

Chinese Bulletin of Botany ›› 2015, Vol. 50 ›› Issue (2): 217-226.DOI: 10.3724/SP.J.1259.2015.00217 cstr: 32102.14.SP.J.1259.2015.00217
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Wen Fan1, Ying Xu1, Ting Xu1, Jing Xu1, 2, Takahiro Yonezawa1, Jiyin Gao3, Wenju Zhang1, *
Received:2014-03-14
Accepted:2014-05-09
Online:2015-03-01
Published:2015-04-10
Contact:
Zhang Wenju
About author:? These authors contributed equally to this paper
Wen Fan, Ying Xu, Ting Xu, Jing Xu, Takahiro Yonezawa, Jiyin Gao, Wenju Zhang. Intragenomic Polymorphism of the Internal Transcribed Spacer Region of Ribosomal DNA in Camellia hongkongensis (Theaceae) and Species Identification[J]. Chinese Bulletin of Botany, 2015, 50(2): 217-226.
Add to citation manager EndNote|Ris|BibTeX
URL: https://www.chinbullbotany.com/EN/10.3724/SP.J.1259.2015.00217
| Typea | rDNA region | Haptotypesb | Genetic distance | h±SD | π | Lengths (bp) | GC contents | ΔG (kcal∙mol-1)c |
|---|---|---|---|---|---|---|---|---|
| All | ITS1-5.8S-ITS2 | 47 | 0.210 5±0.011 7 | 0.991±0.008 | 0.166 72 | 589-687 | 0.557 | |
| ITS1 | 0.297 4±0.023 0 | 0.958±0.015 | 0.242 21 | 221-291 | 0.550 | |||
| 5.8S | 0.083 1±0.011 5 | 0.961±0.013 | 0.065 27 | 139-163 | 0.501 | |||
| ITS2 | 0.240 1±0.020 1 | 0.962±0.012 | 0.171 55 | 198-233 | 0.610 | |||
| F | ITS1-5.8S-ITS2 | 10 | 0.010 3±0.001 9 | 0.978±0.054 | 0.009 74 | 674-687 | 0.686 | |
| ITS1 | 0.007 0±0.002 1 | 0.667±0.163 | 0.006 47 | 281-291 | 0.719 | |||
| 5.8S | 0.008 4±0.003 6 | 0.667±0.163 | 0.008 32 | 163 | 0.543 | -36.90, -47.42 | ||
| ITS2 | 0.015 7±0.004 2 | 0.933±0.077 | 0.014 69 | 230-233 | 0.751 | |||
| P | ITS1-5.8S-ITS2 | 37 | 0.212 2±0.012 0 | 0.994±0.009 | 0.164 71 | 589-632 | 0.531 | |
| ITS1 | 0.283 5±0.023 0 | 0.964±0.015 | 0.225 14 | 221-258 | 0.515 | |||
| 5.8S | 0.095 8±0.013 3 | 0.959±0.014 | 0.073 47 | 139-163 | 0.491 | |||
| ITS2 | 0.248 3±0.020 9 | 0.953±0.016 | 0.176 47 | 198-223 | 0.581 | |||
| Pa1d | ITS1-5.8S-ITS2 | 5 | 0.003 6±0.001 6 | 1.000±0.126 | 0.003 58 | 614 | 0.523 | |
| ITS1 | 0.002 6±0.002 6 | 0.600±0.175 | 0.002 61 | 230 | 0.523 | |||
| 5.8S | 0.004 9±0.003 5 | 0.700±0.218 | 0.004 91 | 163 | 0.461 | -24.90, -36.00 | ||
| ITS2 | 0.003 6±0.002 6 | 0.700±0.218 | 0.003 62 | 221 | 0.567 | |||
| Pa2 | ITS1-5.8S-ITS2 | 8 | 0.006 1±0.002 0 | 1.000±0.063 | 0.006 06 | 607 | 0.541 | |
| ITS1 | 0.005 8±0.003 0 | 0.857±0.108 | 0.005 77 | 223 | 0.535 | |||
| 5.8S | 0.005 1±0.003 7 | 0.679±0.122 | 0.005 04 | 163 | 0.512 | -30.22, -35.70 | ||
| ITS2 | 0.007 2±0.003 7 | 0.750±0.139 | 0.007 11 | 221 | 0.568 | |||
| Pa3 | ITS1-5.8S-ITS2 | 5 | 0.005 8±0.002 1 | 1.000±0.126 | 0.005 77 | 589 | 0.529 | |
| ITS1 | 0.003 5±0.002 5 | 0.700±0.218 | 0.003 51 | 228 | 0.528 | |||
| 5.8S | 0.009 9±0.005 0 | 0.900±0.161 | 0.009 82 | 163 | 0.490 | -20.80, -23.62 | ||
| ITS2 | 0.005 1±0.003 6 | 0.800±0.164 | 0.005 05 | 198 | 0.564 | |||
| Pa4 | ITS1-5.8S-ITS2 | 9 | 0.011 3±0.002 9 | 0.972±0.064 | 0.011 11 | 610-611 | 0.577 | |
| ITS1 | 0.006 7±0.003 6 | 0.833±0.098 | 0.006 70 | 224-225 | 0.584 | |||
| 5.8S | 0.020 0±0.008 3 | 0.806±0.120 | 0.019 43 | 163 | 0.501 | -23.10, -34.50 | ||
| ITS2 | 0.009 6±0.004 6 | 0.694±0.147 | 0.009 47 | 223 | 0.624 | |||
| Pb | ITS1-5.8S-ITS2 | 9 | 0.151 4±0.009 5 | 1.000±0.052 | 0.121 31 | 590-632 | 0.582 | |
| ITS1 | 0.174 4±0.017 4 | 1.000±0.052 | 0.139 17 | 221-258 | 0.587 | |||
| 5.8S | 0.088 8±0.013 5 | 0.917±0.092 | 0.076 94 | 139-163 | 0.499 | -15.22, -42.12 | ||
| ITS2 | 0.181 0±0.019 4 | 0.917±0.092 | 0.136 33 | 206-221 | 0.644 |
Table 1 Characteristics of the ITS sequences of Camellia hongkongensis
| Typea | rDNA region | Haptotypesb | Genetic distance | h±SD | π | Lengths (bp) | GC contents | ΔG (kcal∙mol-1)c |
|---|---|---|---|---|---|---|---|---|
| All | ITS1-5.8S-ITS2 | 47 | 0.210 5±0.011 7 | 0.991±0.008 | 0.166 72 | 589-687 | 0.557 | |
| ITS1 | 0.297 4±0.023 0 | 0.958±0.015 | 0.242 21 | 221-291 | 0.550 | |||
| 5.8S | 0.083 1±0.011 5 | 0.961±0.013 | 0.065 27 | 139-163 | 0.501 | |||
| ITS2 | 0.240 1±0.020 1 | 0.962±0.012 | 0.171 55 | 198-233 | 0.610 | |||
| F | ITS1-5.8S-ITS2 | 10 | 0.010 3±0.001 9 | 0.978±0.054 | 0.009 74 | 674-687 | 0.686 | |
| ITS1 | 0.007 0±0.002 1 | 0.667±0.163 | 0.006 47 | 281-291 | 0.719 | |||
| 5.8S | 0.008 4±0.003 6 | 0.667±0.163 | 0.008 32 | 163 | 0.543 | -36.90, -47.42 | ||
| ITS2 | 0.015 7±0.004 2 | 0.933±0.077 | 0.014 69 | 230-233 | 0.751 | |||
| P | ITS1-5.8S-ITS2 | 37 | 0.212 2±0.012 0 | 0.994±0.009 | 0.164 71 | 589-632 | 0.531 | |
| ITS1 | 0.283 5±0.023 0 | 0.964±0.015 | 0.225 14 | 221-258 | 0.515 | |||
| 5.8S | 0.095 8±0.013 3 | 0.959±0.014 | 0.073 47 | 139-163 | 0.491 | |||
| ITS2 | 0.248 3±0.020 9 | 0.953±0.016 | 0.176 47 | 198-223 | 0.581 | |||
| Pa1d | ITS1-5.8S-ITS2 | 5 | 0.003 6±0.001 6 | 1.000±0.126 | 0.003 58 | 614 | 0.523 | |
| ITS1 | 0.002 6±0.002 6 | 0.600±0.175 | 0.002 61 | 230 | 0.523 | |||
| 5.8S | 0.004 9±0.003 5 | 0.700±0.218 | 0.004 91 | 163 | 0.461 | -24.90, -36.00 | ||
| ITS2 | 0.003 6±0.002 6 | 0.700±0.218 | 0.003 62 | 221 | 0.567 | |||
| Pa2 | ITS1-5.8S-ITS2 | 8 | 0.006 1±0.002 0 | 1.000±0.063 | 0.006 06 | 607 | 0.541 | |
| ITS1 | 0.005 8±0.003 0 | 0.857±0.108 | 0.005 77 | 223 | 0.535 | |||
| 5.8S | 0.005 1±0.003 7 | 0.679±0.122 | 0.005 04 | 163 | 0.512 | -30.22, -35.70 | ||
| ITS2 | 0.007 2±0.003 7 | 0.750±0.139 | 0.007 11 | 221 | 0.568 | |||
| Pa3 | ITS1-5.8S-ITS2 | 5 | 0.005 8±0.002 1 | 1.000±0.126 | 0.005 77 | 589 | 0.529 | |
| ITS1 | 0.003 5±0.002 5 | 0.700±0.218 | 0.003 51 | 228 | 0.528 | |||
| 5.8S | 0.009 9±0.005 0 | 0.900±0.161 | 0.009 82 | 163 | 0.490 | -20.80, -23.62 | ||
| ITS2 | 0.005 1±0.003 6 | 0.800±0.164 | 0.005 05 | 198 | 0.564 | |||
| Pa4 | ITS1-5.8S-ITS2 | 9 | 0.011 3±0.002 9 | 0.972±0.064 | 0.011 11 | 610-611 | 0.577 | |
| ITS1 | 0.006 7±0.003 6 | 0.833±0.098 | 0.006 70 | 224-225 | 0.584 | |||
| 5.8S | 0.020 0±0.008 3 | 0.806±0.120 | 0.019 43 | 163 | 0.501 | -23.10, -34.50 | ||
| ITS2 | 0.009 6±0.004 6 | 0.694±0.147 | 0.009 47 | 223 | 0.624 | |||
| Pb | ITS1-5.8S-ITS2 | 9 | 0.151 4±0.009 5 | 1.000±0.052 | 0.121 31 | 590-632 | 0.582 | |
| ITS1 | 0.174 4±0.017 4 | 1.000±0.052 | 0.139 17 | 221-258 | 0.587 | |||
| 5.8S | 0.088 8±0.013 5 | 0.917±0.092 | 0.076 94 | 139-163 | 0.499 | -15.22, -42.12 | ||
| ITS2 | 0.181 0±0.019 4 | 0.917±0.092 | 0.136 33 | 206-221 | 0.644 |
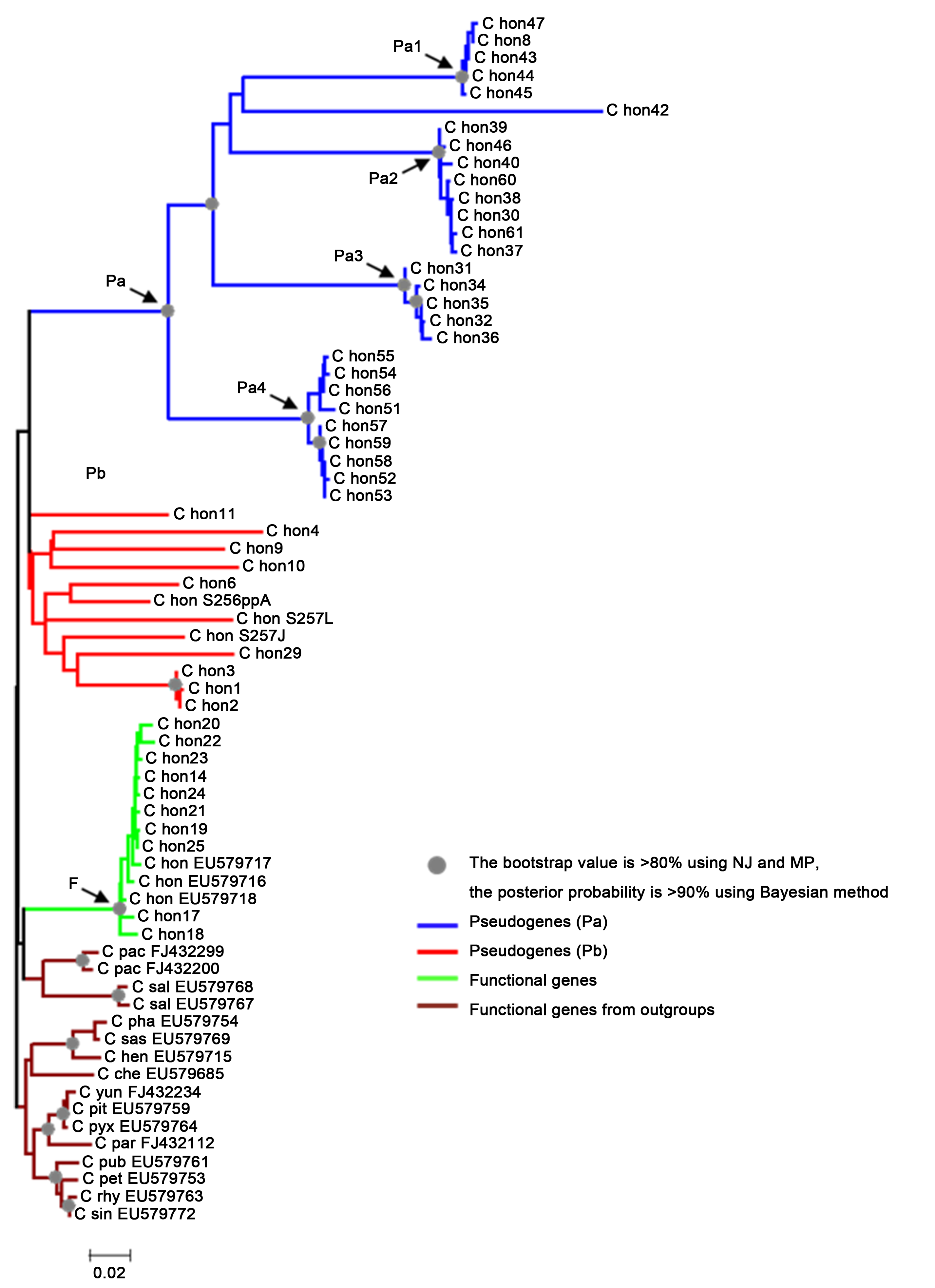
Figure 1 The Neighbor-joining tree of the ITS sequences of Camellia hongkongensis and its outgroups Among 53 sequences of C. honghkongensis, 47 sequences were obtained in this study, 3 sequences from GenBank, and 3 sequences (C hon S256ppA, C hon S257L, C hon S257J) from Dr. Tsou. Outgroups consist of 14 species (only show the first 3 letters of their specific names in this tree), and their sequences were obtained from GenBank
| 1 | 闵天禄 (2000). 世界山茶属的研究. 昆明: 云南科学技术出版社. pp. 4-308. |
| 2 | 闵天禄, 张文驹 (1996). 山茶属植物的进化与分布. 云南植物研究 18, 1-13. |
| 3 | 唐绍清, 施苏华, 钟杨, 王燕 (2004). 基于ITS序列探讨山茶属金花茶组的系统发育关系. 广西植物 24, 488-492. |
| 4 | 徐颖, 徐晶, 高继银, 张文驹 (2011). 山茶属植物ITS的多态性——一个广泛逃离一致性进化的实例. 植物学报 46, 162-169. |
| 5 | 杨俊波, 李洪涛, 杨世雄, 李德铢, 杨莹燕 (2006). 四个DNA片段在山茶属分子系统学研究中的应用. 云南植物研究 28, 108-114. |
| 6 | 张宏达, 任善湘 (1998). 中国植物志(第49卷第3分册). 北京: 科学出版社. pp. 6-170. |
| 7 | Akaike H (1974). A new look at the statistical model identification.IEEE Trans Automat Contr 19, 716-723. |
| 8 | Álvarez I, Wendel JF (2003). Ribosomal ITS sequences and plant phylogenetic inference.Mol Phylogenet Evol 29, 417-434. |
| 9 | Bailey CD, Carr TG, Harris SA, Hughes CE (2003). Characterization of angiosperm nrDNA polymorphism, para- logy, and pseudogenes.Mol Phylogenet Evol 29, 435-455. |
| 10 | Baldwin BG, Sanderson MJ, Porter JM, Wojciechowski MF, Campbell CS, Donoghue MJ (1995). The ITS region of nuclear ribosomal DNA: a valuable source of evidence on angiosperm phylogeny.Ann Missouri Bot Gard 82, 247-277. |
| 11 | Buckler ES 4th, Holtsford TP (1996). Zea ribosomal repeat evolution and substitution patterns.Mol Biol Evol 13, 623-632. |
| 12 | Carranza S, Baguñà J, Riutort M (1999). Origin and evolution of paralogous rRNA gene clusters within the flatworm family Dugesiidae (Platyhelminthes, Tricladida).J Mol Evol 49, 250-259. |
| 13 | Copenhaver GP, Pikaard CS (1996). Two-dimensional RFLP analyses reveal megabase-sized clusters of rRNA gene variants in Arabidopsis thaliana, suggesting local spreading of variants as the mode for gene homogenization during concerted evolution.Plant J 9, 273-282. |
| 14 | Dover G (1982). Molecular drive: a cohesive mode of species evolution.Nature 299, 111-117. |
| 15 | Doyle JJ, Doyle JL (1987). A rapid DNA isolation procedure for small quantities of fresh leaf tissue.Phytochem Bull 19, 11-15. |
| 16 | Eickbush TH, Eickbush DG (2007). Finely orchestrated movements: evolution of the ribosomal RNA genes.Genetics 175, 477-485. |
| 17 | Ganley ARD, Kobayashi T (2007). Highly efficient concerted evolution in the ribosomal DNA repeats: total rDNA repeat variation revealed by whole-genome shotgun sequence data.Genome Res 17, 184-191. |
| 18 | Gu ZJ, Hua X (2003). Physical mapping of the 18S-26S rDNA by fluorescent in situ hybridization (FISH) in Camellia reticulata polyploid complex (Theaceae).Plant Sci 164, 279-285. |
| 19 | Hamada M, Sato K, Kiryu H, Mituyama T, Asai K (2009). Predictions of RNA secondary structure by combining homologous sequence information.Bioinformatics 25, i330-i338. |
| 20 | Hillis DM, Moritz C, Porter CA, Baker RJ (1991). Evidence for biased gene conversion in concerted evolution of ribosomal DNA.Science 251, 308-310. |
| 21 | Keller I, Chintauan-Marquier IC, Veltsos P, Nichols RA (2006). Ribosomal DNA in the grasshopper Podisma pedestris: escape from concerted evolution.Genetics 174, 863-874. |
| 22 | Kovarik A, Pires JC, Leitch AR, Lim KY, Sherwood AM, Matyasek R, Rocca J, Soltis DE, Soltis PS (2005). Rapid concerted evolution of nuclear ribosomal DNA in two Tragopogon allopolyploids of recent and recurrent origin.Genetics 169, 931-944. |
| 23 | Librado P, Rozas J (2009). DnaSP v5: a software for comprehensive analysis of DNA polymorphism data.Bioinformatics 25, 1451-1452. |
| 24 | Long EO, Dawid IB (1980). Repeated genes in eukaryotes.Annu Rev Biochem 49, 727-764. |
| 25 | Márquez LM, Miller DJ, MacKenzie JB, Van Oppen MJH (2003). Pseudogenes contribute to the extreme diversity of nuclear ribosomal DNA in the hard coral Acropora.Mol Biol Evol 20, 1077-1086. |
| 26 | Martin DP, Lemey P, Lott M, Moulton V, Posada D, Lefeuvre P (2010). RDP3: a flexible and fast computer program for analyzing recombination.Bioinformatics 26, 2462-2463. |
| 27 | Martin DP, Williamson C, Posada D (2005). RDP2: recombination detection and analysis from sequence align- ments.Bioinformatics 21, 260-262. |
| 28 | Mayol M, Rosselló JA (2001). Why nuclear ribosomal DNA spacers (ITS) tell different stories in Quercus.Mol Phylogenet Evol 19, 167-176. |
| 29 | Muir G, Fleming CC, Schlötterer C (2001). Three divergent rDNA clusters predate the species divergence in Quercus petraea (matt.) liebl. and Quercus robur L.Mol Biol Evol 18, 112-119. |
| 30 | Nylander JAA (2004). MrModeltest v2. Program distributed by the author. Evolutionary Biology Centre, Uppsala University. |
| 31 | Odorico DM, Miller DJ (1997). Variation in the ribosomal internal transcribed spacers and 5.8S rDNA among five species of Acropora (Cnidaria; Scleractinia): patterns of variation consistent with reticulate evolution.Mol Biol Evol 14, 465-473. |
| 32 | Padidam M, Sawyer S, Fauquet CM (1999). Possible emergence of new geminiviruses by frequent recombination.Virology 265, 218-225. |
| 33 | Parkin EJ, Butlin RK (2004). Within- and between- individual sequence variation among ITS1 copies in the meadow grasshopper Chorthippus parallelus indicates frequent intrachromosomal gene conversion.Mol Biol Evol 21, 1595-1601. |
| 34 | Parks CR (1990). Cross-compatibility studies in the genus Camellia. A presentation for the ICS congress at Maizura.Int Camellia J 22, 37-54. |
| 35 | Pedersen C, Linde-Laursen I (1994). Chromosomal locations of four minor rDNA loci and a marker microsatellite sequence in barley.Chromosome Res 2, 65-71. |
| 36 | Posada D, Crandall KA (2001). Evaluation of methods for detecting recombination from DNA sequences: computer simulations.Proc Natl Acad Sci USA 98, 13757-13762. |
| 37 | Prokopowich CD, Gregory TR, Crease TJ (2003). The correlation between rDNA copy number and genome size in eukaryotes.Genome 46, 48-50. |
| 38 | Rannala B, Yang ZH (1996). Probability distribution of molecular evolutionary trees: a new method of phylogenetic inference.J Mol Evol 43, 304-311. |
| 39 | Ronquist F, Huelsenbeck JP (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models.Bioinformatics 19, 1572-1574. |
| 40 | Saitou N, Nei M (1987). The neighbor-joining method: a new method for reconstructing phylogenetic trees.Mol Biol Evol 4, 406-425. |
| 41 | Schlötterer C, Tautz D (1994). Chromosomal homogeneity of Drosophila ribosomal DNA arrays suggests intrachromosomal exchanges drive concerted evolution.Curr Biol 4, 777-783. |
| 42 | Smith JM (1992). Analyzing the mosaic structure of genes.J Mol Evol 34, 126-129. |
| 43 | Swofford DL (2002). PAUP*: Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer Associates, Sunderland, MA. |
| 44 | Swofford DL, Berlocher SH (1987). Inferring evolutionary trees from gene frequency data under the principle of maximum parsimony.Syst Zool 36, 293-325. |
| 45 | Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods.Mol Biol Evol 28, 2731-2739. |
| 46 | Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997). The CLUSTAL_X windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools.Nucleic Acids Res 25, 4876-4882. |
| 47 | Van Oppen MJH, Willis BL, Van Rheede T, Miller DJ (2002). Spawning times, reproductive compatibilities and genetic structuring in the Acropora aspera group: evidence for natural hybridization and semi-permeable species boundaries in corals.Mol Ecol 11, 1363-1376. |
| 48 | Vijayan K, Zhang WJ, Tsou CH (2009). Molecular taxo- nomy of Camellia (Theaceae) inferred from nrITS sequences.Am J Bot 96, 1348-1360. |
| 49 | Xiao LQ, Möller M, Zhu H (2010). High nrDNA ITS polymorphism in the ancient extant seed plant Cycas: incomplete concerted evolution and the origin of pseudogenes.Mol Phylogenet Evol 55, 168-177. |
| 50 | Xu JP, Zhang QQ, Xu XF, Wang ZG, Qi J (2009). Intragenomic variability and pseudogenes of ribosomal DNA in Stone flounder Kareius bicoloratus.Mol Phylogenet Evol 52, 157-166. |
| 51 | Young ND, Healy J (2003). GapCoder automates the use of indel characters in phylogenetic analysis.BMC Bioinformatics 4, 6. |
| 52 | Zheng XY, Cai DY, Yao LH, Teng YW (2008). Non-concerted ITS evolution, early origin and phylogenetic utility of ITS pseudogenes in Pyrus.Mol Phylogenet Evol 48, 892-903. |
| 53 | Zimmer EA, Martin SL, Beverley SM, Kan YW, Wilson AC (1980). Rapid duplication and loss of genes coding for the alpha chains of hemoglobin.Proc Natl Acad Sci USA 77, 2158-2162. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||