

Chinese Bulletin of Botany ›› 2017, Vol. 52 ›› Issue (5): 550-559.DOI: 10.11983/CBB16208 cstr: 32102.14.CBB16208
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Gan Huang, Xiao Wang, Xuefeng Jin, Xiaojing Wang, Yaqin Wang*
Received:2016-11-04
Accepted:2017-03-06
Online:2017-09-01
Published:2017-07-10
Contact:
Yaqin Wang
Gan Huang, Xiao Wang, Xuefeng Jin, Xiaojing Wang, Yaqin Wang. GRXC9 Negatively Regulates Leaf Size in Arabidopsis[J]. Chinese Bulletin of Botany, 2017, 52(5): 550-559.
| Primer name | Primer sequences (5'-3') |
|---|---|
| GRXC9-F | AACGATTTCTTGCCCGGGTTATGCAAGG |
| GRXC9-R | GGATCCTCACAACCACAGAGCCCCAACT |
| LB | GCCTTTTCAGAAATGGATAAATAGCCTT- GCTTCC |
| GRXC9-LP | GGAAGAAATGGGTGACATGAG |
| GRXC9-RP | TCTTGCACAAGAAATCGTTCC |
| ROT3-F | AGATTTCGTCAGCGGAAAGA |
| ROT3-R | CCAAAGGGTGTGAAGCAAAT |
| LNG1-F | ATGGAGAAGACGCAGCATT |
| LNG1-R | GACTGCTTCTCGAACCCAAG |
| LNG2-F | GAAGGAAGAGGAGCGGCTAT |
| LNG2-R | CCGCTTCTGAATTTCACCAT |
| AN-F | AAACCTGGGGCTTTTCTTGT |
| AN-R | CCTGTTGCCTACTGGTGGAT |
| ACTIN-F | CTACGAGCAGGAACTCGAGA |
| ACTIN-R | GATGGACCTGACTCGTCATAC |
Table 1 The sequences of primers
| Primer name | Primer sequences (5'-3') |
|---|---|
| GRXC9-F | AACGATTTCTTGCCCGGGTTATGCAAGG |
| GRXC9-R | GGATCCTCACAACCACAGAGCCCCAACT |
| LB | GCCTTTTCAGAAATGGATAAATAGCCTT- GCTTCC |
| GRXC9-LP | GGAAGAAATGGGTGACATGAG |
| GRXC9-RP | TCTTGCACAAGAAATCGTTCC |
| ROT3-F | AGATTTCGTCAGCGGAAAGA |
| ROT3-R | CCAAAGGGTGTGAAGCAAAT |
| LNG1-F | ATGGAGAAGACGCAGCATT |
| LNG1-R | GACTGCTTCTCGAACCCAAG |
| LNG2-F | GAAGGAAGAGGAGCGGCTAT |
| LNG2-R | CCGCTTCTGAATTTCACCAT |
| AN-F | AAACCTGGGGCTTTTCTTGT |
| AN-R | CCTGTTGCCTACTGGTGGAT |
| ACTIN-F | CTACGAGCAGGAACTCGAGA |
| ACTIN-R | GATGGACCTGACTCGTCATAC |
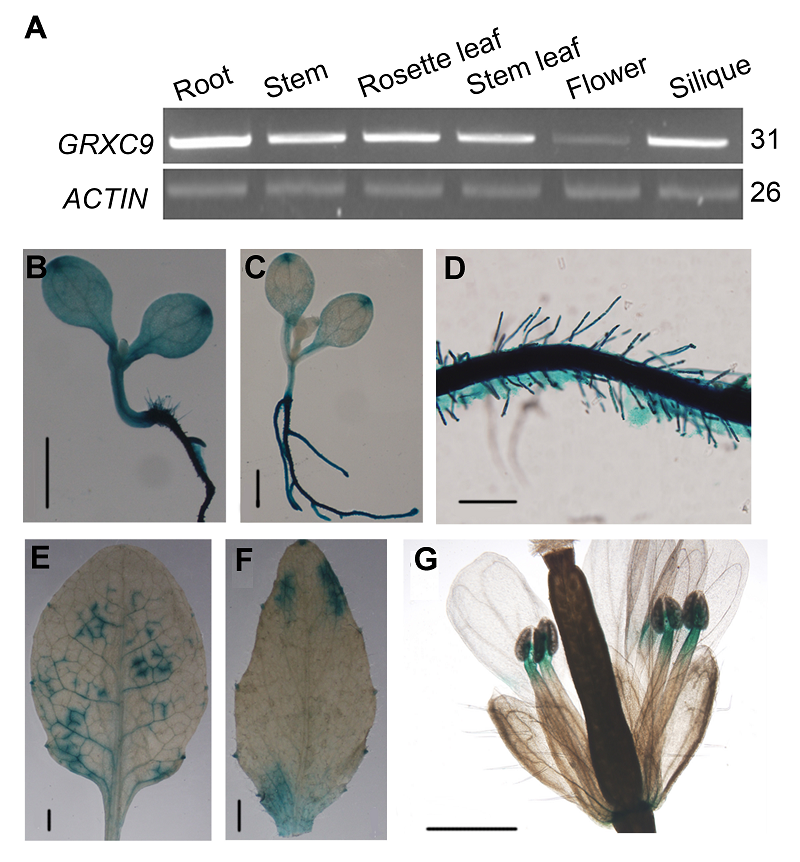
Figure 1 Tissue-specific expression of GRXC9 in Arabi- dopsis thaliana(A) Expression level of GRXC9 in root, stem, rosette leaf, stem leaf, flower and silique (ACTIN was used as internal reference; The number of PCR cycle was listed on the right of the images); (B)-(G) Histochemical localization of GUS activity directed by GRXC9::GUS fusions ((B) Cotyledon stage (Bar=1 mm); (C) Four-leaf stage (Bar=1 mm); (D) Root (Bar=0.1 mm); (E) Rosette leaf; (F) Stem leaf; (G) Flower). (E)-(G) Bar=1 mm
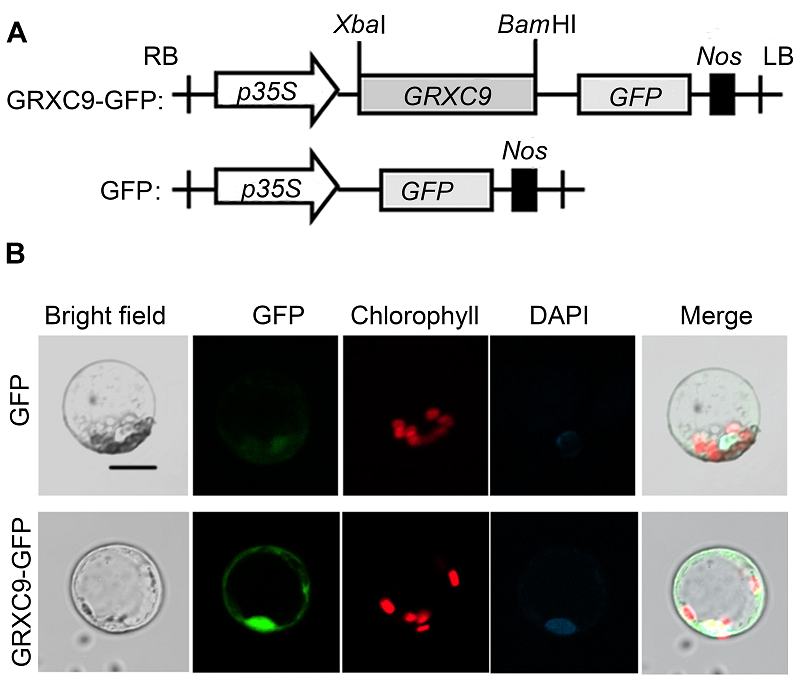
Figure 2 Subcellular localization of GRXC9 of Arabidop- sis thaliana(A) Schematic representation of the p35S::GRXC9-GFP construct; (B) Co-localization of GFP fused to GRXC9 (GFP indicates confocal images under the GFP channel; Bright field indicates confocal images of the same cells with transmitted light; DAPI indicates images under the DAPI channel; Merge indicates the merged images of GFP confocal images and transmitted light) (Bar=10 μm).
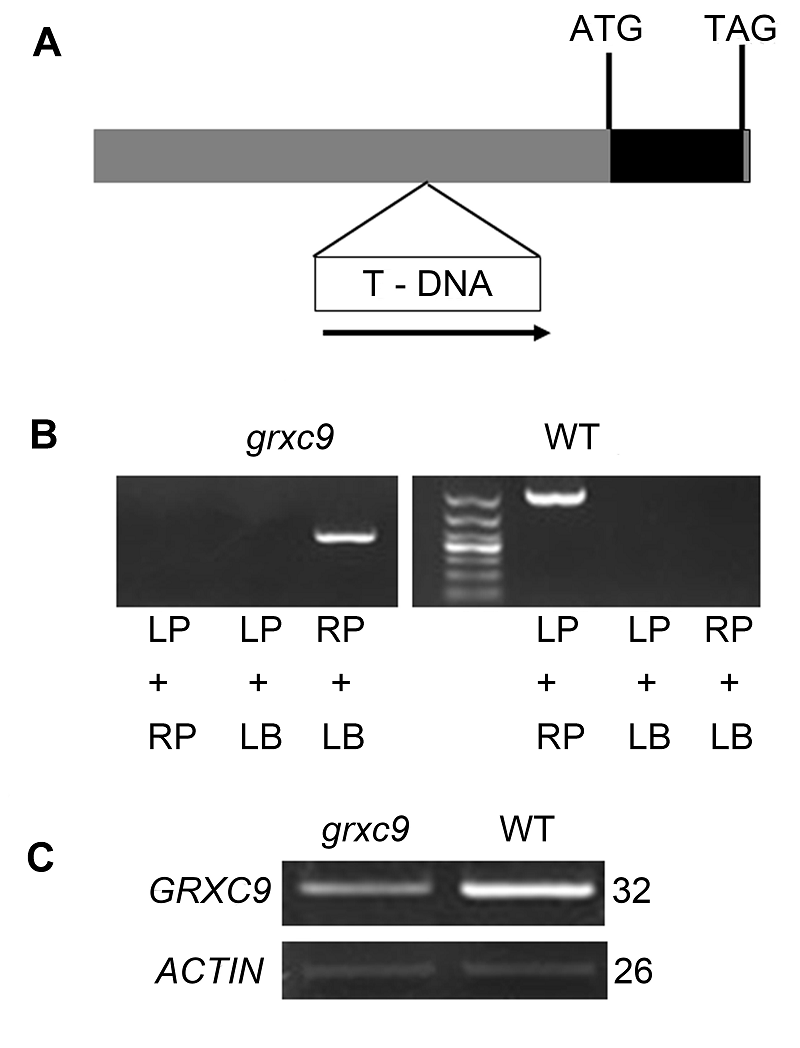
Figure 3 Molecular identification of Arabidopsis grxc9 mu- tant(A) T-DNA insertion sites in grxc9; (B) Identification of grxc9 using the three primer method; (C) Reverse trans- criptase PCR analysis of GRXC9 expression in WT and grxc9
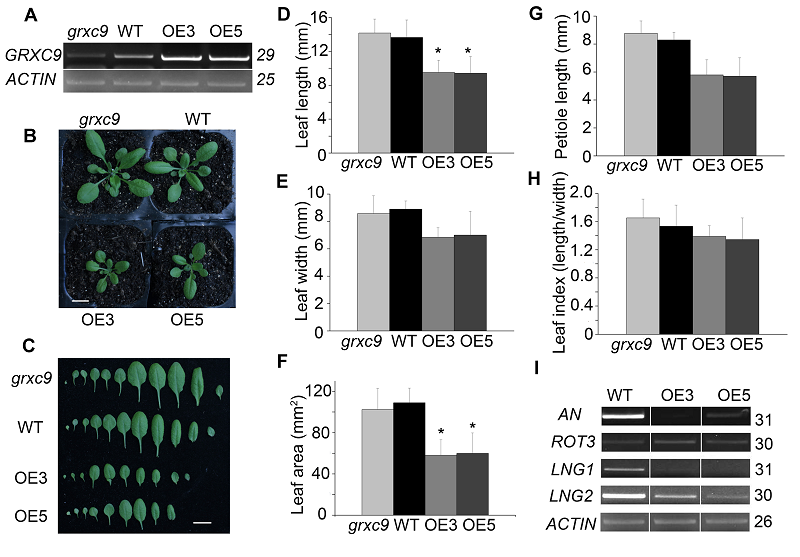
Figure 4 Phenotypic analysis and expression level of leaf size related genes in three genotypes of Arabidopsis thaliana(A) Identification of GRXC9 gene expression in grxc9, 35S::GRXC9 and wild type; (B) The phenotype of three kinds of genotypes after 28 days grown under long-day conditions (Bar=1 cm); (C) The comparison of representative leaves among grxc9, 35S::GRXC9 and wild type (Bar=1 cm); (D)-(H) Statistical analysis of leaf blade length (D), width (E), area (F), leaf petiole length (G) and leaf index (H) among grxc9, two transgenic plants and wild type (Value=means±SE, n >10, * Student’s t-test significant difference compared with WT plants (P<0.05) ); (I) Expression level of leaf size related genes
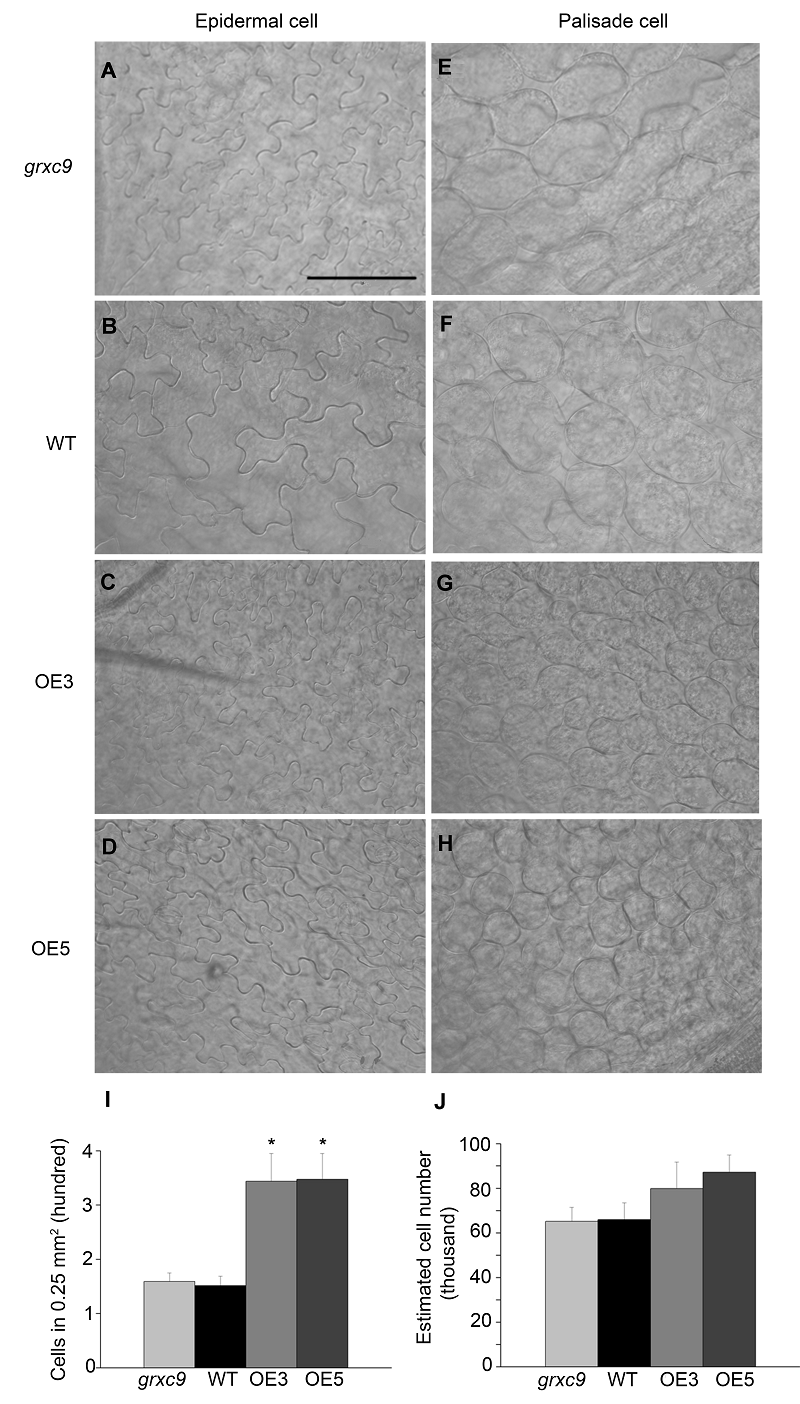
Figure 5 Morphological characterization of epidermal cells and palisade cells in three genotypes of Arabidopsis thaliana(A)-(D) Morphological characterization of epidermal cells in grxc9, two transgenic plants and wild type; (E)-(H) Morphological characterization of palisade cells in grxc9, two transgenic plants and wild type; (I) The statistics of palisade cells in a 0.25 mm2 area of grxc9, two transgenic plants and wild type; (J) Based on average cell number per 0.25 mm2 area and leaf area, the total number of the first layer of palisade cells was estimated (Value=means±SE, n >10, * Student’s t-test significant difference compared with WT plants (P<0.05))
| [1] | 李凌飞, 彭建宗, 王小菁 (2015). 非洲菊微管相关蛋白基因GMAP65-1功能分析. 植物学报 50, 12-21. |
| [2] | Bai Y, Falk S, Schnittger A, Jakoby MJ, Hülskamp M (2010). Tissue layer specific regulation of leaf length and width in Arabidopsis as revealed by the cell autonomous action of ANGUSTIFOLIA.Plant J 61, 191-199. |
| [3] | Bar M, Ori N (2015). Compound leaf development in model plant species.Curr Opin Plant Biol 23, 61-69. |
| [4] | Barkoulas M, Galinha C, Grigg SP, Tsiantis M (2007). From genes to shape: regulatory interactions in leaf development.Curr Opin Plant Biol 10, 660-666. |
| [5] | Byrne ME (2005). Networks in leaf development.Curr Opin Plant Biol 8, 59-66. |
| [6] | Cheng NH, Hirschi KD (2003). Cloning and characterization of CXIP1, a novel PICOT domain-containing Arabidopsis protein that associates with CAX1.J Biol Chem 278, 6503-6509. |
| [7] | Cheng NH, Liu JZ, Brock A, Nelson RS, Hirschi KD (2006). AtGRXcp, an Arabidopsis chloroplastic glutare- doxin, is critical for protection against protein oxidative da- mage.J Biol Chem 281, 26280-26288. |
| [8] | Clough SJ, Bent AF (1998). Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidop- sis thaliana.Plant J 16, 735-743. |
| [9] | Davies PJ (1995). The plant hormones: their nature, occurr- ence, and functions. In: Davies PJ, ed. Plant Hormones. Berlin: Springer. pp. 1-12. |
| [10] | Folkers U, Kirik V, Schöbinger U, Falk S, Krishnakumar S, Pollock MA, Oppenheimer DG, Day I, Reddy AR, Jür- gens G, Hülskamp M (2002). The cell morphogenesis gene ANGUSTIFOLIA encodes a CtBP/BARS-like protein and is involved in the control of the microtubule cytosk- eleton.EMBO J 21, 1280-1288. |
| [11] | Hanson J, Johannesson H, Engström P (2001). Sugar- dependent alterations in cotyledon and leaf development in transgenic plants expressing the HDZhdip gene ATHB- 13.Plant Mol Biol 45, 247-262. |
| [12] | Herrera-Vásquez A, Carvallo L, Blanco F, Tobar M, Villarroel-Candia E, Vicente-Carbajosa J, Salinas P, Holuigue L (2015). Transcriptional control of glutaredoxin GRXC9 expression by a salicylic acid-dependent and NPR1-independent pathway in Arabidopsis.Plant Mol Biol Rep 33, 624-637. |
| [13] | Holmgren A (1976). Hydrogen donor system for Escherichia coli ribonucleoside-diphosphate reductase dependent up- on glutathione.Proc Natl Acad Sci USA 73, 2275-2279. |
| [14] | Holmgren A (1979a). Glutathione-dependent synthesis of deoxyribonucleotides. Characterization of the enzymatic mechanism of Escherichia coli glutaredoxin.J Biol Chem 254, 3672-3678. |
| [15] | Holmgren A (1979b). Glutathione-dependent synthesis of deoxyribonucleotides. Purification and characterization of glutaredoxin from Escherichia coli.J Biol Chem 254, 3664-3671. |
| [16] | Horiguchi G, Kim GT, Tsukaya H (2005). The transcription factor AtGRF5 and the transcription coactivator AN3 re- gulate cell proliferation in leaf primordia of Arabidopsis tha- liana.Plant J 43, 68-78. |
| [17] | Kim GT, Shoda K, Tsuge T, Cho KH, Uchimiya H, Yokoyama R, Nishitani K, Tsukaya H (2002). The ANGUSTIFOLIA gene of Arabidopsis, a plant CtBP gene, regulates leaf-cell expansion, the arrangement of cortical mi- crotubules in leaf cells and expression of a gene involved in cell-wall formation.EMBO J 21, 1267-1279. |
| [18] | Kim GT, Tsukaya H, Uchimiya H (1998). The ROTUNDI- FOLIA3 gene of Arabidopsis thaliana encodes a new member of the cytochrome P-450 family that is required for the regulated polar elongation of leaf cells.Genes Dev 12, 2381-2391. |
| [19] | Kim JH, Kende H (2004). A transcriptional coactivator, AtGIF1, is involved in regulating leaf growth and morpho- logy in Arabidopsis.Proc Natl Acad Sci USA 101, 13374-13379. |
| [20] | Kondorosi E, Roudier F, Gendreau E (2000). Plant cell- size control: growing by ploidy?Curr Opin Plant Biol 3, 488-492. |
| [21] | Laporte D, Olate E, Salinas P, Salazar M, Jordana X, Holuigue L (2012). Glutaredoxin GRXS13 plays a key role in protection against photooxidative stress in Arabidopsis.J Exp Bot 63, 503-515. |
| [22] | Lee YK, Kim GT, Kim IJ, Park J, Kwak SS, Choi G, Chung WI (2006). LONGIFOLIA1 and LONGIFOLIA2, two homologous genes, regulate longitudinal cell elongation in Arabidopsis.Development 133, 4305-4314. |
| [23] | Lemaire SD (2004). The glutaredoxin family in oxygenic photosynthetic organisms.Photosynth Res 79, 305-318. |
| [24] | Lillig CH, Berndt C, Holmgren A (2008). Glutaredoxin systems.Biochim Biophys Acta 1780, 1304-1317. |
| [25] | Meyer Y, Buchanan BB, Vignols F, Reichheld JP (2009). Thioredoxins and glutaredoxins: unifying elements in redox biology.Annu Rev Genet 43, 335-367. |
| [26] | Mizukami Y (2001). A matter of size: developmental control of organ size in plants.Curr Opin Plant Biol 4, 533-539. |
| [27] | Mizukami Y, Fischer RL (2000). Plant organ size control: AINTEGUMENTA regulates growth and cell numbers during organogenesis.Proc Natl Acad Sci USA 97, 942-947. |
| [28] | Murmu J, Bush MJ, DeLong C, Li ST, Xu ML, Khan M, Malcolmson C, Fobert PR, Zachgo S, Hepworth SR (2010). Arabidopsis basic leucine-zipper transcription factors TGA9 and TGA10 interact with floral glutaredoxins ROXY1 and ROXY2 and are redundantly required for anther development.Plant Physiol 154, 1492-1504. |
| [29] | Narita NN, Moore S, Horiguchi G, Kubo M, Demura T, Fukuda H, Goodrich J, Tsukaya H (2004). Overexpres- sion of a novel small peptide ROTUNDIFOLIA4 decre- ases cell proliferation and alters leaf shape in Arabidopsis thaliana.Plant J 38, 699-713. |
| [30] | Ndamukong I, Al Abdallat A, Thurow C, Fode B, Zander M, Weigel R, Gatz C (2007). SA-inducible Arabidopsis glutaredoxin interacts with TGA factors and suppresses JA-responsive PDF1.2 transcription.Plant J 50, 128-139. |
| [31] | Qiu JL, Jilk R, Marks MD, Szymanski DB (2002). The Ara- bidopsis SPIKE1 gene is required for normal cell shape control and tissue development.Plant Cell 14, 101-118. |
| [32] | Rouhier N, Couturier J, Jacquot JP (2006). Genome-wide analysis of plant glutaredoxin systems.J Exp Bot 57, 1685-1696. |
| [33] | Rouhier N, Gelhaye E, Jacquot JP (2004). Plant glutaredoxins: still mysterious reducing systems.Cell Mol Life Sci 61, 1266-1277. |
| [34] | Rouhier N, Lemaire SD, Jacquot JP (2008). The role of glutathione in photosynthetic organisms: emerging functions for glutaredoxins and glutathionylation.Annu Rev Plant Biol 59, 143-166. |
| [35] | Sugimoto-Shirasu K, Roberts K (2003). “Big it up”: endoreduplication and cell-size control in plants.Curr Opin Plant Biol 6, 544-553. |
| [36] | Tsuge T, Tsukaya H, Uchimiya H (1996). Two independent and polarized processes of cell elongation regulate leaf blade expansion in Arabidopsis thaliana (L) Heynh.Development 122, 1589-1600. |
| [37] | Tsukaya H (2005). Leaf shape: genetic controls and envi- ronmental factors.Inter J Dev Biol 49, 547-555. |
| [38] | Tsukaya H (2006). Mechanism of leaf-shape determination.Annu Rev Plant Biol 57, 477-496. |
| [39] | Verma PK, Verma S, Pande V, Mallick S, Tripathi RD, Dhankher OP, Chakrabarty D (2016). Overexpression of rice glutaredoxin OsGrx_C7 and OsGrx_C2.1 reduces intracellular arsenic accumulation and increases tolerance in Arabidopsis thaliana.Front Plant Sci 7, 740. |
| [40] | Vernoux T, Autran D, Traas J (2000). Developmental control of cell division patterns in the shoot apex. In: Inzé D, ed. The Plant Cell Cycle. Dordrecht: Springer. pp. 25-37. |
| [41] | Xing S, Lauri A, Zachgo S (2006). Redox regulation and flower development: a novel function for glutaredoxins.Plant Biol 8, 547-555. |
| [42] | Xing SP, Rosso MG, Zachgo S (2005). ROXY1, a member of the plant glutaredoxin family, is required for petal deve- lopment in Arabidopsis thaliana.Development 132, 1555-1565. |
| [43] | Xing SP, Zachgo S (2008). ROXY1 and ROXY2, two Arabidopsis glutaredoxin genes, are required for anther development.Plant J 53, 790-801. |
| [44] | Yoo SD, Cho YH, Sheen J (2007). Arabidopsis mesophyll protoplasts: a versatile cell system for transient gene expression analysis.Nat Protoc 2, 1565-1572. |
| [45] | Zander M, Chen SX, Imkampe J, Thurow C, Gatz C (2012). Repression of the Arabidopsis thaliana jasmonic acid/ethylene-induced defense pathway by TGA-interac- ting glutaredoxins depends on their C-terminal ALWL motif.Mol Plant 5, 831-840. |
| [46] | Zhang SD, Shen ZJ, Liu XM, Li Z, Zhang QW, Liu XX (2016). Molecular identification of three novel glutaredoxin genes that play important roles in antioxidant defense in Helicoverpa armigera.Insect Biochem Mol Biol 75, 107-116. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||