

鲁桑叶绿体基因组序列及特征分析
收稿日期: 2016-12-13
录用日期: 2017-03-29
网络出版日期: 2017-03-29
基金资助
西北农林科技大学唐仲英育种基金(No.2013-14)
Complete Chloroplast Genome Sequence and Characteristics Analysis of Morus multicaulis
Received date: 2016-12-13
Accepted date: 2017-03-29
Online published: 2017-03-29
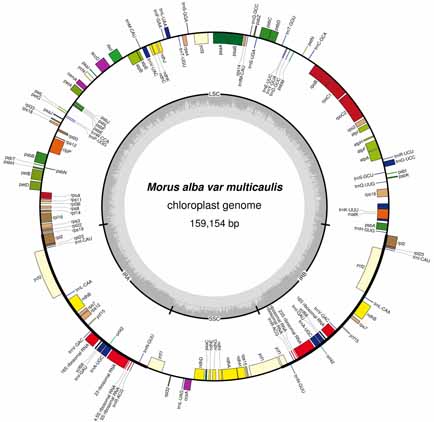
鲁桑(Morus multicaulis)是亚洲地区栽培的重要经济作物。以鲁桑品种日本胡橙为实验材料, 利用高通量测序技术对鲁桑叶绿体基因组进行测序, 获得NCBI登录号(KU355297), 并研究鲁桑的叶绿体基因组结构。结合前人对蒙桑(M. mongolica)、印度桑(M. indica)和川桑(M. notabilis)的研究结果, 对鲁桑的系统进化关系进行了探讨。研究结果表明: 鲁桑叶绿体基因组是一个典型的四部分结构, 全长159 154 bp, 共注释130个基因, 包含85个蛋白质编码基因(18个基因在反向重复区重复)、37个转运RNA (tRNA)基因和8个核糖体RNA (rRNA)基因。生物信息学分析表明, 在鲁桑中共搜索到82个SSR位点, 单核苷酸、二核苷酸、三核苷酸、四核苷酸和五核苷酸重复基序个数分别为63、7、2、9和1个, 并没有发现六核苷酸; 其中单核苷酸重复在鲁桑的叶绿体基因组SSR中占76.8%。采用MEGA 6.0软件, 通过最大似然法和近邻结合法对包括4个桑属物种在内的15个物种的叶绿体基因组序列进行聚类分析, 2种方法得到的聚类结果均为鲁桑和蒙桑聚在一起。研究结果对叶绿体基因组工程研究及桑属种间的分子标记开发和优良品种培育具有一定的参考价值。
李巧丽 , 延娜 , 宋琼 , 郭军战 . 鲁桑叶绿体基因组序列及特征分析[J]. 植物学报, 2018 , 53(1) : 94 -103 . DOI: 10.11983/CBB16247
Mulberry is an economically important crop in Asia. We determined the complete chloroplast sequence of cultivated species of Morus multicaulis. Ribenhuchen was used as experimental material. High-throughput sequencing was used to sequence the chloroplast genome and the genome structure (NCBI No.: KU355297), and we compared the chloroplast genome with those of reported sibling species (Morus mongolica, M. indica, M. notabilis). The chloroplast genome (cpDNA) of M. multicaulis with a typical quadripartite structure is 159 154 bp long. The cpDNA of M. multicaulis contains 130 genes, including 85 protein coding genes (18 genes duplicated in the inverted repeat regions), 37 transfer RNA genes and 8 ribosomal RNA genes. There are 82 simple sequence repeats, and the number of mono-, di-, tri-, tetra-, pentanucleotide repeat motifs is 63, 7, 2, 9, and 1, with no hexanucleotide repeat sequences. Mono-nucleotide repeat sequences accounted for 76.8% of the cpDNA of simple sequence repeats. MEGA 6.0 was used to construct the phylogenetic tree of 15 species and for cluster analysis of Morus plants. M. multicaulis and M. mongolica were clustered into one group. The research results have reference value for chloroplast genome research, molecular marker development and breeding of mulberry.

| [1] | 冯丽春, 杨光伟, 余茂德, 张孝勇, 向怀祥 (1997). 利用RAPD对桑属植物种间亲缘关系的研究. 中国农业科学 30, 52-56. |
| [2] | 黄瑶, 李朝銮, 马诚, 吴乃虎 (1994). 叶绿体DNA及其在植物系统学研究中的应用. 植物学通报 11(2), 11-25. |
| [3] | 徐军望, 冯德江, 宋贵生, 魏晓丽, 陈蕾, 伍晓丽, 李旭刚, 朱桢 (2003). 水稻EPSP合酶第一内含子增强外源基因的表达. 中国科学(C辑) 33, 224-230. |
| [4] | 闫化学, 于杰 (2010). DNA条形码技术在植物中的研究现状. 植物学报 45, 102-108. |
| [5] | 周德贵, 赵琼一, 付崇允, 李宏, 蔡学飞, 罗达, 周少川 (2008). 新一代测序技术及其对水稻分子设计育种的影响. 分子植物育种 6, 619-630. |
| [6] | Allender CJ, Allainguillaume J, Lynn J, King GJ (2007). Simple sequence repeats reveal uneven distribution of genetic diversity in chloroplast genomes of Brassica ole- racea L. and(n=9) wild relatives. Theor Appl Genet 114, 609-618. |
| [7] | Chen C, Zhou W, Huang Y, Wang ZZ (2015). The complete chloroplast genome sequence of the mulberry Morus notabilis(Moreae). Mitochondrial DNA Part A 27, 2856-2857. |
| [8] | Flannery ML, Mitchell FJG, Coyne S, Kavanagh TA, Burke JI, Salamin N, Dowding P, Hodkinson TR (2006). Plastid genome characterisation in Brassica and Brassicaceae using a new set of nine SSRs. Theor Appl Genet 113, 1221-1231. |
| [9] | George B, Bhatt BS, Awasthi M, George B, Singh AK (2015). Comparative analysis of microsatellites in chloroplast genomes of lower and higher plants.Curr Genet 61, 665-677. |
| [10] | Hebert PD, Ratnasingham S, de Waard JR (2003). Barcoding animal life: cytochrome c oxidase subunit 1 divergences among closely related species. Proc R Soc Lond B 270, S96-S99. |
| [11] | Huang YY, Matzke AJM, Matzke M (2013). Complete sequence and comparative analysis of the chloroplast genome of coconut palm ( Cocos nucifera). PLoS One 8, e74736. |
| [12] | Jansen RK, Raubeson LA, Boore JL, dePamphilis CW, Chumley TW, Haberle RC, Wyman SK, Alverson AJ, Peery R, Herman SJ, Fourcade HM, Kuehl JV, McNeal JR, Leebens-Mack J, Cui LY (2005). Methods for obtaining and analyzing whole chloroplast genome sequen- ces.Methods Enzymol 395, 348-384. |
| [13] | Jiao Y, Jia HM, Li XW, Jia HJ, Chen Z, Wang GY, Chai CY, van de Weg E, Gao ZS (2012). Development of simple sequence repeat (SSR) markers from a genome survey of Chinese Bayberry ( Myrica rubra). BMC Genomics 13, 201. |
| [14] | Katti MV, Ranjekar PK, Gupta VS (2001). Differential distribution of simple sequence repeats in eukaryotic genome sequences.Mol Biol Evol 18, 1161-1167. |
| [15] | Kaundun SS, Matsumoto S (2002). Heterologous nuclear and chloroplast microsatellite amplification and variation in tea, Camellia sinensis. Genome 45, 1041-1048. |
| [16] | Kong WQ, Yang JH (2015). The complete chloroplast genome sequence of Morus mongolica and a comparative analysis within the Fabidae clade. Curr Genet 62, 165-172. |
| [17] | Leigh FJ, Mackay I, Oliveira HR, Gosman NE, Horsnell RA, Jones H, White J, Powell W, Brown TA (2013). Using diversity of the chloroplast genome to examine evolutionary history of wheat species.Genet Resour Crop Evol 60, 1831-1842. |
| [18] | Leseberg CH, Duvall MR (2009). The complete chloroplast genome of Coix lacryma-jobi and a comparative molecular evolutionary analysis of plastomes in cereals. J Mol Evol 69, 311-318. |
| [19] | Nazareno AG, Carlsen M, Lohmann LG (2015). Complete chloroplast genome of Tanaecium tetragonolobum: the first Bignoniaceae plastome. PLoS One 10, e0129930. |
| [20] | Plunkett GM, Downie SR (2000). Expansion and contraction of the chloroplast inverted repeat in Apiaceae subfamily Apioideae.Syst Bot 25, 648-667. |
| [21] | Rajendrakumar P, Biswal AK, Balachandran SM, Srinivasarao K, Sundaram RM (2007). Simple sequence repeats in organellar genomes of rice: frequency and distribution in genic and intergenic regions.Bioinformatics 23, 1-4. |
| [22] | Ravi V, Khurana JP, Tyagi AK, Khurana P (2006). The chloroplast genome of mulberry: complete nucleotide sequence, gene organization and comparative analysis.Tree Genet Genomes 3, 49-59. |
| [23] | Ruhlman TA, Jansen RK (2014). The plastid genomes of flowering plants.Methods Mol Biol 1132, 3-38. |
| [24] | Shaw J, Lickey EB, Schilling EE, Small RL (2007). Comparison of whole chloroplast genome sequences to choose noncoding regions for phylogenetic studies in angiosperms: the tortoise and the hare lll.Am J Bot 94, 275-288. |
| [25] | Temnykh S, DeClerck G, Lukashova A, Lipovich L, Cartinhour S, McCouch S (2001). Computational and experimental analysis of microsatellites in rice ( Oryza sativa L.): frequency, length variation, transposon associations, and genetic marker potential. Genome Res 11, 1441-1452. |
| [26] | Zhang HY, Li C, Miao HM, Xiong SJ (2013). Insights from the complete chloroplast genome into the evolution of Se- samum indicum L. PLoS One 8, e80508. |
/
| 〈 |
|
〉 |