

Chinese Bulletin of Botany ›› 2019, Vol. 54 ›› Issue (6): 723-732.DOI: 10.11983/CBB19037 cstr: 32102.14.CBB19037
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Jieli He1,Tiantian Shi2,Ling Chen3,Haigang Wang3,Zhijun Gao4,Meihong Yang1,Ruiyun Wang2,3,*( ),Zhijun Qiao3,*(
),Zhijun Qiao3,*( )
)
Received:2019-02-24
Accepted:2019-06-18
Online:2019-11-01
Published:2020-07-09
Contact:
Ruiyun Wang,Zhijun Qiao
Jieli He,Tiantian Shi,Ling Chen,Haigang Wang,Zhijun Gao,Meihong Yang,Ruiyun Wang,Zhijun Qiao. The Genetic Diversity of Common Millet (Panicum miliaceum) Germplasm Resources Based on the EST-SSR Markers[J]. Chinese Bulletin of Botany, 2019, 54(6): 723-732.
| Ecotope/abroad | Origin | Number of accession | Total |
|---|---|---|---|
| Northwest spring and summer-sowing ecotope (NWSS) | Xinjiang | 4 | 4 |
| Northern spring-sowing ecotope (NSP) | Qinghai | 13 | 48 |
| Gansu | 11 | ||
| Inner Mongolia | 14 | ||
| Shanxi | 10 | ||
| Loess Plateau spring and summer-sowing ecotope (LPSS) | Shanxi | 18 | 37 |
| Shaanxi | 8 | ||
| Ningxia | 11 | ||
| Northeast spring-sowing ecotope (NES) | Heilongjiang | 5 | 9 |
| Jilin | 3 | ||
| Liaoning | 1 | ||
| Northern summer-sowing ecotope (NSU) | Hebei | 9 | 13 |
| Shandong | 2 | ||
| Anhui | 1 | ||
| Henan | 1 | ||
| Southern autumn and winter-sowing ecotope (SAW) | Hainan | 2 | 2 |
| Abroad | Former Soviet Union | 2 | 31 |
| Poland | 2 | ||
| India | 27 | ||
| Total | 144 | ||
Table 1 Distribution of common millet accessions in different ecotopes of China and abroad
| Ecotope/abroad | Origin | Number of accession | Total |
|---|---|---|---|
| Northwest spring and summer-sowing ecotope (NWSS) | Xinjiang | 4 | 4 |
| Northern spring-sowing ecotope (NSP) | Qinghai | 13 | 48 |
| Gansu | 11 | ||
| Inner Mongolia | 14 | ||
| Shanxi | 10 | ||
| Loess Plateau spring and summer-sowing ecotope (LPSS) | Shanxi | 18 | 37 |
| Shaanxi | 8 | ||
| Ningxia | 11 | ||
| Northeast spring-sowing ecotope (NES) | Heilongjiang | 5 | 9 |
| Jilin | 3 | ||
| Liaoning | 1 | ||
| Northern summer-sowing ecotope (NSU) | Hebei | 9 | 13 |
| Shandong | 2 | ||
| Anhui | 1 | ||
| Henan | 1 | ||
| Southern autumn and winter-sowing ecotope (SAW) | Hainan | 2 | 2 |
| Abroad | Former Soviet Union | 2 | 31 |
| Poland | 2 | ||
| India | 27 | ||
| Total | 144 | ||
| Number | Unicode | Accession name | Origin |
|---|---|---|---|
| 1 | 00000177 | Hongmizi | Ningan, Heilongjiang |
| 2 | 00000750 | Baimizi | Shawan, Xinjiang |
| 3 | 00006653 | Jinshu | Hainan |
| 4 | 00007238 | Dahongmizi | Bameng, Inner Mongolia |
| 5 | 00007478 | Baigedami | Huangzhong, Qinghai |
| 6 | No unicode | Hongshuzi | Anyang, Henan |
Table 2 Screening of SSR primers for common millet
| Number | Unicode | Accession name | Origin |
|---|---|---|---|
| 1 | 00000177 | Hongmizi | Ningan, Heilongjiang |
| 2 | 00000750 | Baimizi | Shawan, Xinjiang |
| 3 | 00006653 | Jinshu | Hainan |
| 4 | 00007238 | Dahongmizi | Bameng, Inner Mongolia |
| 5 | 00007478 | Baigedami | Huangzhong, Qinghai |
| 6 | No unicode | Hongshuzi | Anyang, Henan |
| Ecotope/ abroad | Accessions | Na | Ne | I | Ho | He | PIC |
|---|---|---|---|---|---|---|---|
| NWSS | 4 | 2.3375±0.5017 | 2.1517±0.4194 | 0.7644±0.2178 | 0.8042±0.2688 | 0.5944±0.1193 | 0.3551 |
| NSP | 48 | 2.5750±0.4975 | 2.3106±0.3211 | 0.8604±0.1576 | 0.8228±0.1308 | 0.5655±0.0604 | 0.4536 |
| LPSS | 37 | 2.5750±0.4975 | 2.2803±0.3110 | 0.8506±0.1534 | 0.8384±0.1166 | 0.5615±0.0595 | 0.4203 |
| NES | 9 | 2.5125±0.5030 | 2.2435±0.3929 | 0.8289±0.1823 | 0.7937±0.1732 | 0.5737±0.0909 | 0.4212 |
| NSU | 13 | 2.5625±0.4992 | 2.2815±0.3527 | 0.8496±0.1632 | 0.7946±0.1608 | 0.5712±0.0667 | 0.4304 |
| SAW | 2 | 2.2375±0.5092 | 2.0608±0.4387 | 0.7347±0.2316 | 0.7812±0.3265 | 0.6813±0.2006 | 0.2156 |
| Domestic | 113 | 2.5750 ±0.4975 | 2.3122±0.3086 | 0.8628±0.1554 | 0.8200±0.1188 | 0.5625±0.0584 | 0.4651 |
| Abroad | 31 | 2.5750±0.4975 | 2.2464±0.2909 | 0.8387±0.1449 | 0.8540±0.1193 | 0.5571±0.0561 | 0.3896 |
Table 3 Parameters of genetic diversity in different ecotope of common millet
| Ecotope/ abroad | Accessions | Na | Ne | I | Ho | He | PIC |
|---|---|---|---|---|---|---|---|
| NWSS | 4 | 2.3375±0.5017 | 2.1517±0.4194 | 0.7644±0.2178 | 0.8042±0.2688 | 0.5944±0.1193 | 0.3551 |
| NSP | 48 | 2.5750±0.4975 | 2.3106±0.3211 | 0.8604±0.1576 | 0.8228±0.1308 | 0.5655±0.0604 | 0.4536 |
| LPSS | 37 | 2.5750±0.4975 | 2.2803±0.3110 | 0.8506±0.1534 | 0.8384±0.1166 | 0.5615±0.0595 | 0.4203 |
| NES | 9 | 2.5125±0.5030 | 2.2435±0.3929 | 0.8289±0.1823 | 0.7937±0.1732 | 0.5737±0.0909 | 0.4212 |
| NSU | 13 | 2.5625±0.4992 | 2.2815±0.3527 | 0.8496±0.1632 | 0.7946±0.1608 | 0.5712±0.0667 | 0.4304 |
| SAW | 2 | 2.2375±0.5092 | 2.0608±0.4387 | 0.7347±0.2316 | 0.7812±0.3265 | 0.6813±0.2006 | 0.2156 |
| Domestic | 113 | 2.5750 ±0.4975 | 2.3122±0.3086 | 0.8628±0.1554 | 0.8200±0.1188 | 0.5625±0.0584 | 0.4651 |
| Abroad | 31 | 2.5750±0.4975 | 2.2464±0.2909 | 0.8387±0.1449 | 0.8540±0.1193 | 0.5571±0.0561 | 0.3896 |
| Population | NWSS | NSP | LPSS | NES | NSU | SAW | Abroad |
|---|---|---|---|---|---|---|---|
| NWSS | 0.9560 | 0.9614 | 0.9487 | 0.9380 | 0.8694 | 0.9477 | |
| NSP | 0.0449 | 0.9884 | 0.9678 | 0.9794 | 0.9110 | 0.9864 | |
| LPSS | 0.0394 | 0.0117 | 0.9716 | 0.9830 | 0.9116 | 0.9865 | |
| NES | 0.0527 | 0.0327 | 0.0288 | 0.9675 | 0.8974 | 0.9587 | |
| NSU | 0.0640 | 0.0208 | 0.0171 | 0.0331 | 0.9023 | 0.9762 | |
| SAW | 0.1400 | 0.0932 | 0.0926 | 0.1083 | 0.1029 | 0.9102 | |
| Abroad | 0.0537 | 0.0137 | 0.0136 | 0.0422 | 0.0240 | 0.0941 |
Table 4 Parameters of Nei’s genetic distance and Nei’s genetic agreement in common millet populations
| Population | NWSS | NSP | LPSS | NES | NSU | SAW | Abroad |
|---|---|---|---|---|---|---|---|
| NWSS | 0.9560 | 0.9614 | 0.9487 | 0.9380 | 0.8694 | 0.9477 | |
| NSP | 0.0449 | 0.9884 | 0.9678 | 0.9794 | 0.9110 | 0.9864 | |
| LPSS | 0.0394 | 0.0117 | 0.9716 | 0.9830 | 0.9116 | 0.9865 | |
| NES | 0.0527 | 0.0327 | 0.0288 | 0.9675 | 0.8974 | 0.9587 | |
| NSU | 0.0640 | 0.0208 | 0.0171 | 0.0331 | 0.9023 | 0.9762 | |
| SAW | 0.1400 | 0.0932 | 0.0926 | 0.1083 | 0.1029 | 0.9102 | |
| Abroad | 0.0537 | 0.0137 | 0.0136 | 0.0422 | 0.0240 | 0.0941 |
| Group | Accessions | Na | Ne | I | Ho | He | PIC |
|---|---|---|---|---|---|---|---|
| A | 33 | 2.5750±0.4975 | 2.3226±0.3374 | 0.8654±0.1620 | 0.8055±0.1417 | 0.5696±0.0636 | 0.4716 |
| B | 15 | 2.5125±0.5030 | 2.2471±0.3383 | 0.8330±0.1627 | 0.8153±0.1530 | 0.5651±0.0686 | 0.3993 |
| C | 96 | 2.5750±0.4975 | 2.2922±0.2894 | 0.8571±0.1495 | 0.8363±0.1111 | 0.5599±0.0553 | 0.4380 |
| C1 | 70 | 2.5750±0.4975 | 2.3136±0.3098 | 0.8635±0.1551 | 0.8291±0.1199 | 0.5643±0.0584 | 0.4531 |
| C2 | 26 | 2.5625±0.4992 | 2.1912±0.2797 | 0.8163±0.1391 | 0.8555 ±0.1212 | 0.5477±0.0558 | 0.3420 |
| C11 | 37 | 2.5750±0.4975 | 2.3014±0.3143 | 0.8589±0.1561 | 0.8332±0.1182 | 0.5664±0.0603 | 0.4382 |
| C12 | 33 | 2.5750±0.4975 | 2.3028±0.3224 | 0.8583±0.1571 | 0.8255±0.1383 | 0.5654±0.0610 | 0.4348 |
Table 5 Genetic diversity of common millet groups based on UPGMA cluster analysis
| Group | Accessions | Na | Ne | I | Ho | He | PIC |
|---|---|---|---|---|---|---|---|
| A | 33 | 2.5750±0.4975 | 2.3226±0.3374 | 0.8654±0.1620 | 0.8055±0.1417 | 0.5696±0.0636 | 0.4716 |
| B | 15 | 2.5125±0.5030 | 2.2471±0.3383 | 0.8330±0.1627 | 0.8153±0.1530 | 0.5651±0.0686 | 0.3993 |
| C | 96 | 2.5750±0.4975 | 2.2922±0.2894 | 0.8571±0.1495 | 0.8363±0.1111 | 0.5599±0.0553 | 0.4380 |
| C1 | 70 | 2.5750±0.4975 | 2.3136±0.3098 | 0.8635±0.1551 | 0.8291±0.1199 | 0.5643±0.0584 | 0.4531 |
| C2 | 26 | 2.5625±0.4992 | 2.1912±0.2797 | 0.8163±0.1391 | 0.8555 ±0.1212 | 0.5477±0.0558 | 0.3420 |
| C11 | 37 | 2.5750±0.4975 | 2.3014±0.3143 | 0.8589±0.1561 | 0.8332±0.1182 | 0.5664±0.0603 | 0.4382 |
| C12 | 33 | 2.5750±0.4975 | 2.3028±0.3224 | 0.8583±0.1571 | 0.8255±0.1383 | 0.5654±0.0610 | 0.4348 |
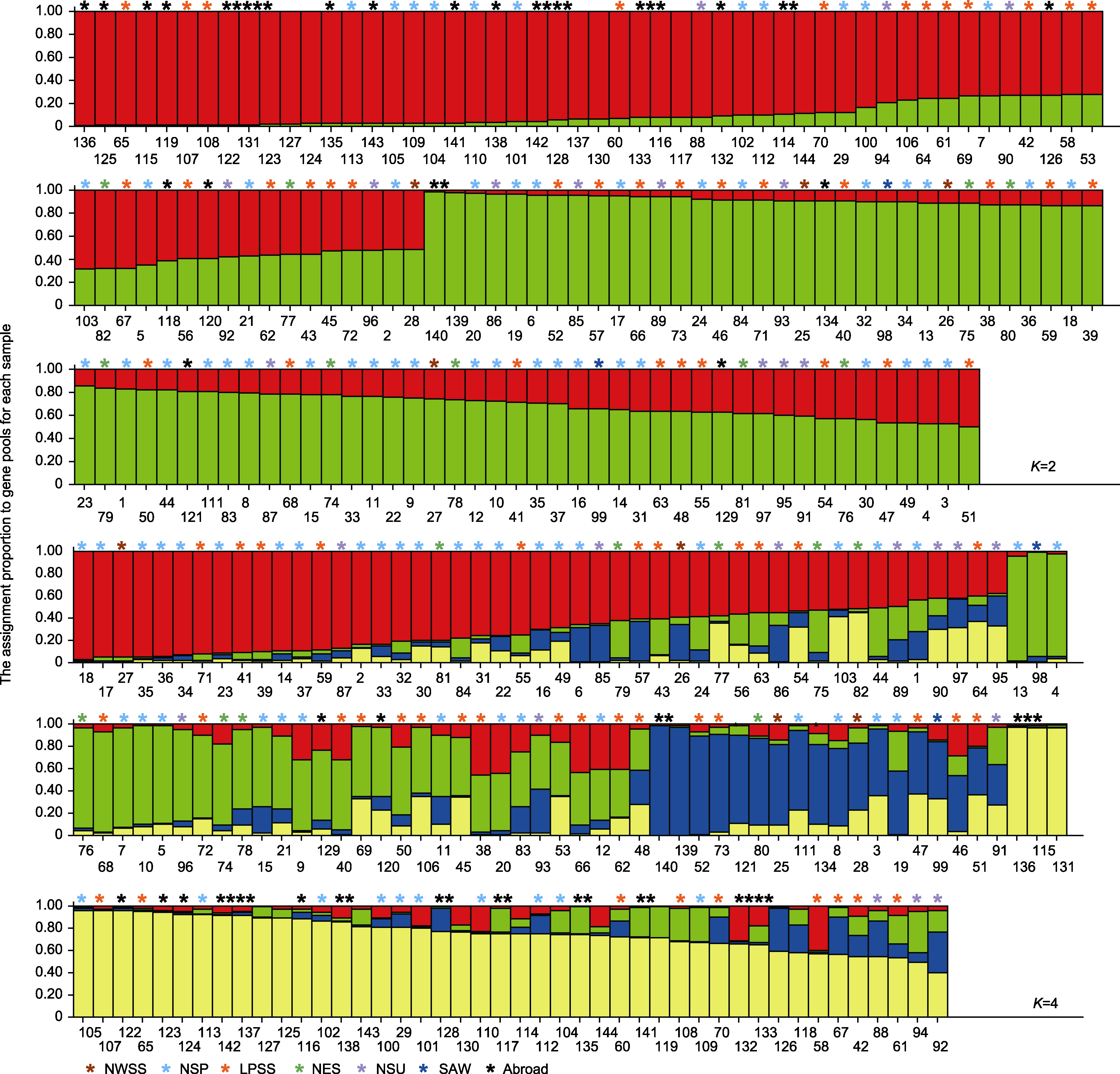
Figure 4 Genetic structure of common millet based on Structure Color represents group; bar and the horizontal coordinate represent origin and its serial number, respectively.
| Cluster | Accessions | Na | Ne | I | Ho | He | PIC | |
|---|---|---|---|---|---|---|---|---|
| K=2 | Red | 68 | 2.5750±0.4975 | 2.2446±0.2626 | 0.8407±0.1403 | 0.8602±0.1056 | 0.5528±0.0520 | 0.3773 |
| Green | 76 | 2.5750±0.4975 | 2.3396±0.3329 | 0.8711±0.1620 | 0.7975±0.1315 | 0.5679±0.0619 | 0.5031 | |
| K=4 | Red | 47 | 2.5750± 0.4975 | 2.3496±0.3944 | 0.8745±0.1768 | 0.7439±0.1668 | 0.5785±0.0743 | 0.5296 |
| Green | 31 | 2.5750± 0.4975 | 2.2844±0.3202 | 0.8516± 0.1549 | 0.8225±0.1399 | 0.5635±0.0607 | 0.4414 | |
| Blue | 19 | 2.5750±0.4975 | 2.2973±0.3199 | 0.8575±0.1568 | 0.8190±0.1311 | 0.5628±0.0608 | 0.4450 | |
| Yellow | 47 | 2.5750±0.4975 | 2.2238±0.2723 | 0.8312±0.1387 | 0.3634±0.2702 | 0.4984±0.1050 | 0.3543 | |
Table 6 Genetic diversity analysis of different cluster based on genetic structure (K=2 and K=4)
| Cluster | Accessions | Na | Ne | I | Ho | He | PIC | |
|---|---|---|---|---|---|---|---|---|
| K=2 | Red | 68 | 2.5750±0.4975 | 2.2446±0.2626 | 0.8407±0.1403 | 0.8602±0.1056 | 0.5528±0.0520 | 0.3773 |
| Green | 76 | 2.5750±0.4975 | 2.3396±0.3329 | 0.8711±0.1620 | 0.7975±0.1315 | 0.5679±0.0619 | 0.5031 | |
| K=4 | Red | 47 | 2.5750± 0.4975 | 2.3496±0.3944 | 0.8745±0.1768 | 0.7439±0.1668 | 0.5785±0.0743 | 0.5296 |
| Green | 31 | 2.5750± 0.4975 | 2.2844±0.3202 | 0.8516± 0.1549 | 0.8225±0.1399 | 0.5635±0.0607 | 0.4414 | |
| Blue | 19 | 2.5750±0.4975 | 2.2973±0.3199 | 0.8575±0.1568 | 0.8190±0.1311 | 0.5628±0.0608 | 0.4450 | |
| Yellow | 47 | 2.5750±0.4975 | 2.2238±0.2723 | 0.8312±0.1387 | 0.3634±0.2702 | 0.4984±0.1050 | 0.3543 | |
| [1] | 董俊丽, 王海岗, 陈凌, 王君杰, 曹晓宁, 王纶, 乔治军 ( 2015). 糜子骨干种质遗传多样性和遗传结构分析. 中国农业科学 48, 3121-3131. |
| [2] | 国家谷子糜子产业技术体系 ( 2018). 中国现代农业产业可持续发展战略研究·谷子糜子分册. 北京: 中国农业出版社. pp. 3-22. |
| [3] | 郭琪, 郭大龙, 郭丽丽, 张琳, 侯小改 ( 2015). SSR分子标记在牡丹亲缘关系研究中的应用与研究进展. 植物学报 50, 652-664. |
| [4] | 连帅, 陆平, 乔治军, 张琦, 张茜, 刘敏轩, 王瑞云 ( 2016). 利用SSR分子标记研究国内外黍稷地方品种和野生资源的遗传多样性. 中国农业科学 49, 3264-3275. |
| [5] | 刘笑瑜 ( 2017). 利用高基元SSR分析中国糜子资源的遗传多样性. 硕士论文. 太谷: 山西农业大学. pp. 22-41. |
| [6] | 刘笑瑜, 王瑞云, 刘敏轩, 邱岩岩, 季煦, 连帅, 乔治军, 王纶, 王海岗 ( 2016). 利用SSR标记分析40份糜子资源的遗传多样性. 分子植物育种 14, 1624-1630. |
| [7] | 王璐琳, 王瑞云, 何杰丽, 薛延桃, 陈凌, 王海岗, 乔治军 ( 2018). 糜子特异性SSR标记的开发. 山西农业科学 46, 1-4, 86. |
| [8] | 王瑞云 (2017). 糜子遗传多样性及进化研究进展. 北京: 中国农业出版社. pp. 20-92. |
| [9] | 王瑞云, 季煦, 陆平, 刘敏轩, 许月, 王纶, 王海岗, 乔治军 ( 2017a). 利用荧光SSR分析中国糜子遗传多样性. 作物学报 43, 530-548. |
| [10] | 王瑞云, 刘笑瑜, 王海岗, 陆平, 刘敏轩, 陈凌, 乔治军 ( 2017b). 用高基元微卫星标记分析中国糜子遗传多样性. 中国农业科学 50, 3848-3859. |
| [11] | 王舒婷, 何杰丽, 石甜甜, 陈凌, 王海岗, 王瑞云, 乔治军 ( 2019). 利用微卫星标记分析山西糜子的遗传多样性. 植物遗传资源学报 20, 69-78. |
| [12] | 王银月, 刘敏轩, 陆平, 乔治军, 杨天育, 李海, 崔喜艳 ( 2014). 构建黍稷分子遗传图谱SSR引物的筛选. 作物杂志 ( 4), 32-38. |
| [13] | 薛延桃, 陆平, 乔治军, 刘敏轩, 王瑞云 ( 2018). 基于SSR标记的黍稷种质资源遗传多样性及亲缘关系研究. 中国农业科学 51, 2846-2859. |
| [14] | 朱宇佳, 焦凯丽, 罗秀俊, 冯尚国, 王慧中 ( 2018). 基于SSR分子标记的酸浆属植物亲缘关系研究. 植物学报 53, 305-312. |
| [15] | Azevedo ALS, Costa PP, Machado JC, Machado MA, Pereira AV, da Silva Lédo FJ ( 2012). Cross species amplification of Pennisetum glaucum microsatellite markers in Pennisetum purpureum and genetic diversity of napier grass accessions. Crop Sci 52, 1776-1785. |
| [16] | Bonman JM, Babiker EM, Cuesta-Marcos A, Esvelt-Klos K, Brown-Guedira G, Chao SM, See D, Chen JL, Akhunov E, Zhang JL, Bockelman HE, Gordon TC ( 2015). Genetic diversity among wheat accessions from the USDA national small grains collection. Crop Sci 55, 1243-1253. |
| [17] | Changmei S, Dorothy J ( 2014). Millet—the frugal grain. Int J Sci Res Rev 3(4), 75-90. |
| [18] | Cho Yl, Chung JW, Lee GA, Ma KH, Dixit A, Gwag JG, Park YJ ( 2010). Development and characterization of twenty-five new polymorphic microsatellite markers in proso millet ( Panicum miliaceum L.). Genes Genomics 32, 267-273. |
| [19] | Courtois B, Frouin J, Greco R, Bruschi G, Droc G, Hamelin C, Ruiz M, Clément G, Evrard JC, Van Coppenole S, Katsantonis D, Oliveira M, Negrão S, Matos C, Cavigiolo S, Lupotto E, Piffanelli P, Ahmadi N ( 2012). Genetic diversity and population structure in a European collection of rice. Crop Sci 52, 1663-1675. |
| [20] | Evanno G, Regnaut S, Goudet J ( 2005). Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14, 2611-2620. |
| [21] | Falush D, Stephens M, Pritchard JK ( 2003). Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164, 1567-1587. |
| [22] | Habiyaremye C, Matanguihan JB, Guedes JD, Ganjyal GM, Whiteman MR, Kidwell KK, Murphy KM ( 2017). Proso millet ( Panicum miliaceum L.) and its potential for cultivation in the pacific northwest, U.S: a review. Front Plant Sci 7, 1961. |
| [23] | Hu XY, Wang JF, Lu P, Zhang HS ( 2009). Assessment of genetic diversity in broomcorn millet ( Panicum miliaceum L.) using SSR markers. J Genet Genomics 36, 491-500. |
| [24] | Hunt HV, Campana MG, Lawes MC, Park YJ, Bower MA, Howe CJ, Jones MK ( 2011). Genetic diversity and phylogeography of broomcorn millet ( Panicum miliaceum L.) across Eurasia. Mol Ecol 20, 4756-4771. |
| [25] | Liu KJ, Muse SV ( 2005). PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21, 2128-2129. |
| [26] | Liu MX, Xu Y, He JH, Zhang S, Wang YY, Lu P ( 2016). Genetic diversity and population structure of broomcorn millet ( Panicum miliaceum L.) cultivars and landraces in China based on microsatellite markers. Int J Mol Sci 17, 370. |
| [27] | Lu HY, Zhang JP, Liu KB, Wu NQ, Li YM, Zhou KS, Ye ML, Zhang TY, Zhang HJ, Yang XY, Shen LC, Xu DK, Li Q ( 2009). Earliest domestication of common millet ( Panicum miliaceum ) in East Asia extended to 10, 000 years ago. Proc Natl Acad Sci USA 106, 7367-7372. |
| [28] | Murray MG, Thompson WF ( 1980). Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res 8, 4321-4325. |
| [29] | Prevost A, Wilkinson MJ ( 1999). A new system of comparing PCR primers applied to ISSR fingerprinting of potato cultivars. Theor Appl Genet 98, 107-112. |
| [30] | Rajput SG, Plyler-Harveson T, Santra DK ( 2014). Development and characterization of SSR markers in proso millet based on switchgrass genomics. Am J Plant Sci 5, 175-186. |
| [31] | Rajput SG, Santra DK ( 2016). Evaluation of genetic diversity of proso millet germplasm available in the United States using simple-sequence repeat markers. Crop Sci 56, 2401-2409. |
| [32] | Rohlf FJ (2002). NTSYS-pc: Numerical Taxonomy and Multivariate Analysis System, Version 2.10. New York: Exter Publishing Ltd. Setauket. |
| [33] | Saha D, Channabyre Gowda MV, Arya L, Verma M, Bansal KC ( 2016). Genetic and genomic resources of small millets. Crit Rev Plant Sci 35, 56-79. |
| [34] | Satya P, Karan M, Jana S, Mitra S, Sharma A, Karmakar PG, Ray DP ( 2015). Start codon targeted (SCoT) polymorphism reveals genetic diversity in wild and domesticated populations of ramie ( Boehmeria nivea L. Gaudich.), a premium textile fiber producing species. Meta Gene 3, 62-70. |
| [35] | Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S ( 2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 665, 2731-2739. |
| [36] | Tiwar G, Singh R, Singh N, Choudhury DR, Paliwal R, Kumar A, Gupta V ( 2016). Study of arbitrarily amplified (RAPD and ISSR) and gene targeted (SCoT and CBDP) markers for genetic diversity and population structure in kalmegh [Andrographis paniculata(Burm. f.) Nees]. Ind Crops Prod 86, 1-11. |
| [37] | Van Inghelandt D, Melchinger AE, Lebreton C, Stich B ( 2010). Population structure and genetic diversity in a commercial maize breeding program assessed with SSR and SNP markers. Theor Appl Genet 120, 1289-1299. |
| [38] | Wang RY, Hunt HV, Qiao ZJ, Wang L, Han YH ( 2016). Diversity and cultivation of broomcorn millet ( Panicum miliaceum L.) in China: a review. Econ Bot 70, 332-342. |
| [39] | Wang RY, Wang HG, Liu XY, Ji X, Chen L, Lu P, Liu MX, Teng B, Qiao ZJ ( 2018). Waxy allelic diversity in common millet(Panicum miliaceum L.) in China. Crop J 6, 377-385. |
| [40] | Yeh FC, Boyle TJB ( 1997). Population genetic analysis of codominant and dominant markers and quantitative traits. Belg J Bot 129, 157-163. |
| [1] | 黄 承玲, Han Li Rong, Ling Qing Hong, Xiong Yang Sheng, Ling Tian Xiao, Xia Guowei Xia Guowei, Ren Chen Zheng, Wei Zhou. Study on genetic conservation of Rhododendron liboense based on SNP molecular markers,a plant species with extremely small populations [J]. , 2025, 49(濒危植物的保护与恢复): 0-. |
| [2] | Zhang Ruli, Li Dezhu, Zhang Yuxiao. Population Genetic Structure and Climate Adaptation Analysis of Brachystachyum densiflorum [J]. Chinese Bulletin of Botany, 2025, 60(3): 407-424. |
| [3] | Zhigang Yang, Pengcheng Zhang, Haiwen Chang, Liru Kang, Yi Zuo, Haoxin Xiang, Fengying Han. Genetic Diversity Analysis of Pepper Germplasms Based on Morphological Traits and SSR Markers [J]. Chinese Bulletin of Botany, 2025, 60(2): 218-234. |
| [4] | Jiachen Wang, Tangjun Xu, Wei Xu, Gaoji Zhang, Yijin You, Honghua Ruan, Hongyi Liu. Impact of urban landscape pattern on the genetic structure of Thereuopoda clunifera population in Nanjing, China [J]. Biodiv Sci, 2025, 33(1): 24251-. |
| [5] | Kexin Cao, Jingwen Wang, Guo Zheng, Pengfeng Wu, Yingbin Li, Shuyan Cui. Effects of precipitation regime change and nitrogen deposition on soil nematode diversity in the grassland of northern China [J]. Biodiv Sci, 2024, 32(3): 23491-. |
| [6] | Zhengyong Duan, Min Ding, Yuzhuo Wang, Yibing Ding, Ling Chen, Ruiyun Wang, Zhijun Qiao. Genome-wide Identification and Expression Analysis of SBP Genes in Panicum miliaceum [J]. Chinese Bulletin of Botany, 2024, 59(2): 231-244. |
| [7] | Linjun He, Wenjing Yang, Yuhao Shi, Kezhemo Ashuo, Yu Fan, Guoyan Wang, Jingji Li, Songlin Shi, Guihua Yi, Peihao Peng. Effects of plant community phylogeny and functional diversity on Ageratina adenophora invasion under fire disturbance [J]. Biodiv Sci, 2024, 32(11): 24269-. |
| [8] | Shiyi Long, Bobo Zhang, Yuchen Xia, Yangfan Fei, Yani Meng, Bingwei Lü, Yueqing Song, Pu Zheng, Taoran Guo, Jian Zhang, Shaopeng Li. Effects of diversity and temporal stability of native communities on the biomass of invasive species Solidago canadensis [J]. Biodiv Sci, 2024, 32(11): 24263-. |
| [9] | Qingduo Li, Dongmei Li. Analysis for the prevalence of global bat-borne Bartonella [J]. Biodiv Sci, 2023, 31(9): 23166-. |
| [10] | Chen Feng, Jie Zhang, Hongwen Huang. Parallel situ conservation: A new plant conservation strategy to integrate in situ and ex situ conservation of plants [J]. Biodiv Sci, 2023, 31(9): 23184-. |
| [11] | Hailing Qi, Pengzhen Fan, Yuehua Wang, Jie Liu. Genetic diversity and population structure of Juglans regia from six provinces in northern China [J]. Biodiv Sci, 2023, 31(8): 23120-. |
| [12] | Fei Xiong, Hongyan Liu, Dongdong Zhai, Xinbin Duan, Huiwu Tian, Daqing Chen. Population genetic structure of Pelteobagrus vachelli in the upper Yangtze River based on genome re-sequencing [J]. Biodiv Sci, 2023, 31(4): 22391-. |
| [13] | Yuanyuan Xiao, Wei Feng, Yangui Qiao, Yuqing Zhang, Shugao Qin. Effects of soil microbial community characteristics on soil multifunctionality in sand-fixation shrublands [J]. Biodiv Sci, 2023, 31(4): 22585-. |
| [14] | Yiyue He, Yuying Liu, Fubin Zhang, Qiang Qin, Yu Zeng, Zhenyu Lü, Kun Yang. Genetic diversity and population structure of Saurogobio dabryi under cascade water conservancy projects in the Jialing River [J]. Biodiv Sci, 2023, 31(11): 23160-. |
| [15] | Weiyue Sun, Jiangping Shu, Yufeng Gu, Morigengaowa, Xiajin Du, Baodong Liu, Yuehong Yan. Conservation genomics analysis revealed the endangered mechanism of Adiantum nelumboides [J]. Biodiv Sci, 2022, 30(7): 21508-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||