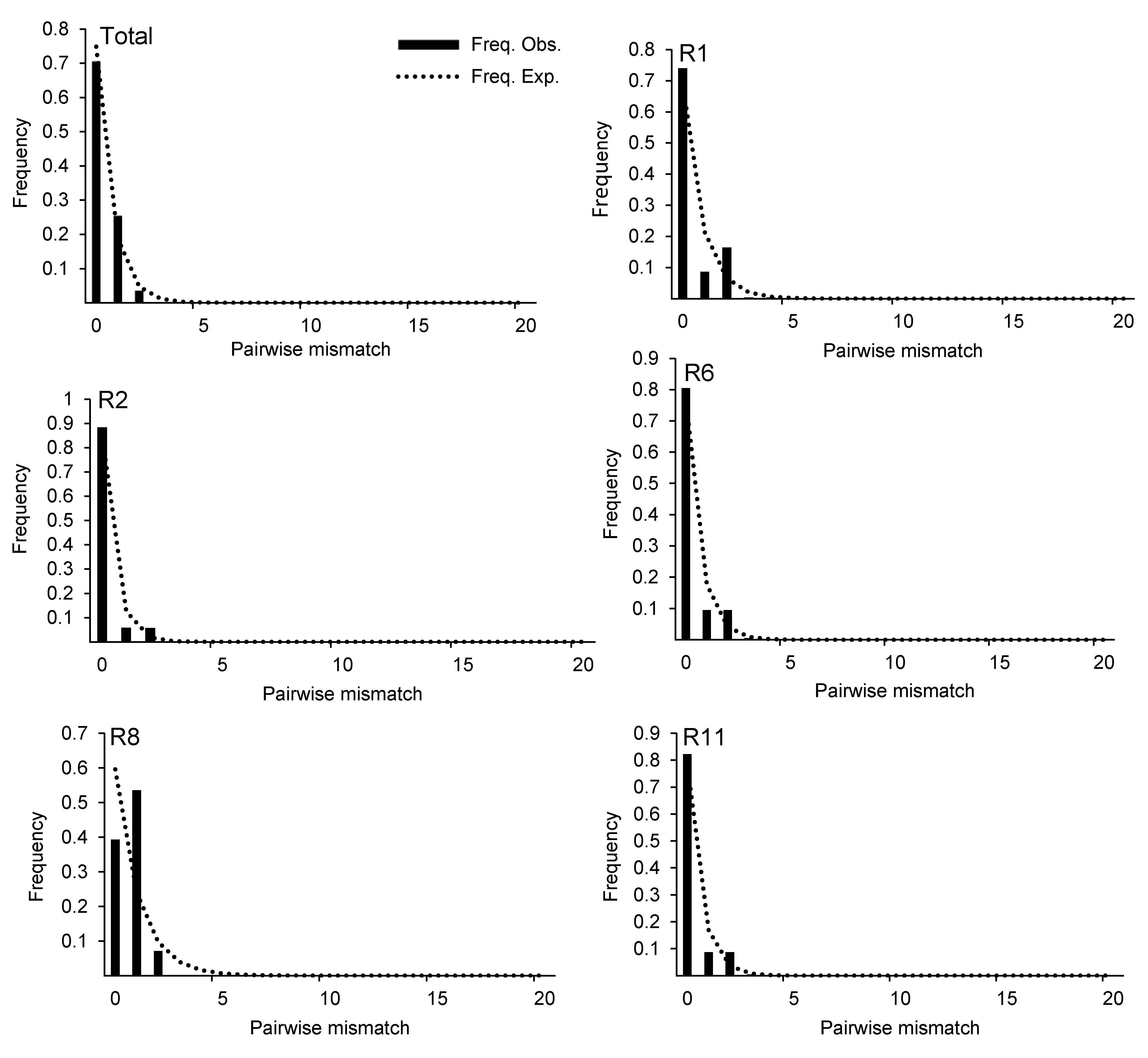
Figure 3 Mismatch distribution analysis for the populations of Cynomorium songaricum based on ITS sequenceCylindricality represents the expected distribution of variation sites under the population expansion model; dotted line represents the actual distribution of variation sites.
| [1] | 陈贵林, 岳鑫, 刘广达 (2011). 锁阳愈伤组织体系的建立及遗传多样性研究. 见: 第十届全国药用植物及植物药学术研讨会论文集. 昆明: 中国植物学会. pp. 26. |
| [2] | 陈叶, 高海宁, 高宏, 韩多宏, 罗光宏, 张勇 (2013). 甘肃河西走廊道地药材锁阳的分布和利用. 中兽医医药杂志 32, 77-79. |
| [3] | 郝媛媛, 岳利军, 康建军, 王锁民 (2012). “沙漠人参”肉苁蓉和锁阳研究进展. 草业学报21, 286-293. |
| [4] | 黄建峰, 李朗, 李捷 (2016). 樟属植物ITS序列多态性分析. 植物学报 51, 609-619. |
| [5] | 李洪芹, 马昌豪, 彭艳丽 (2015). 山东玫瑰花核糖体rDNA ITS序列分析初步探究. 天津中医药大学学报 34, 104-107. |
| [6] | 李金博, 高丽华, 周美亮, 李诗刚, 彭昭良, 吴燕民 (2016). 15个白三叶品种的ISSR和ITS遗传多样性分析. 草业科学 33, 1147-1153. |
| [7] | 李学营, 彭建营, 白瑞霞 (2005). 基于核rDNA的ITS序列在种子植物系统发育研究中的应用. 西北植物学报 25, 829-834. |
| [8] | 宁淑萍, 颜海飞, 郝刚, 葛学军 (2008). 植物DNA条形码研究进展. 生物多样性 16, 417-425. |
| [9] | 宋宇婷, 马大龙, 隋心, 穆立蔷 (2011). 黑龙江、吉林地区松茸ITS序列遗传多样性分析. 中国酿造 30, 133-136. |
| [10] | 岳鑫, 段园园, 陈贵林 (2013). 锁阳愈伤组织诱导和增殖及不定根分化. 植物生理学报 49, 1421-1426. |
| [11] | 中国科学院中国植物志编辑委员会 (2000). 中国植物志(第五十三卷第二分册). 北京: 科学出版社. pp. 152-154. |
| [12] | Avise JC (2000).Phylogeography: the History and Formation of Species. Cambridge: Harvard University Press. pp. 134-135. |
| [13] | Bandelt HJ, Forster P, Röhl A (1999). Median-joining networks for inferring intraspecific phylogenies.Mol Biol Evol 16, 37-48. |
| [14] | Barkman TJ, McNeal JR, Lim SH, Coat G, Croom HB, Young ND, Depamphilis CW (2007). Mitochondrial DNA suggests at least 11 origins of parasitism in angiosperms and reveals genomic chimerism in parasitic plants.BMC Evol Biol 7, 248. |
| [15] | Cibrián-Jaramillo A, Bacon CD, Garwood NC, Bateman RM, Thomas MM, Russell S, Bailey CD, Hahn WJ, Bridgewater SG, DeSalle R (2009). Population genetics of the understory fishtail palm Chamaedorea ernestiau- gusti in Belize: high genetic connectivity with local differen- tiation. BMC Genet 10, 65. |
| [16] | Cooke DEL, Duncan JM (1997). Phylogenetic analysis of Phytophthora species based on ITS1 and ITS2 sequences of the ribosomal RNA gene repeat. Mycol Res 101, 667-677. |
| [17] | Cosacov A, Johnson LA, Paiaro V, Cocucci AA, Córdoba FE, Sérsic AN, Crisci J (2013). Precipitation rather than temperature influenced the phylogeography of the endemic shrub Anarthrophyllum desideratum in the Patagonian steppe. J Biogeogra 40, 168-182. |
| [18] | Excoffier L (2004). Patterns of DNA sequence diversity and genetic structure after a range expansion: lessons from the infinite-island model.Mol Ecol 13, 853-864. |
| [19] | Excoffier L, Laval G, Schneider S (2007). Arlequin (version 3.0): an integrated software package for population genetics data analysis.Evol Bioinform 1, 47-50. |
| [20] | Excoffier L, Smouse PE, Quattro JM (1992). Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data.Genetics 131, 479-491. |
| [21] | Hall TA (1999). BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/ 98/NT. In: Information Retrieval Ltd., Nucleic Acids Symposium Series, vol. 41. London: IRL Press. pp. 95-98. |
| [22] | Hou DY, Song JY, Shi LC, Ma XC, Xin TY, Han JP, Xiao W, Sun ZY, Cheng RY, Yao H (2013a). Stability and accuracy assessment of identification of traditional Chinese materia medica using DNA barcoding: a case study on Flos Lonicerae Japonicae.Biomed Res Int 2013, 549037. |
| [23] | Hou DY, Song JY, Yao H, Han JP, Pang XH, Shi LC, Wang XC, Chen SL (2013b). Molecular identification of corni fructus and its adulterants by ITS/ITS2 sequences.Chin J Nat Med 11, 121-127. |
| [24] | Kumar S, Stecher G, Tamura K (2016). MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger data- sets.Mol Biol Evol 33, 1870-1874. |
| [25] | Lahaye R, Van der Bank M, Bogarin D, Warner J, Pupulin F, Gigot G, Maurin O, Duthoit S, Barraclough TG, Savolainen V (2008). DNA barcoding the floras of biodiversity hotspots.Proc Natl Acad Sci USA 105, 2923-2928. |
| [26] | Librado P, Rozas J (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data.Bioinformatics 25, 1451-1452. |
| [27] | Liu GD, Chen GL, Li W, Li CX (2013). Genetic and phytochemical diversities of Cynomorium songaricum Rupr. in Northwest China indicated by ISSR markers and HPLC- fingerprinting. Biochem Syst Ecol 48, 34-41. |
| [28] | Newmaster SG, Fazekas AJ, Steeves RAD, Janovec J (2008). Testing candidate plant barcode regions in the My- risticaceae.Mol Ecol Res 8, 480-490. |
| [29] | Nickrent DL, Der JP, Anderson FE (2005). Discovery of the photosynthetic relatives of the “Maltese mushroom” Cynomorium. BMC Evol Biol 5, 38. |
| [30] | Pons O, Petit RJ (1996). Measwring and testing genetic differentiation with ordered Versus unordered alleles. Genetics 144, 1237-1245. |
| [31] | Rogers AR, Harpending H (1992). Population growth makes waves in the distribution of pairwise genetic differences.Mol Biol Evol 9, 552-569. |
| [32] | Slatkin M, Hudson RR (1991). Pairwise comparisons of mitochondrial DNA sequences in stable and exponentially growing populations.Genetics 129, 555-562. |
| [33] | Song JY, Shi LC, Li DZ, Sun YZ, Niu YY, Chen ZD, Luo HM, Pang XH, Sun ZY, Liu C, Lv AP, Deng YP, Larson-Rabin Z, Wilkinson M, Chen SL (2012). Extensive pyrosequencing reveals frequent intra-genomic variations of internal transcribed spacer regions of nuclear ribosomal DNA.PLoS One 7, e43971. |
| [34] | Swafford DL (2002). PAUP*: Phylogenetic Analysis Using Parsimony (* and Other Methods). Version 4.0 10b. Sunderland, MA:Sinauer Associates. |
| [35] | Zhang ZH, Li CQ, Li JH (2009). Phylogenetic placement of Cynomorium in Rosales inferred from sequences of the inverted repeat region of the chloroplast genome. J Syst Evol 47, 297-304. |
| [1] | TONG Jin-Lian, ZHANG Bo-Na, TANG TANG Lu-yao, YE Lin-Feng, LI Shu-Wen, LI Yan, Zhong-Yuan WANG. Regional differentiation of functional trait network of C4 plants Setaria viridis along precipitation gradient [J]. Chin J Plant Ecol, 2025, 49(预发表): 1-. |
| [2] | Jing Zhang Li JunPan Han XU Yi-De LI Hai ShengHe. Comparison of plant biomass in conifer and broadleaved mixed artificial forests in south subtropical area and analyses of the influential factors [J]. Chin J Plant Ecol, 2025, 49(化学计量与功能性状): 0-0. |
| [3] | Dai Lijun, Xiang Lingyi, Jian Chen, WANG Xiaofeng. The alteration of backwater dynamics in the Three Gorges has intensified the local differentiation of functional traits among typical herbaceous plants within the riparian zone of small watersheds [J]. Chin J Plant Ecol, 2025, 49(化学计量与功能性状): 1-. |
| [4] | 黄 承玲, Han Li Rong, Ling Qing Hong, Xiong Yang Sheng, Ling Tian Xiao, Xia Guowei Xia Guowei, Ren Chen Zheng, Wei Zhou. Study on genetic conservation of Rhododendron liboense based on SNP molecular markers,a plant species with extremely small populations [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [5] | Min Luo, Yongchuan Yang, Cheng Jin, Lihua Zhou, Yuxiao Long. Composition profile and response to human activity of mammals in the urban forests of central Chongqing [J]. Biodiv Sci, 2025, 33(5): 24402-. |
| [6] | LI Meng-Qi, MIAO Ling-Feng, LI Da-Dong, LONG Yi-Fan, YE Bing-Bing, YANG Fan. Response of mangrove fine root functional traits to sediment nutrient changes at different tide levels in Dongzhaigang, Hainan, China [J]. Chin J Plant Ecol, 2025, 49(4): 552-561. |
| [7] | GUO Li-Qi, YAN Xiao-Lei, CAO Lei, GAO Jing, LIU Rui-Qiang, ZHOU Xu-Hui. Effects of mycorrhizal types and root traits of tree species on rhizosphere microbial network complexity [J]. Chin J Plant Ecol, 2025, 49(4): 573-584. |
| [8] | Zhang Ruli, Li Dezhu, Zhang Yuxiao. Population Genetic Structure and Climate Adaptation Analysis of Brachystachyum densiflorum [J]. Chinese Bulletin of Botany, 2025, 60(3): 407-424. |
| [9] | Zhang Songqi, Lu Yi, Chen Bingyao, Yang Guang, Wang Yanping, Chen Chuanwu. A dataset on the morphological, life-history, and ecological traits of cetaceans worldwide [J]. Biodiv Sci, 2025, 33(2): 24442-. |
| [10] | Zhao Yifan, Wang Yanping. A database of life-history, ecological, and biogeographical traits of snakes worldwide [J]. Biodiv Sci, 2025, 33(2): 24476-. |
| [11] | Zhigang Yang, Pengcheng Zhang, Haiwen Chang, Liru Kang, Yi Zuo, Haoxin Xiang, Fengying Han. Genetic Diversity Analysis of Pepper Germplasms Based on Morphological Traits and SSR Markers [J]. Chinese Bulletin of Botany, 2025, 60(2): 218-234. |
| [12] | LI Shu-Wen, TANG Lu-Yao, ZHANG Bo-Na, YE Lin-Feng, TONG Jin-Lian, XIE Jiang-Bo, LI Yan, WANG Zhong-Yuan. Regional differentiation of cooperative relationships between Ulmus pumila branches and leaves along precipitation gradients [J]. Chin J Plant Ecol, 2025, 49(2): 282-294. |
| [13] | Jiachen Wang, Tangjun Xu, Wei Xu, Gaoji Zhang, Yijin You, Honghua Ruan, Hongyi Liu. Impact of urban landscape pattern on the genetic structure of Thereuopoda clunifera population in Nanjing, China [J]. Biodiv Sci, 2025, 33(1): 24251-. |
| [14] | LIAO Su-Hui, NI Long-Kang, QIN Jia-Shuang, TAN Yu, GU Da-Xing. Hydraulic regulation strategies of karst forest species exhibit variation across different successional stages in the mid-subtropical zone [J]. Chin J Plant Ecol, 2024, 48(9): 1223-1231. |
| [15] | Chuan Jin, Zijia Zhang, Kai Di, Weirong Zhang, Dong Qiao, Siyuan Cheng, Zhongmin Hu. A dataset on fluorescence, photosynthesis gas exchange, and leaf traits of Hainan tropical rainforest plant species [J]. Biodiv Sci, 2024, 32(9): 24139-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||