

Chinese Bulletin of Botany ›› 2023, Vol. 58 ›› Issue (4): 590-601.DOI: 10.11983/CBB22102 cstr: 32102.14.CBB22102
• TECHNIQUES AND METHODS • Previous Articles Next Articles
Pengyun Luo1,2,3,4, Hongping Qian1,2,3,4, Yan Liu5, Changwen Xu1,2,3,4, Yaning Cui1,2,3,4( )
)
Received:2022-06-06
Accepted:2022-11-12
Online:2023-07-01
Published:2022-11-29
Contact:
*E-mail: cuiyaning@bjfu.edu.cn
Pengyun Luo, Hongping Qian, Yan Liu, Changwen Xu, Yaning Cui. Regulation of Plasma Membrane Protein Dynamics and Its Research Methods[J]. Chinese Bulletin of Botany, 2023, 58(4): 590-601.
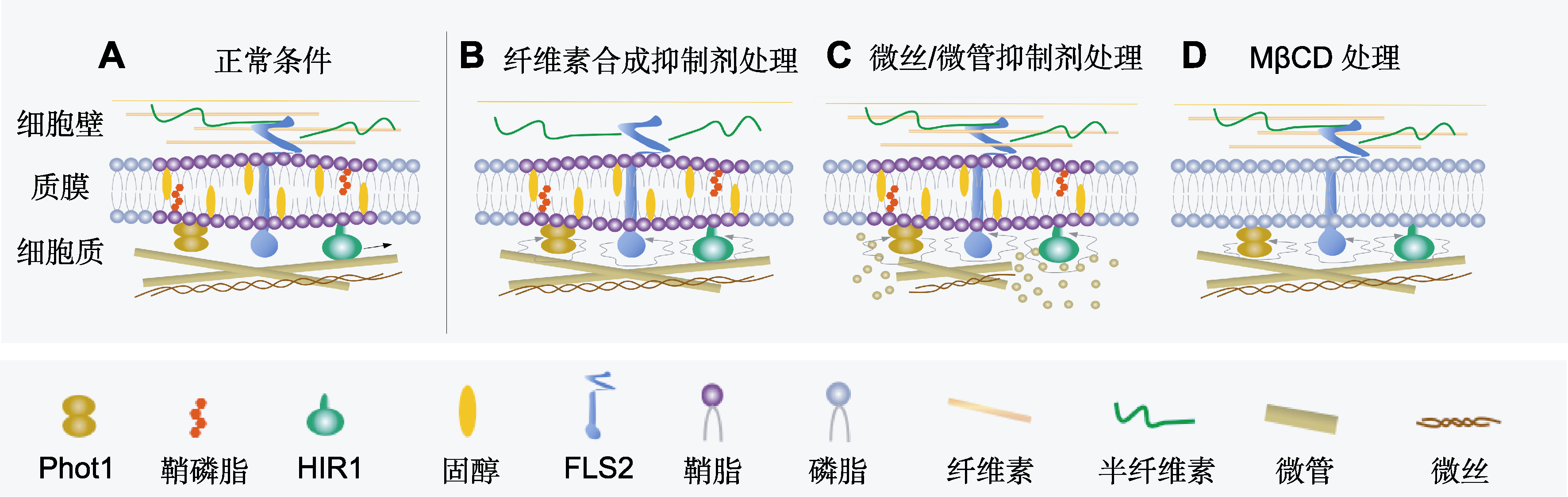
Figure 1 Factors affecting the dynamics of plasma membrane protein (A) Plasma membrane proteins are relatively stationary on the membrane under normal conditions; (B) The lateral diffusion of plasma membrane proteins was changed when the cell wall was damaged by cellulose synthesis inhibitors; (C) Microfilament/microtubule inhibitor treatment disrupts the cytoskeleton and changes the lateral diffusion movement of plasma membrane proteins; (D) MβCD treatment destroys the membrane raft and changes the lateral diffusion of plasma membrane proteins.
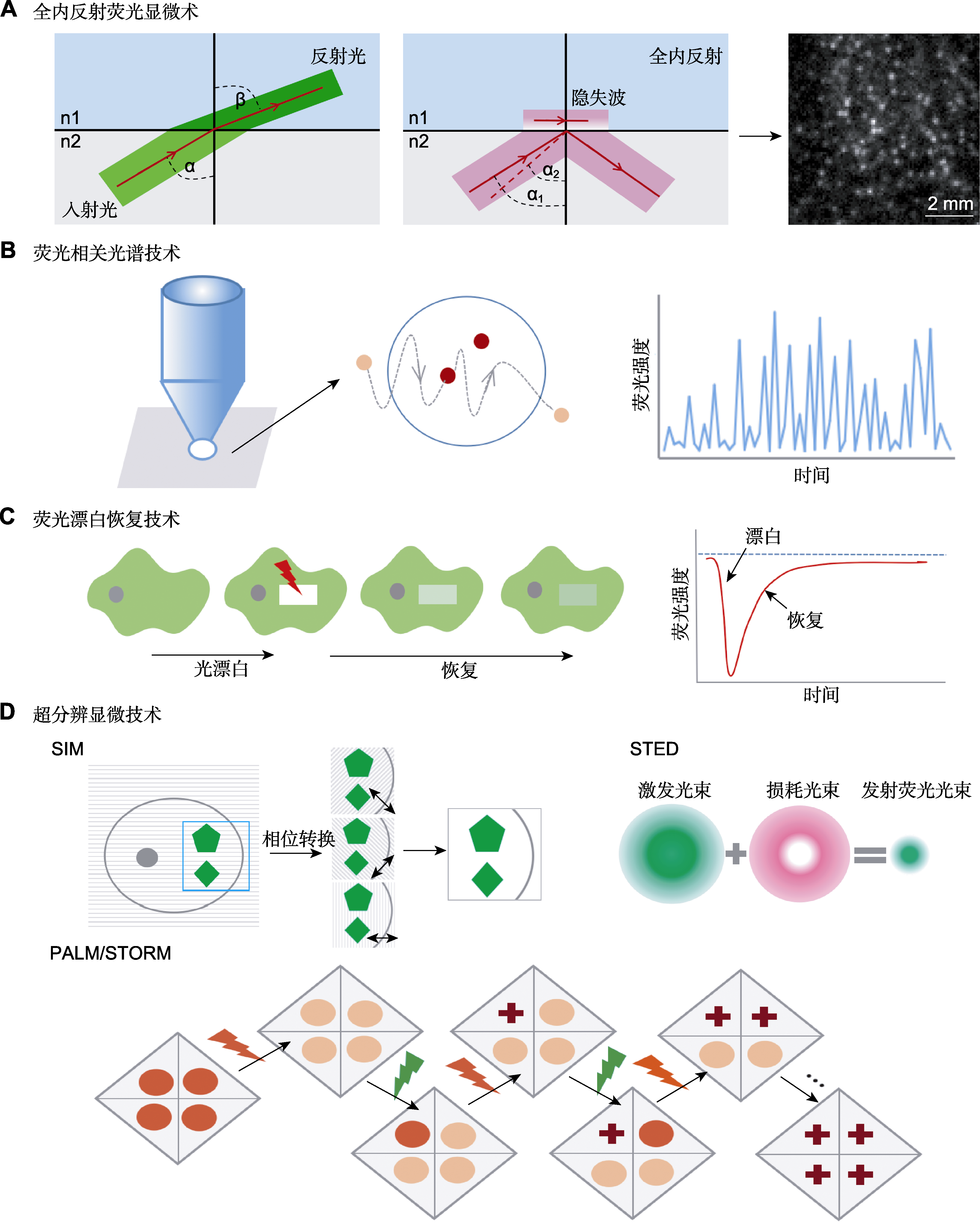
Figure 2 Technology of plasma membrane protein dynamics (A) From refraction to total internal reflection (TIR): the incident beam approaching the interface between two media with different indices of refraction n1 and n2 at an angle α will be refracted according to Snell’s law, beam propagation after the interface occurs thus at a different angle, if the incident angle α1 is greater than or equal to the critical angle α2, the beam does not propagate in the optically less-dense medium, but instead undergoes total reflection at the interface; (B) The diffusion of single labeled molecules in and out of the focal volume in fluorescence correlation spectroscopy (FCS) induces fluctuations in fluorescence with time, the number of particles and their diffusion coefficients can be calculated by fitting the fluorescence fluctuation curve to an adequate diffusion model; (C) A region of interest (ROI) is selected bleached with an intense laser beam, and the fluorescence recovery in the ROI is measured over time; (D) Schematic diagram of the principles of super-resolution microscopic imaging techniques. FCS, SIM, STED, PALM and STORM see Table 1.
| 技术名称 | 适用对象 | 优点 | 缺点 |
|---|---|---|---|
| TIRFM | 膜蛋白和脂类等分子 | 只激发靠近界面100-200 nm处的荧光分子, 信噪比显著提高 | 成像受衍射极限的限制, 且轴向分辨率局限于宽场范围 |
| FCS | 可自由扩散的荧光分子 | 灵敏度高, 空间分辨率高, 非侵入性检测等 | 不适用于高浓度和扩散速度慢的样本, 高激发功率下荧光团会出现光漂白 |
| FRAP | 活细胞内分子 | 可用于功能性测量活细胞中的蛋白质迁移率 | 只能检测荧光分子的群体移动, 不能观察单个分子的移动 |
| SIM | 活细胞内分子 | 成像速度快, 普适性强, 无须特殊的染料标记 | 硬件装置昂贵, 成像时对光条件、原始数据的质量和数量要求严苛, 捕捉动态性较高的目标时会出现引入动态伪影的问题, 对活细胞内动态性较高的亚细胞结构长时程成像存在明显的局限性 |
| STED | 活体生物样品 | 荧光探针无光开关的特殊要求, 时间分辨率高 | |
| PALM | 细胞外源表达的荧光蛋白 | 可用于活细胞测量, 分辨率可达1 nm量级 | |
| STORM | 内源性蛋白物质 | 可研究细胞内源性蛋白的超分辨率定位 |
Table 1 Comparison of methods on plasma membrane protein dynamics research
| 技术名称 | 适用对象 | 优点 | 缺点 |
|---|---|---|---|
| TIRFM | 膜蛋白和脂类等分子 | 只激发靠近界面100-200 nm处的荧光分子, 信噪比显著提高 | 成像受衍射极限的限制, 且轴向分辨率局限于宽场范围 |
| FCS | 可自由扩散的荧光分子 | 灵敏度高, 空间分辨率高, 非侵入性检测等 | 不适用于高浓度和扩散速度慢的样本, 高激发功率下荧光团会出现光漂白 |
| FRAP | 活细胞内分子 | 可用于功能性测量活细胞中的蛋白质迁移率 | 只能检测荧光分子的群体移动, 不能观察单个分子的移动 |
| SIM | 活细胞内分子 | 成像速度快, 普适性强, 无须特殊的染料标记 | 硬件装置昂贵, 成像时对光条件、原始数据的质量和数量要求严苛, 捕捉动态性较高的目标时会出现引入动态伪影的问题, 对活细胞内动态性较高的亚细胞结构长时程成像存在明显的局限性 |
| STED | 活体生物样品 | 荧光探针无光开关的特殊要求, 时间分辨率高 | |
| PALM | 细胞外源表达的荧光蛋白 | 可用于活细胞测量, 分辨率可达1 nm量级 | |
| STORM | 内源性蛋白物质 | 可研究细胞内源性蛋白的超分辨率定位 |
| [1] | 李晓娟 (2012). 拟南芥质膜水通道蛋白活性调控多重机制的单分子研究. 博士论文. 北京: 中国科学院大学. pp. 1-93. |
| [2] |
刘玥, 尹悦佳, 梁重阳, 黄殿帅, 王阳, 刘艳芝, 窦瑶, 冯树丹, 郝东云 (2015). 3D-SIM结构照明超分辨率显微镜实现蛋白质在植物亚细胞器内的定位. 植物学报 50, 495-503.
DOI |
| [3] |
钱虹萍, 陈博, 林金星, 崔亚宁 (2021). RNA聚合酶II动态调控及其成像技术的研究进展. 生物技术通报 37, 293-302.
DOI |
| [4] |
邱丽丽, 赵琪, 张玉红, 戴绍军 (2017). 植物质膜蛋白质组的逆境应答研究进展. 植物学报 52, 128-147.
DOI |
| [5] |
苏泊丹, 林金星, 肖建伟 (2016). 泛素介导的植物膜蛋白转运及其研究方法. 植物学报 51, 387-395.
DOI |
| [6] |
Ananthanarayanan V (2021). Cytoskeletal mechanics. J Indian Inst Sci 101, 3.
DOI |
| [7] |
Chu TTH, Hoang TG, Trinh DC, Bureau C, Meynard D, Vernet A, Ingouff M, Do NV, Périn C, Guiderdoni E, Gantet P, Maurel C, Luu DT (2018). Sub-cellular markers highlight intracellular dynamics of membrane proteins in response to abiotic treatments in rice. Rice (NY) 11, 23.
DOI PMID |
| [8] | Cui YN, Li XJ, Yu M, Li RL, Fan LS, Zhu YF, Lin JX (2018a). Sterols regulate endocytic pathways during flg22- induced defense responses in Arabidopsis. Development 145, dev165688. |
| [9] |
Cui YN, Yu M, Yao XM, Xing JJ, Lin JX, Li XJ (2018b). Single-particle tracking for the quantification of membrane protein dynamics in living plant cells. Mol Plant 11, 1315-1327.
DOI URL |
| [10] |
Cui YN, Zhao YX, Lu YQ, Su X, Chen YY, Shen YB, Lin JX, Li XJ (2021). In vivo single-particle tracking of the aquaporin AtPIP2;1 in stomata reveals cell type-specific dynamics. Plant Physiol 185, 1666-1681.
DOI URL |
| [11] |
Donnert G, Keller J, Medda R, Andrei MA, Rizzoli SO, Lührmann R, Jahn R, Eggeling C, Hell SW (2006). Macromolecular-scale resolution in biological fluorescence microscopy. Proc Natl Acad Sci USA 103, 11440-11445.
PMID |
| [12] |
Douglass AD, Vale RD (2005). Single-molecule microscopy reveals plasma membrane microdomains created by protein-protein networks that exclude or trap signaling molecules in T cells. Cell 121, 937-950.
DOI PMID |
| [13] |
Gutierrez R, Grossmann G, Frommer WB, Ehrhardt DW (2010). Opportunities to explore plant membrane organization with super-resolution microscopy. Plant Physiol 154, 463-466.
DOI PMID |
| [14] |
Hao HQ, Fan LS, Chen T, Li RL, Li XJ, He QH, Botella MA, Lin JX (2014). Clathrin and membrane microdomains cooperatively regulate RbohD dynamics and activity in Arabidopsis. Plant Cell 26, 1729-1745.
DOI URL |
| [15] |
Hoffmann N, King S, Samuels AL, McFarlane HE (2021). Subcellular coordination of plant cell wall synthesis. Dev Cell 56, 933-948.
DOI URL |
| [16] |
Jaqaman K, Grinstein S (2012). Regulation from within: the cytoskeleton in transmembrane signaling. Trends Cell Biol 22, 515-526.
DOI PMID |
| [17] | Jozefowicz AM, Matros A, Witzel K, Mock HP (2018). Mini-scale isolation and preparation of plasma membrane proteins from potato roots for LC/MS analysis. In: Mock HP, Matros A, Witzel K, eds. Plant Membrane Proteomics. New York: Humana Press. pp. 195-204. |
| [18] |
Kipper FC, Tamajusuku ASK, Minussi DC, Vargas JE, Battastini AMO, Kaczmarek E, Robson SC, Lenz G, Wink MR (2018). Analysis of NTPDase2 in the cell membrane using fluorescence recovery after photobleaching (FRAP). Cytometry A 93, 232-238.
DOI URL |
| [19] |
Kleine-Vehn J, Wabnik K, Martinière A, Łangowski Ł, Willig K, Naramoto S, Leitner J, Tanaka H, Jakobs S, Robert S, Luschnig C, Govaerts W, Hell SW, Runions J, Friml J (2011). Recycling, clustering, and endocytosis jointly maintain PIN auxin carrier polarity at the plasma membrane. Mol Syst Biol 7, 540.
DOI PMID |
| [20] |
Komis G, Mistrik M, Šamajová O, Doskočilová A, Ovečka M, Illés P, Bartek J, Šamaj J (2014). Dynamics and organization of cortical microtubules as revealed by superresolution structured illumination microscopy. Plant Physiol 165, 129-148.
DOI PMID |
| [21] |
Kure JL, Andersen CB, Mortensen KI, Wiseman PW, Arnspang EC (2020). Revealing plasma membrane nano-domains with diffusion analysis methods. Membranes (Basel) 10, 314.
DOI URL |
| [22] |
Le Gall H, Philippe F, Domon JM, Gillet F, Pelloux J, Rayon C (2015). Cell wall metabolism in response to abiotic stress. Plants (Basel) 4, 112-166.
DOI URL |
| [23] |
Li XJ, Luu DT, Maurel C, Lin JX (2013). Probing plasma membrane dynamics at the single-molecule level. Trends Plant Sci 18, 617-624.
DOI PMID |
| [24] |
Li XJ, Wang XH, Yang Y, Li RL, He QH, Fang XH, Luu DT, Maurel C, Lin JX (2011). Single-molecule analysis of PIP2;1 dynamics and partitioning reveals multiple modes of Arabidopsis plasma membrane aquaporin regulation. Plant Cell 23, 3780-3797.
DOI URL |
| [25] |
Lv XQ, Jing YP, Xiao JW, Zhang YD, Zhu YF, Julian R, Lin JX (2017). Membrane microdomains and the cytoskeleton constrain AtHIR1 dynamics and facilitate the formation of an AtHIR1-associated immune complex. Plant J 90, 3-16.
DOI URL |
| [26] |
Machta BB, Papanikolaou S, Sethna J, Veatch S (2011). Minimal model of plasma membrane heterogeneity requires coupling cortical actin to criticality. Biophys J 100, 1668-1677.
DOI PMID |
| [27] |
Malinovsky FG, Fange lJU, Willats WGT (2014). The role of the cell wall in plant immunity. Front Plant Sci 5, 178.
DOI PMID |
| [28] |
Martínez-Muñoz L, Rodriguez-Frade JM, Barroso R, Sorzano CÓS, Torreno-Pina JA, Sañtiago CA, Manzo C, Lucas P, García-Cuesta EM, Gutierrez E, Barrio L, Vargas J, Cascio G, Carrasco YR, Sánchez-Madrid F, García-Parajo MF, Mellado M (2018). Separating actin-dependent chemokine receptor nanoclustering from dimerization indicates a role for clustering in CXCR4 signaling and function. Mol Cell 70, 106-119.
DOI PMID |
| [29] |
Martinière A, Lavagi I, Nageswaran G, Rolfe DJ, Maneta- Peyret L, Luu DT, Botchway SW, Webb SED, Mongrand S, Maurel C, Martin-Fernandez ML, Kleine-Vehn J, Friml J, Moreau P, Runions J (2012). Cell wall constrains lateral diffusion of plant plasma-membrane proteins. Proc Natl Acad Sci USA 109, 12805-12810.
DOI PMID |
| [30] |
Mattila PK, Batista FD, Treanor B (2016). Dynamics of the actin cytoskeleton mediates receptor cross talk: an emerging concept in tuning receptor signaling. J Cell Biol 212, 267-280.
DOI PMID |
| [31] |
McKenna JF, Rolfe DJ, Webb SED, Tolmie AF, Botchway SW, Martin-Fernandez ML, Hawes C, Runions J (2019). The cell wall regulates dynamics and size of plasma-membrane nanodomains in Arabidopsis. Proc Natl Acad Sci USA 116, 12857-12862.
DOI PMID |
| [32] |
Mudumbi KC, Schirmer EC, Yang WD (2016). Single-point single-molecule FRAP distinguishes inner and outer nuclear membrane protein distribution. Nat Commun 7, 12562.
DOI PMID |
| [33] | Mudumbi KC, Yang WD (2017). Determination of membrane protein distribution on the nuclear envelope by single-point single-molecule FRAP. Curr Protoc Cell Biol 76, 21.11.1-21.11.13. |
| [34] |
Nederveen-Schippers LM, Pathak P, Keizer-Gunnink I, Westpha lAH, Van Haastert PJM, Borst JW, Kortholt A, Skakun V (2021). Combined FCS and PCH analysis to quantify protein dimerization in living cells. Int J Mol Sci 22, 7300.
DOI URL |
| [35] |
Pinaud F, Michalet X, Iyer G, Margeat E, Moore HP, Weiss S (2009). Dynamic partitioning of a glycosyl-phosphatidylinositol-anchored protein in glycosphingolipid-rich microdomains imaged by single-quantum dot tracking. Traffic 10, 691-712.
DOI PMID |
| [36] |
Rostoks N, Schmierer D, Kudrna D, Kleinhofs A (2003). Barley putative hypersensitive induced reaction genes: genetic mapping, sequence analyses and differential expression in disease lesion mimic mutants. Theor Appl Genet 107, 1094-1101.
DOI PMID |
| [37] |
Schneider F, Hernandez-Varas P, Lagerholm BC, Shrestha D, Sezgin E, Roberti MJ, Ossato G, Hecht F, Eggeling C, Urbančič I (2020). High photon count rates improve the quality of super-resolution fluorescence fluctuation spectroscopy. J Phys D Appl Phys 53, 164003.
DOI |
| [38] |
Somerville C, Bauer S, Brininstool G, Facette M, Hamann T, Milne J, Osborne E, Paredez A, Persson S, Raab T, Vorwerk S, Youngs H (2004). Toward a systems approach to understanding plant cell walls. Science 306, 2206-2211.
DOI PMID |
| [39] |
Son S, Oh CJ, An CS (2014). Arabidopsis thaliana remorins interact with SnRK1 and play a role in susceptibility to beet curly top virus and beet severe curly top virus. Plant Pathol J 30, 269-278.
DOI URL |
| [40] |
Su BD, Zhang X, Li L, Abbas S, Yu M, Cui YN, Baluška F, Hwang I, Shan XY, Lin JX (2021). Dynamic spatial reorganization of BSK1 complexes in the plasma membrane underpins signal-specific activation for growth and immunity. Mol Plant 14, 588-603.
DOI PMID |
| [41] | Sundd P, Gutierrez E, Pospieszalska M, Groisman A, Ley K (2010). Stressed and compressed molecular bonds revealed in footprints of rolling neutrophils using total internal reflection fluorescence microscopy. Biophys J 98, 595a. |
| [42] | Tingey M, Li YC, Yang WD (2021). Protocol for single-molecule fluorescence recovery after photobleaching microscopy to analyze the dynamics and spatial locations of nuclear transmembrane proteins in live cells. STAR Pro-toc 2, 100490. |
| [43] |
Villarruel C, Dawson SP (2020). Quantification of fluctua-tions from fluorescence correlation spectroscopy experi-ments in reaction-diffusion systems. Phys Rev E 102, 052407.
DOI PMID |
| [44] |
Wan YL, Ash III WM, Fan LS, Hao HQ, Kim MK, Lin JX (2011). Variable-angle total internal reflection fluorescence microscopy of intact cells of Arabidopsis thaliana. Plant Methods 7, 27.
DOI |
| [45] |
Wang L, Li H, Lv XQ, Chen T, Li RL, Xue YQ, Jiang JJ, Jin B, Baluška F, Šamaj J, Wang XL, Lin JX (2015). Spatiotemporal dynamics of the BRI1 receptor and its regulation by membrane microdomains in living Arabidopsis cells. Mol Plant 8, 1334-1349.
DOI PMID |
| [46] |
Wang QL, Zhao YY, Luo WX, Li RL, He QH, Fang XH, Michele RD, Ast C, Von Wirén N, Lin JX (2013). Single- particle analysis reveals shutoff control of the Arabidopsis ammonium transporter AMT1;3 by clustering and inter-nalization . Proc Natl Acad Sci USA 110, 13204-13209.
DOI URL |
| [47] | Wohland T, Bag N, Ma XX, Huang SR (2013). Epidermal growth factor receptor localization in cell membranes in-vestigated by imaging FCS. Biophys J 104, 681a. |
| [48] | Wollert T, Langford GM (2022). Super-resolution imaging of the actin cytoskeleton in living cells using TIRF-SIM. In: Gavin RH, ed. Cytoskeleton. New York: Humana. pp. 3-24. |
| [49] |
Xing JJ, Li XJ, Wang XH, Lv XQ, Wang L, Zhang L, Zhu YF, Shen QH, Baluška F, Šamaj J, Lin JX (2019). Se-cretion of phospholipase D deltafunctions as a regulatory mechanism in plant innate immunity. Plant Cell 31, 3015-3032.
DOI URL |
| [50] |
Xu CW, Abbas S, Qian HP, Yu M, Zhang X, Li XJ, Cui YN, Lin JX (2022). Environmental cues contribute to dynamic plasma membrane organization of nanodomains containing flotillin-1 and hypersensitive induced reaction-1 pro-teins in Arabidopsis thaliana. Front Plant Sci 13, 897594.
DOI URL |
| [51] |
Xue YQ, Xing JJ, Wan YL, Lv XQ, Fan LS, Zhang YD, Song K, Wang L, Wang XH, Deng X, Baluška F, Christie JM, Lin JX (2018). Arabidopsis blue light receptor phototropin 1 undergoes blue light-induced activation in membrane microdomains. Mol Plant 11, 846-859.
DOI URL |
| [52] |
Yasui M, Hiroshima M, Kozuka J, Sako Y, Ueda M (2018). Automated single-molecule imaging in living cells. Nat Commun 9, 3061.
DOI PMID |
| [53] | Yu M, Li RL, Cui YN, Chen WJ, Li B, Zhang X, Bu YF, Cao YY, Xing JJ, Jewaria PK, Li XJ, Bhalerao RP, Yu F, Lin JX (2020). The RALF1-FERONIA interaction modulates endocytosis to mediate control of root growth in Arabidop-sis. Development 147, dev189902. |
| [54] |
Yu M, Liu HJ, Dong ZY, Xiao JW, Su BD, Fan LS, Komis G, Šamaj J, Lin JX, Li RL (2017). The dynamics and endocytosis of Flot1 protein in response to flg22 in Arabidopsis. J Plant Physiol 215, 73-84.
DOI URL |
| [55] |
Zhang X, Cui YN, Yu M, Su BD, Gong W, Baluška F, Ko-mis G, Šamaj J, Shan XY, Lin JX (2019). Phosphoryla-tion-mediated dynamics of nitrate transceptor NRT1.1 regulate auxin flux and nitrate signaling in lateral root growth. Plant Physiol 181, 480-498.
DOI PMID |
| [1] | TANG Yuan-Xiang, XIONG Shi-Chen, ZHU Hong-Feng, ZHANG Xin-Sheng, YOU Cheng-Ming, LIU Si-Ning, TAN Bo, XU Zhen-Feng. Effects of long-term nitrogen addition on leaf litter production and carbon, nitrogen and phosphorus return of the dominant tree species in broadleaf evergreen forests on the western margin of Sichuan Basin [J]. Chin J Plant Ecol, 2025, 49(5): 720-731. |
| [2] | Li Yanpeng, Chen Jie, Lu Chunyang, Xu Han. Community characteristics of a 64-ha secondary forest dynamics plot in a tropical montane rainforest in Jianfengling, Hainan [J]. Biodiv Sci, 2025, 33(2): 24445-. |
| [3] | Xiuxiu Chen, Ling Tang, Wenjia Hu, Zhaolin Yang, Xin Deng, Xiaohua Wang. Quantitative Analysis of Plasma Membrane Order in Live Plant Cells [J]. Chinese Bulletin of Botany, 2025, 60(1): 90-100. |
| [4] | Rongjiao Li, Jianghai Dong, Wenfang Zheng, Ruyuan Liu, Lijuan Zhao, Ruihe Gao. Soil faunal diversity characteristics and influencing factors across the altitude gradient in the poplar-birch forest of Guandi Mountains [J]. Biodiv Sci, 2024, 32(9): 24070-. |
| [5] | LIU Shi-Ling, YANG Bao-Guo, ZHENG Lu, SHU Wei-Wei, MIN Hui-Lin, ZHANG Pei, LI Hua, YANG Kun, ZHOU Bing-Jiang, TIAN Zu-Wei. Seasonal stem radial growth of Castanopsis hystrix plantation and its response to climatic factors in Guangxi, China [J]. Chin J Plant Ecol, 2024, 48(8): 1021-1034. |
| [6] | Jianwei Cheng, Manhou Xu, Yongjing Dou, Yadong Wang, Yanan Wang, Xinmin Liu, Frank Yonghong Li. Seasonal dynamics of arthropod communities during horse dung decomposition in Inner Mongolian grasslands [J]. Biodiv Sci, 2024, 32(6): 24018-. |
| [7] | Jiaxin Wei, Zhiguo Jiang, Linsen Yang, Huanhuan Xiong, Jiaojiao Jin, Fanglin Luo, Jiehua Li, Hao Wu, Yaozhan Xu, Xiujuan Qiao, Xinzeng Wei, Hui Yao, Huiliang Yu, Jingyuan Yang, Mingxi Jiang. Community composition and structure in a 25 ha mid-subtropical mountain deciduous broad-leaved forest dynamics plot in Shennongjia, Hubei, China [J]. Biodiv Sci, 2024, 32(3): 23338-. |
| [8] | Xiaolin Liu, Yougui Wu, Minhua Zhang, Xiaorong Chen, Zhicheng Zhu, Dingyun Chen, Shu Dong, Buhang Li, Bingyang Ding, Yu Liu. Community composition and structure of a 25-ha forest dynamics plot of subtropical forest in Baishanzu, Zhejiang Province [J]. Biodiv Sci, 2024, 32(2): 23294-. |
| [9] | Kailun Xu, Xiaorong Chen, Minhua Zhang, Wanwan Yu, Sumei Wu, Zhicheng Zhu, Dingyun Chen, Rongguang Lan, Shu Dong, Yu Liu. Succession and topography jointly influence the diversity of plant sexual systems in the Baishanzu forest community [J]. Biodiv Sci, 2024, 32(12): 24338-. |
| [10] | XU Ming-Ze, ZHAO Hong-Xian, LI Cheng, LI Man-Le, TIAN Yun, LIU Peng, ZHA Tian-Shan. Characteristics of seasonal leaf trait network and its drivers in Artemisia ordosica in the Mau Us Sandy Land [J]. Chin J Plant Ecol, 2024, 48(12): 1650-1665. |
| [11] | LAN Guang-Fei, ZHANG Qiang, CHEN Xiang-Biao, CHEN Shi-Dong, XIONG De-Cheng, LIU Xiao-Fei, YANG Zhi-Jie, YANG Yu-Sheng. Seasonal dynamics of litterfall of a Castanopsis kawakamii evergreen broadleaf forest in mid-subtropical China and their influencing factors [J]. Chin J Plant Ecol, 2024, 48(12): 1589-1601. |
| [12] | Weiwei Wang, Xiangcheng Mi, Ningning Wang, Haibao Ren, Zhixi Tang, Zhuning Zhang, Keping Ma, Lei Chen. Plant diversity dataset of the 24-ha Zhejiang Gutianshan subtropical evergreen broad-leaved forest dynamics plot (2005-2020) [J]. Biodiv Sci, 2024, 32(12): 24417-. |
| [13] | Yasu Cao, Min Fan, Yu Peng, Jiaxun Xin, Nanyi Peng. Effects of landscape pattern dynamics on plant species and functional diversity in Hunshandak Sandland [J]. Biodiv Sci, 2023, 31(8): 23048-. |
| [14] | Zhehan Chen, Jin Yin, Ji Ye, Dongwei Liu, Zikun Mao, Shuai Fang, Fei Lin, Xugao Wang. Effects of simulated warming on seasonal dynamics of herbaceous diversity in temperate secondary forests in Northeast China [J]. Biodiv Sci, 2023, 31(5): 23059-. |
| [15] | HE Chun-Mei, LI Yu-Shan, YIN Qiu-Long, JIA Shi-Hong, HAO Zhan-Qing. Diameter structure and quantitative characteristics of dominant tree species in warm temperate deciduous broadleaf forest plot in Huangguan of Qinling Mountains, China [J]. Chin J Plant Ecol, 2023, 47(12): 1658-1667. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||