

石刁柏核质体DNA的生物信息学分析及染色体定位
收稿日期: 2018-05-11
录用日期: 2018-08-06
网络出版日期: 2018-12-07
基金资助
国家自然科学基金(31470334和河南省高校科技创新团队支持计划No.17IRTSTHN017)
Bioinformatics Analysis and Chromosome Location of Nuclear Integrants of Plastid DNA in Asparagus officinalis
Received date: 2018-05-11
Accepted date: 2018-08-06
Online published: 2018-12-07
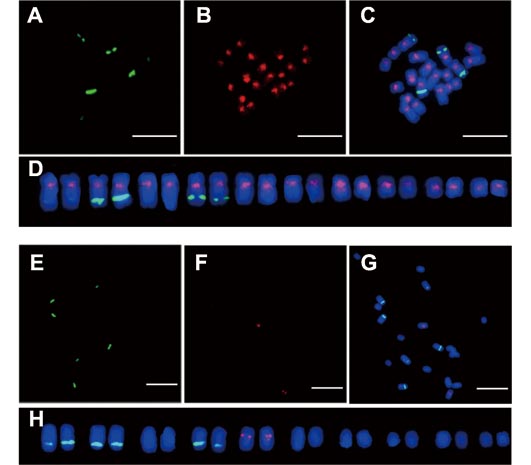
在植物基因组中, 叶绿体DNA (cpDNA)序列可以向核基因组转移成为核质体DNA (NUPT)。NUPTs在植物染色体(包括性染色体)的演化过程中具有重要作用, 但目前相关研究比较缺乏。以雌雄异株植物石刁柏(Asparagus officinalis)为材料, 采用生物信息学方法对其核基因组NUPTs进行注释及分析, 并选取叶绿体基因组反向重复区(IR) 2个片段进行染色体定位。结果表明, 石刁柏核基因组中有2 239个NUPTs序列的插入, 总长度为565 970 bp, 占核基因组的0.047%。不同染色体上插入的NUPTs数量存在较大差异, Y染色体上的NUPTs数量、密度及总长度均高于其它染色体, 表明NUPTs在石刁柏性(Y)染色体上累积的更多。石刁柏叶绿体基因组中的IR区、大单拷贝区(LSC)和小单拷贝区(SSC)序列均能够向核基因组转移, 但IR区序列转移频率更高。此外, 对2个IR区的叶绿体序列进行荧光原位杂交, 其中AocpIR1主要分布在所有染色体的着丝粒部位, 而AocpIR2特异性分布在Y染色体上。研究结果为深入揭示石刁柏基因组的结构及其性染色体的演化奠定了坚实的基础。
程广前,贾克利,李娜,邓传良,李书粉,高武军 . 石刁柏核质体DNA的生物信息学分析及染色体定位[J]. 植物学报, 2019 , 54(3) : 328 -334 . DOI: 10.11983/CBB18117
Chloroplast DNA (cpDNA) can transfer into the plant nuclear genome to form nuclear integrants of plastid DNA (NUPTs). NUPTs may play a role in plant sex chromosome evolution. However, few studies have focused on this research area. In this study, we annotated and analyzed NUPTs in the genome of dioecious Asparagus officinalis and located two cpDNA fragments on chromosomes. The nuclear genome of A. officinalis contains 2 239 NUPT insertions, with total length 565 970 bp, accounting for 0.047% of the genome. The amount of NUPTs differs among chromosomes; the number, density, and the average length of NUPTs on the Y chromosome were all higher than those on the other chromosomes, which indicates more accumulation of NUPTs on sex chromosomes. All regions of inverted repeats (IRs) and small and large single copy regions of cpDNA could transfer into nuclear DNA; however, the IR region showed the highest transfer frequency. Furthermore, FISH analysis of two cpDNA sequences from IR regions showed that AocpIR1 distributed mainly on the centromeres of all chromosomes, whereas AocpIR2 specifically located on sex chromosomes of A. officinalis. The data provide an important foundation for determining the genome structure and sex chromosome evolution of A. officinalis.

| [1] | 高东迎, 何冰, 孙立华 ( 2007). 水稻转座子研究进展. 植物学通报 24, 667-676. |
| [2] | 李巧丽, 延娜, 宋琼, 郭军战 ( 2018). 鲁桑叶绿体基因组序列及特征分析. 植物学报 53, 94-103. |
| [3] | 李书粉, 李旭, 王冰肖, 袁金红, 邓传良, 高武军 ( 2016). 石刁柏雄性偏向核质体DNA的克隆与分析. 西北植物学报 36, 2385-2390. |
| [4] | Abbott JK, Nordén AK, Hansson B ( 2017). Sex chromosome evolution: historical insights and future perspectives. Proc Roy Soc B 284, 20162806. |
| [5] | Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ ( 1990). Basic local alignment search tool. J Mol Biol 215, 403-410. |
| [6] | Doyle JJ, Doyle JL ( 1987). A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 19, 11-15. |
| [7] | Feschotte C, Pritham EJ ( 2007). DNA transposons and the evolution of eukaryotic genomes. Annu Rev Genet 41, 331-368. |
| [8] | Guo XY, Ruan SL, Hu WM, Cai DG, Fan LJ ( 2008). Chloroplast DNA insertions into the nuclear genome of rice: the genes, sites and ages of insertion involved. Funct Integr Genomics 8, 101-108. |
| [9] | Harkess A, Zhou JS, Xu CY, Bowers JE, van der Hulst R, Ayyampalayam S, Mercati F, Riccardi P, McKain MR, Kakrana A, Tang HB, Ray J, Groenendijk J, Arikit S, Mathioni SM, Nakano M, Shan HY, Telgmann-Rauber A, Kanno A, Yue Z, Chen HX, Li WQ, Chen YL, Xu XY, Zhang YP, Luo SC, Chen HL, Gao JM, Mao ZC, Pires JC, Luo MZ, Kudrna D, Wing RA, Meyers BC, Yi KX, Kong HZ, Lavrijsen P, Sunseri F, Falavigna A, Ye Y, Leebens-Mack JH, Chen GY ( 2017). The asparagus genome sheds light on the origin and evolution of a young Y chromosome. Nat Commun 8, 1279. |
| [10] | Kejnovsky E, Hobza R, Cermak T, Kubat Z, Vyskot B ( 2009). The role of repetitive DNA in structure and evolution of sex chromosomes in plants. Heredity 102, 533-541. |
| [11] | Kejnovsky E, Kubat Z, Hobza R, Lengerova M, Sato S, Tabata S, Fukui K, Matsunaga S, Vyskot B ( 2006). Accumulation of chloroplast DNA sequences on the Y chromosome of Silene latifolia. Genetica 128, 167-175. |
| [12] | Kleine T, Maier UG, Leister D ( 2009). DNA transfer from organelles to the nucleus: the idiosyncratic genetics of endosymbiosis. Annu Rev Plant Biol 60, 115-138. |
| [13] | Li SF, Zhang GJ, Yuan JH, Deng CL, Gao WJ ( 2016). Repetitive sequences and epigenetic modification: inseparable partners play important roles in the evolution of plant sex chromosomes. Planta 243, 1083-1095. |
| [14] | L?ptien H ( 1979). Identification of the sex chromosome pair in asparagus ( Asparagus officinalis L.). Z Pflanzenzüecht 82, 162-173. |
| [15] | Martis MM, Klemme S, Banaei-Moghaddam AM, Blattner FR, Macas J, Schmutzer T, Scholz U, Gundlach H, Wicker T, ?imková H, Novák P, Neumann P, Kubaláková M, Bauer E, Haseneyer G, Fuchs J, Dole?el J, Stein N, Mayer KFX, Houben A ( 2012). Selfish supernumerary chromosome reveals its origin as a mosaic of host genome and organellar sequences. Proc Natl Acad Sci USA 109, 13343-13346. |
| [16] | Matsuo M, Ito Y, Yamauchi R, Obokata J ( 2005). The rice nuclear genome continuously integrates, shuffles, and eli- minates the chloroplast genome to cause chloroplast- nuclear DNA flux. Plant Cell 17, 665-675. |
| [17] | Michalovova M, Vyskot B, Kejnovsky E ( 2013). Analysis of plastid and mitochondrial DNA insertions in the nucleus (NUPTs and NUMTs) of six plant species: size, relative age and chromosomal localization. Heredity 111, 314-320. |
| [18] | Richly E, Leister D ( 2004). NUMTs in sequenced eukaryotic genomes. Mol Biol Evol 21, 1081-1084. |
| [19] | Sheng WT, Chai XW, Rao YS, Tu XT, Du SG ( 2017). Complete chloroplast genome sequence of asparagus ( Asparagus officinalis L.) and its phylogenetic position within asparagales. J Plant Breed Genet 5, 121-128. |
| [20] | Steflova P, Hobza R, Vyskot B, Kejnovsky E ( 2014). Strong accumulation of chloroplast DNA in the Y chromosomes of Rumex acetosa and Silene latifolia. Cytogenet Genome Res 142, 59-65. |
| [21] | Timmis JN, Ayliffe MA, Huang CY, Martin W ( 2004). Endosymbiotic gene transfer: organelle genomes forge eukaryotic chromosomes. Nat Rev Genet 5, 123-135. |
| [22] | VanBuren R, Ming R ( 2013). Organelle DNA accumulation in the recently evolved papaya sex chromosomes. Mol Genet Genomics 288, 277-284. |
| [23] | Wang RJ, Cheng CL, Chang CC, Wu CL, Su TM, Chaw SM ( 2008). Dynamics and evolution of the inverted repeat-large single copy junctions in the chloroplast genomes of monocots. BMC Evol Biol 8, 36. |
| [24] | Yoshida T, Furihata HY, Kawabe A ( 2014). Patterns of genomic integration of nuclear chloroplast DNA fragments in plant species. DNA Res 21, 127-140. |
/
| 〈 |
|
〉 |