

黑涩楠叶绿体全基因组的结构和比较分析及系统进化推断
收稿日期: 2024-09-24
录用日期: 2025-02-09
网络出版日期: 2025-02-10
基金资助
新优植物品种引进及推广示范(FY2024013)
Structural and Comparative Analysis of the Complete Chloroplast Genome of the Aronia melanocarpa and Its Phylogenetic Inference
Received date: 2024-09-24
Accepted date: 2025-02-09
Online published: 2025-02-10

王传永 , 庄典 , 宋正达 , 翟恒华 , 李乃伟 , 张凡 . 黑涩楠叶绿体全基因组的结构和比较分析及系统进化推断[J]. 植物学报, 2025 , 60(4) : 573 -585 . DOI: 10.11983/CBB24146
INTRODUCTION Aronia melanocarpa also known as black chokeberry, belongs to the genus Aronia (Rosaceae). In addition to A. melanocarpa, Aronia includes A. arbutifolia or red chokeberry and A. prunifolia or purple chokeberry, both distributed naturally in North American, and an additional cultivated taxon, A. mitschurinii or Mitschurin’s chokeberry, originating from Europe. However, the species boundaries and relationships among the species of Aronia are not clear. Moreover, the taxonomic history of Aroniais complex, as species of this genus have formerly been placed in many different genera, such as Mespilus, Pyrus, Adenorachis, Sorbus, and Photinia. In the present study, we first sequenced and characterized the complete chloroplast (cp) genome of A. melanocarpa and compared its sequence with those of the cp genomes from 13 species of the family Rosaceae. The aims of this study were: (1) to increase our understanding of the structural patterns of complete cp genome of A. melanocarpa; (2) to investigate the phylogenetic relationships of A. melanocarpa with other Rosaceae species based on their cp genomes.
RATIONALE The chloroplast is a unique and essential organelle in green plants with vital roles in photosynthesis and carbon fixation. Comparative analyses of cp genomes between different plant species reveal intra- and inter-species rearrangements that have occurred during evolution, such as inverted repeat (IR) contraction and expansion. Based on these characteristics, the cp genome has been wildly used for species identification, phylogenetic analysis, and exploring the genetic basis of environmental adaptation.
RESULTS The complete A. melanocarpa cp genome was sequenced, analyzed, and compared with that from 13 other species in the Rosaceae. The cp genome is 159 772 bp and has a total guanine-cytosine (GC) content of 36.6%. It exhibits a typical quadripartite structure with four separate regions, including a large single copy (LSC) region of 87 810 bp and a small single copy (SSC) region of 19 200 bp separated by two inverted repeats (IRa and IRb) regions of 26 381 bp each. A total of 132 genes were annotated, including 87 protein-coding genes, 37 tRNAs, and eight rRNAs, with 22 duplicates in the IR regions. In total, 76 simple sequence repeats (SSRs) and 50 long repeats were detected. Phylogenetic analysis indicated that A. melanocarpa is most closely related to A. arbutifolia and forms a sister clade to Cydonia oblonga with weak support.
CONCLUSION We analyzed the complete cp genome of A. melanocarpa by using Illumina high-throughput sequencing technology. The sequence of A. melanocarpa cp genome could be further used for the development of molecular markers. Highly variable regions were detected in intergenic regions, such as trnK-rps16, rps16-trnQ, trnG-atpA, petN-psbM, trnT-psbD, psbZ-trnG, trnT-trnL, ndhC-trnV and accD-psaI, which might be useful for broad applications in genetic research studies as well as phylogenetic studies. Phylogenetic construction results strongly supported that A. melanocarpa was closest related to A. arbutifolia, followed by C. oblonga with weak support. This newly available genomic data for A. melanocarpa will provide a basis for future research on the population genetics and phylogenomics and will benefit the breeding studies and utilization of the genus Aronia.
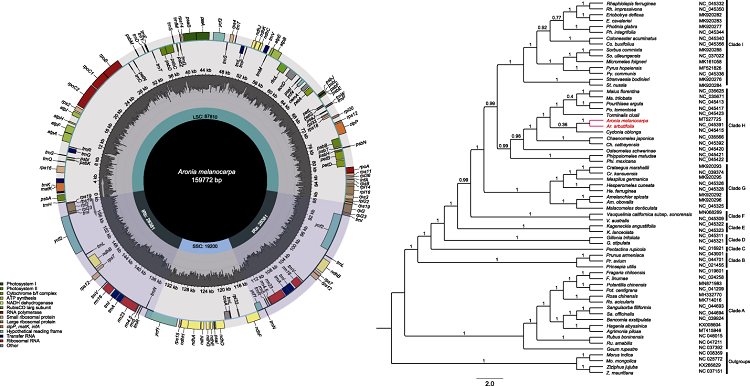
Map of the chloroplast genome of Aronia melanocarpa and phylogenetic analyses among the 60 Rosaceae species using their complete chloroplast genomes. Aronia formed a clade with Dichotomanthes and Pourthiaea based on cpDNA tree. Moreover, A. melanocarpa is most closely related to A. arbutifolia and forms a sister clade to Cydonia oblonga with weak support.

| [1] | Amiryousefi A, Hyv?nen J, Poczai P (2018). IRscope: an online program to visualize the junction sites of chloroplast genomes. Bioinformatics 34, 3030-3031. |
| [2] | Benson G (1999). Tandem repeats finder: a program to analyze DNA sequences. Nucleic Acids Res 27, 573-580. |
| [3] | Brand M (2010). Aronia: native shrubs with untapped potential. Arnoldia 67, 14-25. |
| [4] | Campbell CS, Evans RC, Morgan DR, Dickinson TA, Arsenault MP (2007). Phylogeny of subtribe Pyrinae (formerly the Maloideae, Rosaceae): limited resolution of a complex evolutionary history. Plant Syst Evol 266, 119-145. |
| [5] | Chiapella JO, Barfuss MHJ, Xue ZQ, Greimler J (2019). The plastid genome of Deschampsia cespitosa (Poaceae). Molecules 24, 216. |
| [6] | Chu ZZ, Gulbar Yisilam, Qu ZZ, Tian XM (2023). Comparative analyses on the chloroplast genome of three sympatric Atraphaxis species. Chin Bull Bot 58, 417-432. (in Chinese) |
| 褚振州, 古丽巴哈尔·依斯拉木, 屈泽众, 田新民 (2023). 同域分布的3种木蓼属植物叶绿体基因组比较. 植物学报 58, 417-432. | |
| [7] | Connolly BA (2014). Collection, Description, Taxonomic Relationships, Fruit Biochemistry, and Utilization of Aronia melanocarpa, A. arbutifolia, A. prunifolia, and A. mitschurinii. PhD dissertation. Connecticut: University of Connecticut. pp. 1-1. |
| [8] | Daniell H, Lin CS, Yu M, Chang WJ (2016). Chloroplast genomes: diversity, evolution, and applications in genetic engineering. Genome Biol 17, 134. |
| [9] | Dierckxsens N, Mardulyn P, Smits G (2017). NOVOPlasty: de novo assembly of organelle genomes from whole genome data. Nucleic Acids Res 45, e18. |
| [10] | Domarew CA, Holt RR, Goodman-Snitkoff G (2002). A study of Russian phytomedicine and commonly used herbal remedies. J Herb Pharmacother 2, 31-48. |
| [11] | Frazer KA, Pachter L, Poliakov A, Rubin EM, Dubchak I (2004). VISTA: computational tools for comparative genomics. Nucleic Acids Res 32, W273-W279. |
| [12] | Fu PC, Zhang YZ, Geng HM, Chen SL (2016). The complete chloroplast genome sequence of Gentiana lawrencei var. farreri (Gentianaceae) and comparative analysis with its congeneric species. PeerJ 4, e2540. |
| [13] | Guo HJ, Liu JS, Luo L, Wei XP, Zhang J, Qi YD, Zhang BG, Liu HT, Xiao PG (2017). Complete chloroplast genome sequences of Schisandra chinensis: genome structure, comparative analysis, and phylogenetic relationship of basal angiosperms. Sci China Life Sci 60, 1286-1290. |
| [14] | Guo W, Yu YJ, Shen RJ, Liao WB, Chin SW, Potter D (2011). A phylogeny of Photinia sensu lato (Rosaceae) and related genera based on nrITS and cpDNA analysis. Plant Syst Evol 291, 91-102. |
| [15] | Hardin JW (1973). The enigmatic chokeberries (Aronia, Rosaceae). Bull Torrey Bot Club 100, 178-184. |
| [16] | He Y, Xiao HT, Deng C, Xiong L, Yang J, Peng C (2016). The complete chloroplast genome sequences of the medicinal plant Pogostemon cablin. Int J Mol Sci 17, 820. |
| [17] | Hildebrand M, Hallick RB, Passavant CW, Bourque DP (1988). Trans-splicing in chloroplasts: the rps 12 loci of Nicotiana tabacum. Proc Natl Acad Sci USA 85, 372-376. |
| [18] | Katoh K, Rozewicki J, Yamada KD (2019). MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20, 1160-1166. |
| [19] | Kulling SE, Rawel HM (2008). Chokeberry (Aronia melanocarpa)—a review on the characteristic components and potential health effects. Planta Med 74, 1625-1634. |
| [20] | Kurtz S, Choudhuri JV, Ohlebusch E, Schleiermacher C, Stoye J, Giegerich R (2001). REPuter: the manifold applications of repeat analysis on a genomic scale. Nucleic Acids Res 29, 4633-4642. |
| [21] | Li QY, Guo W, Liao WB, Macklin JA, Li JH (2012). Generic limits of Pyrinae: insights from nuclear ribosomal DNA sequences. Bot Stud 53, 151-164. |
| [22] | Li YF, Sylvester SP, Li M, Zhang C, Li X, Duan YF, Wang XR (2019). The complete plastid genome of Magnolia zenii and genetic comparison to Magnoliaceae species. Molecules 24, 261. |
| [23] | Lim JD, Cha HS, Choung MG, Choi RN, Choi DJ, Youn AR (2014). Antioxidant activities of acidic ethanol extract and the anthocyanin rich fraction from Aronia melanocarpa. Korean J Food Cookery Sci 30, 573-578. |
| [24] | Liu BB, Liu GN, Hong DY, Wen J (2020). Eriobotrya belongs to Rhaphiolepis (Maleae, Rosaceae): evidence from chloroplast genome and nuclear ribosomal DNA data. Front Plant Sci 10, 1731. |
| [25] | Liu HY, Yu Y, Deng YQ, Li J, Huang ZX, Zhou SD (2018). The chloroplast genome of Lilium henrici: genome structure and comparative analysis. Molecules 23, 1276. |
| [26] | Liu QP, Xue QZ (2005). Comparative studies on codon usage pattern of chloroplasts and their host nuclear genes in four plant species. J Genet 84, 55-62. |
| [27] | Liu YC, Lin BY, Lin JY, Wu WL, Chang CC (2016). Evaluation of chloroplast DNA markers for intraspecific identification of Phalaenopsis equestris cultivars. Sci Hortic 203, 86-94. |
| [28] | Lohse M, Drechsel O, Bock R (2007). OrganellarGenomeDRAW (OGDRAW): a tool for the easy generation of high-quality custom graphical maps of plastid and mitochondrial genomes. Curr Genet 52, 267-274. |
| [29] | Mayor C, Brudno M, Schwartz JR, Poliakov A, Rubin EM, Frazer KA, Pachter LS, Dubchak I (2000). VISTA: visualizing global DNA sequence alignments of arbitrary length. Bioinformatics 16, 1046-1047. |
| [30] | Morton BR (2003). The role of context-dependent mutations in generating compositional and codon usage bias in grass chloroplast DNA. J Mol Evol 56, 616-629. |
| [31] | Mudunuri SB, Nagarajaram HA (2007). IMEx: imperfect microsatellite extractor. Bioinformatics 23, 1181-1187. |
| [32] | Palmer JD (1987). Chloroplast DNA evolution and biosystematic uses of chloroplast DNA variation. Am Nat 130, S6-S29. |
| [33] | Price MN, Dehal PS, Arkin AP (2010). FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490. |
| [34] | Ravi V, Khurana JP, Tyagi AK, Khurana P (2008). An update on chloroplast genomes. Plant Syst Evol 271, 101-122. |
| [35] | Robertson KR, Phipps JB, Rohrer JR, Smith PG (1991). A synopsis of genera in Maloideae (Rosaceae). Syst Bot 16, 376-394. |
| [36] | Seidemann J (1993). Chockberries a fruit little known till now. Deutsch Lebensm Rundsch (Ger) 89, 149-151. |
| [37] | Sharp PM, Li WH (1987). The codon adaptation index—a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15, 1281-1295. |
| [38] | Shi LC, Chen HM, Jiang M, Wang LQ, Wu X, Huang LF, Liu C (2019). CPGAVAS2, an integrated plastome sequence annotator and analyzer. Nucleic Acids Res 47, W65-W73. |
| [39] | Shipunov A, Gladkova S, Timoshina P, Lee HJ, Choi J, Despiegelaere S, Connolly B (2019). Mysterious chokeberries: new data on the diversity and phylogeny of Aronia Medik. (Rosaceae). Eur J Taxon 570, 1-14. |
| [40] | Shukla S, Mehta A (2015). Anticancer potential of medicinal plants and their phytochemicals: a review. Braz J Bot 38, 199-210. |
| [41] | Szopa A, Kokotkiewicz A, Kubica P, Banaszczak P, Wojtanowska-Kro?niak A, Kro?niak M, Marzec-Wróblewska U, Badura A, Zagrodzki P, Bucinski A, Luczkiewicz M, Ekiert H (2017). Comparative analysis of different groups of phenolic compounds in fruit and leaf extracts of Aronia sp.: A. melanocarpa, A. arbutifolia, and A. × prunifolia and their antioxidant activities. Eur Food Res Technol 243, 1645-1657. |
| [42] | Wang T, Zeng XY, Hu YQ, Li N (2023). Optimization of brewing technology and antioxidant activity of wild cherry berry wine. Food Ind 44(6), 26-30. (in Chinese) |
| 汪娣, 曾雪莹, 胡玉清, 李楠 (2023). 野樱莓果酒酿造工艺优化及抗氧化活性分析. 食品工业 44(6), 26-30. | |
| [43] | Wyman SK, Jansen RK, Boore JL (2004). Automatic annotation of organellar genomes with DOGMA. Bioinformatics 20, 3252-3255. |
| [44] | Xu C, Dong WP, Li WQ, Lu YZ, Xie XM, Jin XB, Shi JP, He KH, Suo ZL (2017). Comparative analysis of six Lagerstroemia complete chloroplast genomes. Front Plant Sci 8, 15. |
| [45] | Yang L, Ding X, Jiang ZP, Chen WW, Chen S, Ma N, Xu SY, Wang JY (2024). Optimization of fermentation technology of Aronia melanocarpa pomace enzyme and study on its antioxidant activity. Chin Cond 49(9), 29-37. (in Chinese) |
| 杨柳, 丁雪, 姜志鹏, 陈文文, 陈生, 马宁, 徐圣喻, 王静怡 (2024). 野樱莓果渣酵素发酵工艺优化及抗氧化活性研究. 中国调味品 49(9), 29-37. | |
| [46] | Zhao F, Drew BT, Chen YP, Hu GX, Li B, Xiang CL (2020). The chloroplast genome of Salvia: genomic characterization and phylogenetic analysis. Int J Mol Sci 181, 812-830. |
| [47] | Zhao JL, Qi B, Ding LJ, Tang XQ (2010). Based on RSCU and QRSCU research codon bias of F/10 and G/11 Xylanase. J Food Sci Biotechnol 29, 755-764. |
| [48] | Zhao YM, Yang ZY, Zhao YP, Li XL, Zhao ZX, Zhao GF (2019). Chloroplast genome structural characteristics and phylogenetic relationships of Oleaceae. Chin Bull Bot 54, 441-454. (in Chinese) |
| 赵月梅, 杨振艳, 赵永平, 李筱玲, 赵志新, 赵桂仿 (2019). 木犀科植物叶绿体基因组结构特征和系统发育关系. 植物学报 54, 441-454. |
/
| 〈 |
|
〉 |