

植物卷须发生及调控机制研究进展
收稿日期: 2025-04-11
录用日期: 2025-07-29
网络出版日期: 2025-07-30
基金资助
国家重点研发计划(2024YFF1307604)
Research Progress in the Development and Regulatory Mechanisms of Plant Tendrils
Received date: 2025-04-11
Accepted date: 2025-07-29
Online published: 2025-07-30
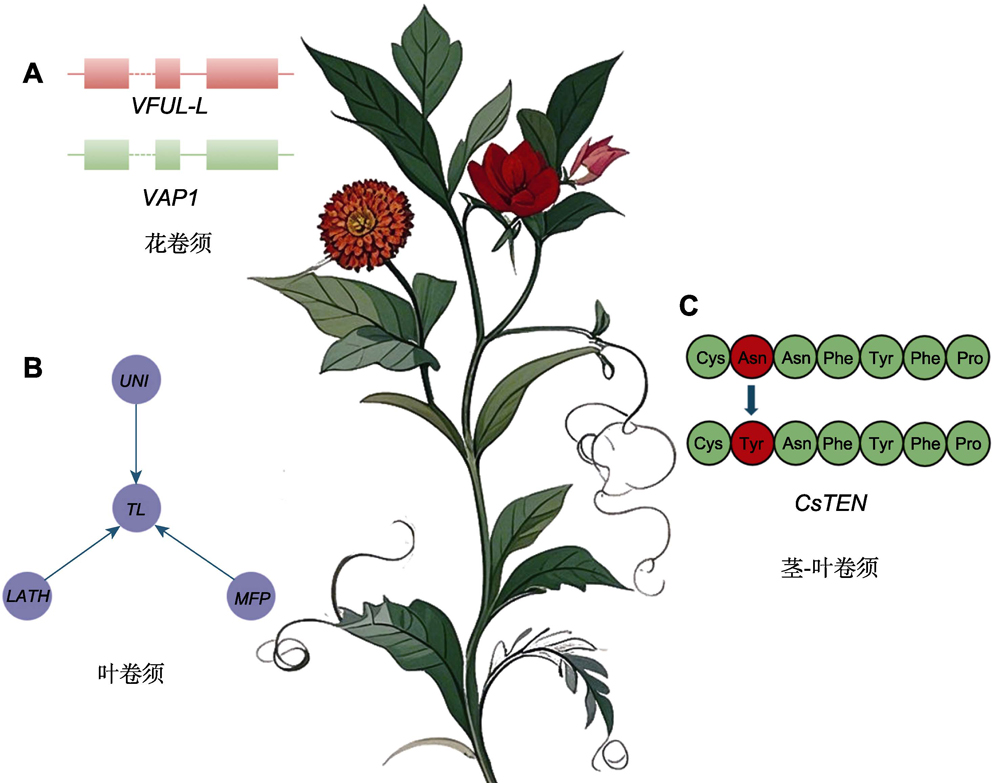
植物卷须是一种特化的攀缘器官, 在植物的生存与环境适应中发挥关键作用。通过提供结构支撑、增强光能捕获能力以及降低地面资源竞争, 卷须显著提升了植物的生态适应性。卷须由花序、叶片和枝茎等器官衍生而来, 由TCP、HD-ZIP和MADS-box等基因家族调控, 并受到生长素、赤霉素、细胞分裂素和茉莉酸等植物激素的影响。卷须在功能、形态及分子机制上表现出趋同进化现象, 并呈现独立演化特征, 这反映了植物对生存环境的适应策略。该文系统综述了植物卷须的生物学特性和发育的分子机制, 展望了未来卷须形成与调控机制研究应关注跨物种的演化机制、环境信号与植物激素的相互调控等方面。
罗号东 , 刘勇波 . 植物卷须发生及调控机制研究进展[J]. 植物学报, 2025 , 60(6) : 993 -1004 . DOI: 10.11983/CBB25064
Tendrils are specialized climbing organs that play a key role in the survival and environmental adaptation of plants. Through structural support, light capture, and resources competition, tendrils significantly improve the ecological adaptability of plants. Tendrils are derived from inflorescences, leaves, stems and other organs. The occurrence of tendrils is regulated by gene families such as TCP, HD-ZIP, and MADS-box families, and is influenced by auxin, gibberellin, cytokinin, jasmonic acid and other plant hormones. Tendrils exhibit convergent evolution in function, morphology, and molecular mechanisms, and display characteristics of independent evolution, reflecting the adaptive strategies of plants to their living environment. This review systematically synthesizes the biological characteristics and molecular mechanisms of development in plant tendrils. Future studies will focus on evolutionary mechanisms across species and interactions between environmental signals and plant hormones for plant tendrils.

| [1] | Allfrey VG, Faulkner R, Mirsky AE (1964). Acetylation and methylation of histones and their possible role in the regulation of RNA synthesis. Proc Natl Acad Sci USA 51, 786-794. |
| [2] | Ameha M, Skirvin RM, Mitiku G, Bullock D, Hofmann P (1998). In vitro tendril and flower development in cucumber (Cucumis sativus) may be regulated by gibberellins. J Hortic Sci Biotechnol 73, 159-163. |
| [3] | Bai F, DeMason DA (2006). Hormone interactions and regulation of Unifoliata, PsPK2, PsPIN1 and LE gene expression in pea (Pisum sativum) shoot tips. Plant Cell Physiol 47, 935-948. |
| [4] | Bharathan G, Goliber TE, Moore C, Kessler S, Pham T, Sinha NR (2002). Homologies in leaf form inferred from KNOXI gene expression during development. Science 296, 1858-1860. |
| [5] | Boss PK, Thomas MR (2002). Association of dwarfism and floral induction with a grape ‘green revolution’ mutation. Nature 416, 847-850. |
| [6] | Bucher P (1990). Weight matrix descriptions of four eukaryotic RNA polymerase II promoter elements derived from 502 unrelated promoter sequences. J Mol Biol 212, 563-578. |
| [7] | Calonje M, Cubas P, Martínez-Zapater JM, Carmona MJ (2004). Floral meristem identity genes are expressed during tendril development in grapevine. Plant Physiol 135, 1491-1501. |
| [8] | Carmona MJ, Cubas P, Calonje M, Martínez-Zapater JM (2007). Flowering transition in grapevine (Vitis vinifera L.). Can J Bot 85, 701-711. |
| [9] | Chen FF, Fu BB, Pan YP, Zhang CW, Wen HF, Weng YQ, Chen P, Li YH (2017). Fine mapping identifies CsGCN5 encoding a histone acetyltransferase as putative candidate gene for tendril-less1 mutation (td-1) in cucumber. Theor Appl Genet 130, 1549-1558. |
| [10] | Chen JY, Wang WQ, Luo SY, Yang LX, Wang HJ, Wu TY, Zhu QK (2025). Expression pattern and metabolic correlation analysis of TCP gene family in Bergenia purpurascens. Chin Bull Bot doi: 10.11983/CBB25008. (in Chinese) |
| 陈靖彧, 王文庆, 罗诗语, 杨路祥, 汪慧骏, 吴天宇, 朱乾坤 (2025). 岩白菜TCP基因家族的表达模式及代谢关联分析. 植物学报 doi: 10.11983/CBB25008. | |
| [11] | Chen Y, Wen HF, Pan J, Du H, Zhang KY, Zhang LY, Yu Y, He HL, Cai R, Pan JS, Wang G (2021). CsUFO is involved in the formation of flowers and tendrils in cucumber. Theor Appl Genet 134, 2141-2150. |
| [12] | Cutri L (2009). Estudo da Fun??o do Gene LEAFY (LFY) em Duas Especies de Passiflora. Master’s thesis. Campinas, SP: Universidade Estadual de Campinas. |
| [13] | Cutri L, Dornelas MC (2012). PASSIOMA: exploring expressed sequence tags during flower development in Passiflora spp. Comp Funct Genomics 2012, 510549. |
| [14] | Dai YJ, Luo XF, Zhou WG, Chen F, Shuai HY, Yang WY, Shu K (2019). Plant systemic signaling under biotic and abiotic stresses conditions. Chin Bull Bot 54, 255-264. (in Chinese) |
| 代宇佳, 罗晓峰, 周文冠, 陈锋, 帅海威, 杨文钰, 舒凯 (2019). 生物和非生物逆境胁迫下的植物系统信号. 植物学报 54, 255-264. | |
| [15] | DeMason DA, Chetty V, Barkawi LS, Liu X, Cohen JD (2013). Unifoliata-Afila interactions in pea leaf morphogenesis. Am J Bot 100, 478-495. |
| [16] | DeMason DA, Chetty VJ (2011). Interactions between GA, auxin, and UNI expression controlling shoot ontogeny, leaf morphogenesis, and auxin response in Pisum sativum (Fabaceae): or how the uni-tac mutant is rescued. Am J Bot 98, 775-791. |
| [17] | Eshed Y, Baum SF, Perea JV, Bowman JL (2001). Establishment of polarity in lateral organs of plants. Curr Biol 11, 1251-1260. |
| [18] | Gélinas-Marion A, Eléou?t MP, Cook SD, Vander Schoor JK, Abel SAG, Nichols DS, Smith JA, Hofer JMI, Ross JJ (2023). Plant development in the garden pea as revealed by mutations in the Crd/PsYUC1 gene. Genes 14, 2115. |
| [19] | Gourlay CW, Hofer JM, Ellis TH (2000). Pea compound leaf architecture is regulated by interactions among the genes UNIFOLIATA, COCHLEATA, AFILA, and TENDRIL-LESS. Plant Cell 12, 1279-1294. |
| [20] | Guo J, Xu WB, Hu Y, Huang J, Zhao YY, Zhang L, Huang CH, Ma H (2020). Phylotranscriptomics in Cucurbitaceae reveal multiple whole-genome duplications and key morphological and molecular innovations. Mol Plant 13, 1117-1133. |
| [21] | Han XR, Yuan X, Gao J, Wang LS, Wang XH, Jia WQ, Fu ZZ, Zhang HC (2025). Identification of TCP gene family and functional analysis of TCP15a of tree peony. Chin Bull Bot doi: 10.11983/CBB25005. (in Chinese) |
| 韩新蕊, 袁欣, 高杰, 王亮生, 王晓晖, 贾文庆, 符真珠, 张和臣 (2025). 牡丹TCP基因家族鉴定及TCP15a功能分析. 植物学报 doi: 10.11983/CBB25005. | |
| [22] | Hofer J, Turner L, Hellens R, Ambrose M, Matthews P, Michael A, Ellis N (1997). UNIFOLIATA regulates leaf and flower morphogenesis in pea. Curr Biol 7, 581-587. |
| [23] | Hofer J, Turner L, Moreau C, Ambrose M, Isaac P, Butcher S, Weller J, Dupin A, Dalmais M, Le Signor C, Bendahmane A, Ellis N (2009). Tendril-less regulates tendril formation in pea leaves. Plant Cell 21, 420-428. |
| [24] | Hong ZZ, Wang XR, Fan ZP, Wang JH, Yang AY, Yan GC, He Y, Wang HS, Zhu ZJ, Xu YM (2024). The intrinsic developmental age signal defines an age-dependent climbing behavior in cucumber. Hortic Plant J 10, 797-808. |
| [25] | Hong ZZ, Wang XR, Yang AY, Yan GC, He Y, Zhu ZJ, Xu YM (2023). Tendril morphogenesis is regulated by a CsaTEN-CsaUFO module in cucumber. New Phytol 239, 364-373. |
| [26] | Immink RG, Tonaco IA, De Folter S, Shchennikova A, Van Dijk AD, Busscher-Lange J, Borst JW, Angenent GC (2009). SEPALLATA3: the ‘glue’ for MADS box transcription factor complex formation. Genome Biol 10, R24. |
| [27] | Jaffe MJ, Galston AW (1968). The physiology of tendrils. Annu Rev Plant Biol 19, 417-434. |
| [28] | Jiang YX, Zhang AR, He WJ, Li QQ, Zhao BS, Zhao HJ, Ke XB, Guo YL, Sun PY, Yang TW, Wang Z, Jiang B, Shen JJ, Li Z (2023). GRAS family member LATERAL SUPPRESSOR regulates the initiation and morphogenesis of watermelon lateral organs. Plant Physiol 193, 2592-2604. |
| [29] | Kim M, Pham T, Hamidi A, McCormick S, Kuzoff RK, Sinha N (2003). Reduced leaf complexity in tomato wiry mutants suggests a role for PHAN and KNOX genes in generating compound leaves. Development 130, 4405-4415. |
| [30] | Kosugi S, Ohashi Y (2002). DNA binding and dimerization specificity and potential targets for the TCP protein family. Plant J 30, 337-348. |
| [31] | Kotoda N, Wada M, Komori S, Kidou S, Abe K, Masuda T, Soejima J (2000). Expression pattern of homologues of floral meristem identity genes LFY and AP1 during flower development in apple. J Am Soc Hortic Sci 125, 398-403. |
| [32] | Krings M, Kerp H, Taylor TN, Taylor EL (2003). How Paleozoic vines and lianas got off the ground: on scrambling and climbing carboniferous-early permian pteridosperms. Bot Rev 69, 204-224. |
| [33] | Kumar S, Kumar Rai S, Pandey-Rai S, Srivastava S, Singh D (2004). Regulation of unipinnate character in the distal tendrilled domain of compound leaf-blade by the gene MULTIFOLIATE PINNA (MFP) in pea Pisum sativum. Plant Sci 166, 929-940. |
| [34] | Lescot M (2002). PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res 30, 325-327. |
| [35] | Li BQ, Yan JH, Li H, Xin W, Tian YH, Yang ZB, Tang WX (2022). Changes of small GTPases activity during cucumber tendril Winding. Chin Bull Bot 57, 299-307. (in Chinese) |
| 李彬琪, 闫佳慧, 李豪, 辛伟, 田云鹤, 杨贞标, 唐文鑫 (2022). 黄瓜卷须缠绕过程中小G蛋白活性变化. 植物学报 57, 299-307. | |
| [36] | Li CJ, Bangerth F (2003). Stimulatory effect of cytokinins and interaction with IAA on the release of lateral buds of pea plants from apical dominance. J Plant Physiol 160, 1059-1063. |
| [37] | Li YY, Qi YH (2022). Advances in biological functions of Aux/IAA gene family in plants. Chin Bull Bot 57, 30-41. (in Chinese) |
| 李艳艳, 齐艳华 (2022). 植物Aux/IAA基因家族生物学功能研究进展. 植物学报 57, 30-41. | |
| [38] | Lin H, Niu LF, McHale NA, Ohme-Takagi M, Mysore KS, Tadege M (2013). Evolutionarily conserved repressive activity of WOX proteins mediates leaf blade outgrowth and floral organ development in plants. Proc Natl Acad Sci USA 110, 366-371. |
| [39] | Liu XF, Hao N, Li HY, Ge DF, Du YL, Liu RY, Wen CL, Li YH, Zhang XL, Wu T (2019). PINOID is required for lateral organ morphogenesis and ovule development in cucumber. J Exp Bot 70, 5715-5730. |
| [40] | López-Juez E, Buurmeijer WF, Heeringa GH, Kendrick RE, Wesselius JC (1990). Response of light-grown wild- type and long-hypocotyl mutant cucumber plants to end- of-day far-red light. Photochem Photobiol 52, 143-149. |
| [41] | Mishra RK, Chaudhary S, Kumar A, Kumar S (2009). Effects of MULTIFOLIATE-PINNA, AFILA, TENDRIL-LESS and UNIFOLIATA genes on leafblade architecture in Pisum sativum. Planta 230, 177-190. |
| [42] | Mizuno S, Sonoda M, Tamura Y, Nishino E, Suzuki H, Sato T, Oizumi T (2015). Chiba Tendril-Less locus determines tendril organ identity in melon (Cucumis melo L.) and potentially encodes a tendril-specific TCP homolog. J Plant Res 128, 941-951. |
| [43] | Narasimhan S (2024). Ultrastructural studies on tendrils of plant climbers reveals a hierarchical tissue organization: a microscopic investigation. J Integr Sci Technol 12, 760. |
| [44] | Nicolas M, Cubas P (2015). The role of TCP transcription factors in shaping flower structure, leaf morphology, and plant architecture. In: Gonzalez DH, ed. Plant Transcription Factors: Evolutionary, Structural and Functional Aspects, Amsterdam: Academic Press. pp. 249-267. |
| [45] | Pajoro A, Biewers S, Dougali E, Leal Valentim F, Mendes MA, Porri A, Coupland G, Van de Peer Y, van Dijk ADJ, Colombo L, Davies B, Angenent GC (2014). The (r)evolution of gene regulatory networks controlling Arabidopsis plant reproduction: a two-decade history. J Exp Bot 65, 4731-4745. |
| [46] | Pratt C (1974). Vegetative anatomy of cultivated grapes—a review. Am J Enol Vitic 25, 131-150. |
| [47] | Prenner G (2014). Floral ontogeny in Passiflora lobata (Malpighiales, Passifloraceae) reveals a rare pattern in petal formation and provides new evidence for interpretation of the tendril and corona. Plant Syst Evol 300, 1285-1297. |
| [48] | Scorza LCT, Hernandes-Lopes J, Melo-de-Pinna GFA, Dornelas MC (2017). Expression patterns of Passiflora edulis APETALA1/FRUITFULL homologues shed light onto tendril and corona identities. EvoDevo 8, 3. |
| [49] | Shen JJ, Jiang YX, Pan J, Sun LH, Li QQ, He WJ, Sun PY, Zhao BS, Zhao HJ, Ke XB, Guo YL, Yang TW, Li Z (2024). The GRAS transcription factor CsTL regulates tendril formation in cucumber. Plant Cell 36, 2818-2833. |
| [50] | Shuai HY, Meng YJ, Chen F, Zhou WG, Luo XF, Yang WY, Shu K (2018). Phytohormone-mediated plant shade responses. Chin Bull Bot 53, 139-148. (in Chinese) |
| 帅海威, 孟永杰, 陈锋, 周文冠, 罗晓峰, 杨文钰, 舒凯 (2018). 植物荫蔽胁迫的激素信号响应. 植物学报 53, 139-148. | |
| [51] | Song MF, Zha GH, Chen JF, Lou QF (2022). Research progress on molecular basis of plant architecture related traits in cucumber. Acta Hortic Sin 49, 2683-2702. (in Chinese) |
| 宋蒙飞, 查高辉, 陈劲枫, 娄群峰 (2022). 黄瓜株型性状分子基础研究进展. 园艺学报 49, 2683-2702. | |
| [52] | Sousa-Baena MS, Lohmann LG, Hernandes-Lopes J, Sinha NR (2018a). The molecular control of tendril development in angiosperms. New Phytol 218, 944-958. |
| [53] | Sousa-Baena MS, Lohmann LG, Rossi M, Sinha NR (2014a). Acquisition and diversification of tendrilled leaves in Bignonieae (Bignoniaceae) involved changes in expression patterns of SHOOTMERISTEMLESS (STM), LEAFY/ FLORICAULA (LFY/FLO), and PHANTASTICA (PHAN). New Phytol 201, 993-1008. |
| [54] | Sousa-Baena MS, Sinha NR, Hernandes-Lopes J, Lohmann LG (2018b). Convergent evolution and the diverse ontogenetic origins of tendrils in angiosperms. Front Plant Sci 9, 403. |
| [55] | Sousa-Baena MS, Sinha NR, Lohmann LG (2014b). Evolution and development of tendrils in Bignonieae (Lamiales, Bignoniaceae). Ann Mo Bot Gard 99, 323-347. |
| [56] | Srinivasan C, Mullins MG (1976). Reproductive anatomy of the grape-vine (Vitis vinifera L.): origin and development of the anlage and its derivatives. Ann Bot 40, 1079-1084. |
| [57] | Srinivasan C, Mullins MG (1978). Control of flowering in the grapevine (Vitis vinifera L.): formation of inflorescences in vitro by isolated tendrils. Plant Physiol 61, 127-130. |
| [58] | Sung SK, Yu GH, An G (1999). Characterization of MdMADS2, a member of the SQUAMOSA subfamily of genes, in apple. Plant Physiol 120, 969-978. |
| [59] | Tadege M, Lin H, Bedair M, Berbel A, Wen JQ, Rojas CM, Niu LF, Tang YH, Sumner L, Ratet P, McHale NA, Madue?o F, Mysore KS (2011). STENOFOLIA regulates blade outgrowth and leaf vascular patterning in Medicago truncatula and Nicotiana sylvestris. Plant Cell 23, 2125-2142. |
| [60] | Tian L, Chen ZJ (2001). Blocking histone deacetylation in Arabidopsis induces pleiotropic effects on plant gene regulation and development. Proc Natl Acad Sci USA 98, 200-205. |
| [61] | Turner BM (1991). Histone acetylation and control of gene expression. J Cell Sci 99, 13-20. |
| [62] | Vaughn KC, Bowling AJ (2010). Biology and physiology of vines. In: Janick J, ed. Horticultural Reviews. NJ: John Wiley & Sons Inc. pp. 1-21. |
| [63] | Viola IL, Reinheimer R, Ripoll R, Manassero NGU, Gonzalez DH (2012). Determinants of the DNA binding specificity of class I and class II TCP transcription factors. J Biol Chem 287, 347-356. |
| [64] | Waites R, Hudson A (1995). Phantastica: a gene required for dorsoventrality of leaves in Antirrhinum majus. Development 121, 2143-2154. |
| [65] | Walton EF, Podivinsky E, Wu RM (2001). Bimodal patterns of floral gene expression over the two seasons that kiwifruit flowers develop. Physiol Plant 111, 396-404. |
| [66] | Wang SH, Yang XY, Xu MN, Lin XZ, Lin T, Qi JJ, Shao GJ, Tian NN, Yang Q, Zhang ZH, Huang SW (2015). A rare SNP identified a TCP transcription factor essential for tendril development in cucumber. Mol Plant 8, 1795-1808. |
| [67] | Wang SW, Yang AY, Wang HS, Xu YM (2021). Identification and expression analysis of miR156/157-SPL pathway genes in cucumber. Acta Hortic Sin 48, 2227-2238. (in Chinese) |
| 汪淑雯, 杨爱怡, 王华森, 徐云敏 (2021). 黄瓜miR156/ 157-SPL途径基因鉴定和表达分析. 园艺学报 48, 2227-2238. | |
| [68] | Weigel D, Alvarez J, Smyth DR, Yanofsky MF, Meyerowitz EM (1992). LEAFY controls floral meristem identity in Arabidopsis. Cell 69, 843-859. |
| [69] | Weiler EW, Albrecht T, Groth B, Xia ZQ, Luxem M, Li? H, Andert L, Spengler P (1993). Evidence for the involvement of jasmonates and their octadecanoid precursors in the tendril coiling response of Bryonia dioica. Phytochemistry 32, 591-600. |
| [70] | Yan CH, Li GR, Hu YS, Zhou RJ, Hu HL, Miao WD, Zhu ZG (2017). Cloning and expression analysis of CIPK genes in grapevine. Acta Hortic Sin 44, 1463-1476. (in Chinese) |
| 闫朝辉, 李桂荣, 扈岩松, 周瑞金, 扈惠灵, 苗卫东, 朱自果 (2017). 欧洲葡萄中CIPK基因的克隆及表达分析. 园艺学报 44, 1463-1476. | |
| [71] | Yang XY, Yan JB, Zhang Z, Lin T, Xin TX, Wang BW, Wang SH, Zhao JC, Zhang ZH, Lucas WJ, Li GH, Huang SW (2020). Regulation of plant architecture by a new histone acetyltransferase targeting gene bodies. Nat Plants 6, 809-822. |
| [72] | Yi LC, Zhou W, Zhou QL, Chen ZB, Zhang Y, Dai ZY, Wang YQ (2023). Fine mapping identifies ClTFL1 encodes a TERMINAL FLOWER 1 protein as putative candidate gene for inflorescence architecture and tendril development and in watermelon. J Plant Growth Regul 42, 4150-4160. |
| [73] | Yuan Y, Enhebayaer, Qi YH (2023). Research advances in biological functions of GH3 gene family in plants. Chin Bull Bot 58, 770-782. (in Chinese) |
| 园园, 恩和巴雅尔, 齐艳华 (2023). 植物GH3基因家族生物学功能研究进展. 植物学报 58, 770-782. | |
| [74] | Zeng K, Wang SW, Zhou BY, Wang HS, Xu YM (2020). Research progress on tendril formation in Cucurbitaceae horticultural crops. J Zhejiang A F Univ 37, 1216-1223. (in Chinese) |
| 曾康, 汪淑雯, 周冰莹, 王华森, 徐云敏 (2020). 葫芦科园艺作物卷须发生的研究进展. 浙江农林大学学报 37, 1216-1223. | |
| [75] | Zhang SY, Qi Q, Li XH, Gai Y, Jiang XN (2017). Simultaneous determination of jasmonic acid and 12-oxo-phytodienoic acid in different plant materials by HPLC-MS/ MS. Chin Bull Bot 52, 631-636. (in Chinese) |
| 张双玉, 齐芪, 李晓慧, 盖颖, 蒋湘宁 (2017). 茉莉酸及其前体12-氧-植物二烯酸的HPLC-MS/MS检测方法. 植物学报 52, 631-636. | |
| [76] | Zhang Y, Zhang YC, Li B, Tan X, Zhu CP, Wu T, Feng SY, Yang QH, Shen SQ, Yu T, Liu Z, Song XM (2022). Polyploidy events shaped the expansion of transcription factors in Cucurbitaceae and exploitation of genes for tendril development. Hortic Plant J 8, 562-574. |
| [77] | Zhang YP, Mu Q, Li XP, Wang C, Song CN, Fang JG (2013). Advances of studies on grape tendrils. Plant Phys J 49, 234-240. (in Chinese) |
| 张彦苹, 慕茜, 李晓鹏, 王晨, 宋长年, 房经贵 (2013). 葡萄卷须及其相关研究. 植物生理学报 49, 234-240. | |
| [78] | Zhuang D, Ding F, Wu J, Gan DF (2014). Expression of CsaIAAs gene in cucumber induced by 6-BA treatment. J Anhui Agric Univ 41, 260-264. (in Chinese) |
| 庄丹, 丁飞, 吴娇, 甘德芳 (2014). 6-BA诱导黄瓜CsaIAAs基因的表达研究. 安徽农业大学学报 41, 260-264. | |
| [79] | Zhuang LL, Ambrose M, Rameau C, Weng L, Yang J, Hu XH, Luo D, Li X (2012). LATHYROIDES, encoding a WUSCHEL-related homeobox1 transcription factor, controls organ lateral growth, and regulates tendril and dorsal petal identities in garden pea (Pisum sativum L.). Mol Plant 5, 1333-1345. |
/
| 〈 |
|
〉 |