

Chinese Bulletin of Botany ›› 2015, Vol. 50 ›› Issue (5): 539-548.DOI: 10.11983/CBB15076 cstr: 32102.14.CBB15076
Previous Articles Next Articles
Shujia Li, Jin Gao, Jiayang Li, Yonghong Wang*
Received:2015-05-18
Accepted:2015-07-02
Online:2015-09-01
Published:2015-10-09
Contact:
Wang Yonghong
About author:? These authors contributed equally to this paper
Shujia Li, Jin Gao, Jiayang Li, Yonghong Wang. Advances in Regulating Rice Tillers by Strigolactones[J]. Chinese Bulletin of Botany, 2015, 50(5): 539-548.
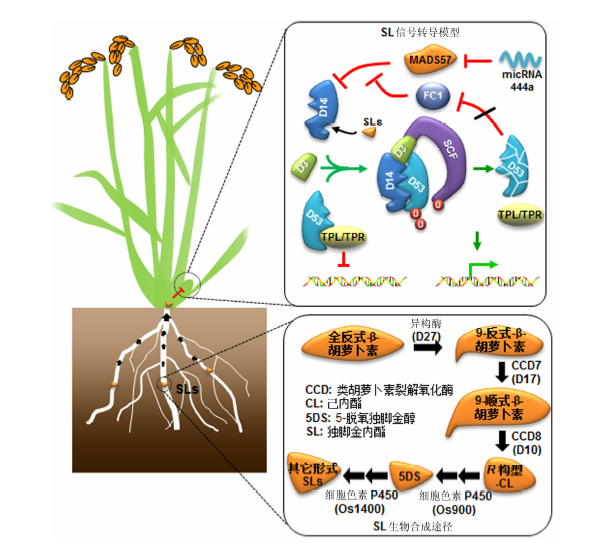
Figure 1 A simplified model of rice tillering inhibition via strigolactonesStrigolactones are produced and exuded mainly from the roots and function as plant hormones inhibiting rice tiller outgrowth. Biosynthesis of strigolactones proceeds through D27, followed by D17, D10, and OsMAX1 (Os900, Os1400). SL signaling is mediated by D3, D14, and D53. The expression of D14 is repressed by OsMADS57, and this is in turn repressed by OsTB1/FC1, which is respressed by D53. CCD: Carotenoid cleavage dioxygenases; CL: Carlactone; 5DS: 5-deoxystrigol; SL: Strigolactone
| 1 | Alder A, Jamil M, Marzorati M, Bruno M, Vermathen M, Bigler P, Ghisla S, Bouwmeester H, Beyer P, Al-Babili S (2012). The path from β-carotene to carlactone, a strigo- lactone-like plant hormone.Science 335, 1348-1351. |
| 2 | Arite T, Iwata H, Ohshima K, Maekawa M, Nakajima M, Kojima M, Sakakibara H, Kyozuka J (2007). DWARF10, an RMS1/MAX4/DAD1 ortholog, controls lateral bud outgrowth in rice.Plant J 51, 1019-1029. |
| 3 | Arite T, Umehara M, Ishikawa S, Hanada A, Maekawa M, Yamaguchi S, Kyozuka J (2009). d14, a strigolactone- insensitive mutant of rice, shows an accelerated outgrowth of tillers.Plant Cell Physiol 50, 1416-1424. |
| 4 | Bennett T, Leyser O (2014). Strigolactone signaling: standing on the shoulders of DWARFs.Curr Opin Plant Biol 22, 7-13. |
| 5 | Beveridge CA (2000). Long-distance signaling and a mutational analysis of branching in pea.Plant Growth Regul 32, 193-203. |
| 6 | Beveridge CA, Kyozuka J (2010). New genes in the strigolactone-related shoot branching pathway.Curr Opin Plant Biol 13, 34-39. |
| 7 | Booker J, Auldridge M, Wills S, McCarty D, Klee H, Leyser O (2004). MAX3/CCD7 is a carotenoid cleavage dioxygenase required for the synthesis of a novel plant signaling molecule.Curr Biol 14, 1232-1238. |
| 8 | Booker J, Sieberer T, Wright W, Williamson L, Willett B, Stirnberg P, Turnbull C, Srinivasan M, Goddard P, Leyser O (2005). MAX1 encodes a cytochrome P450 family member that acts downstream of MAX3/4 to produce a carotenoid-derived branch-inhibiting hormone.Dev Cell 8, 443-449. |
| 9 | Butler LG (1995). Chemical communication between the parasitic weed Striga and its crop host: a new dimension in allelochemistry.Acs Sym Ser 582, 158-168. |
| 10 | Cardoso C, Zhang YX, Jamil M, Hepworth J, Charnikhova T, Dimkpa SON, Meharg C, Wright MH, Liu JW, Meng XB, Wang YH, Li JY, McCouch SR, Leyser O, Price AH, Bouwmeester HJ, Ruyter-Spira C (2014). Natural varia- tion of rice strigolactone biosynthesis is associated with the deletion of two MAX1 orthologs.Proc Natl Acad Sci USA 111, 2379-2384. |
| 11 | Cook CE, Whichard LP, Turner B, Wall ME, Egley GH (1966). Germination of witchweed (Striga lutea Lour.): isolation and properties of a potent stimulant.Science 154, 1189-1190. |
| 12 | Cook CE, Whichard LP, Wall M, Egley GH, Coggon P, Luhan PA, McPhail AT (1972). Germination stimulants. II. Structure of strigol, a potent seed germination stimulant for witchweed (Striga lutea).J Am Chem Soc 94, 6198-6199. |
| 13 | Drummond RSM, Martínez-Sánchez NM, Janssen BJ, Templeton KR, Simons JL, Quinn BD, Karunairetnam S, Snowden KC (2009). Petunia hybrida CAROTENOID CLEAVAGE DIOXYGENASE7 is involved in the production of negative and positive branching signals in Petunia.Plant Physiol 151, 1867-1877. |
| 14 | Dun EA, Brewer PB, Beveridge CA (2009). Strigolactones: discovery of the elusive shoot branching hormone.Trends Plant Sci 14, 364-372. |
| 15 | Foo E, Bullier E, Goussot M, Foucher F, Rameau C, Beveridge CA (2005). The branching gene RAMOSUS1 mediates interactions among two novel signals and auxin in pea.Plant Cell 17, 464-474. |
| 16 | Gao ZY, Qian Q, Liu XH, Yan MX, Feng Q, Dong GJ, Liu J, Han B (2009). Dwarf 88, a novel putative esterase gene affecting architecture of rice plant.Plant Mol Biol 71, 265-276. |
| 17 | Gomez-Roldan V, Fermas S, Brewer PB, Puech-Pagès V, Dun EA, Pillot JP, Letisse F, Matusova R, Danoun S, Portais JC, Bouwmeester H, Bécard G, Beveridge CA, Rameau C, Rochange SF (2008). Strigolactone inhibition of shoot branching.Nature 455, 189-194. |
| 18 | Guo SY, Xu YY, Liu HH, Mao ZW, Zhang C, Ma Y, Zhang QR, Meng Z, Chong K (2013). The interaction between OsMADS57 and OsTB1 modulates rice tillering via DWA- RF14.Nat Commun 4, 1566. |
| 19 | Hamiaux C, Drummond RSM, Janssen BJ, Ledger SE, Cooney JM, Newcomb RD, Snowden KC (2012). DAD2 is an α/β hydrolase likely to be involved in the perception of the plant branching hormone, strigolactone.Curr Biol 22, 2032-2036. |
| 20 | Ishikawa S, Maekawa M, Arite T, Onishi K, Takamure I, Kyozuka J (2005). Suppression of tiller bud activity in tillering dwarf mutants of rice.Plant Cell Physiol 46, 79-86. |
| 21 | Jiang L, Liu X, Xiong GS, Liu HH, Chen FL, Wang L, Meng XB, Liu GF, Yu H, Yuan YD, Yi W, Zhao LH, Ma HL, He YZ, Wu ZS, Melcher K, Qian Q, Xu HE, Wang YH, Li JY (2013). DWARF 53 acts as a repressor of strigolactone signaling in rice.Nature 504, 401-405. |
| 22 | Johnson X, Brcich T, Dun EA, Goussot M, Haurogné K, Beveridge CA, Rameau C (2006). Branching genes are conserved across species. Genes controlling a novel signal in pea are coregulated by other long-distance signals.Plant Physiol 142, 1014-1026. |
| 23 | Kagiyama M, Hirano Y, Mori T, Kim SY, Kyozuka J, Seto Y, Yamaguchi S, Hakoshima T (2013). Structures of D14 and D14L in the strigolactone and karrikin signaling pathways.Genes Cells 18, 147-160. |
| 24 | Kohlen W, Charnikhova T, Liu Q, Bours R, Domagalska MA, Beguerie S, Verstappen F, Leyser O, Bouwmeester H, Ruyter-Spira C (2011). Strigolactones are transported through the xylem and play a key role in shoot architectural response to phosphate deficiency in nonarbuscular mycor- rhizal host Arabidopsis.Plant Physiol 155, 974-987. |
| 25 | Leyser O (2008). Strigolactones and shoot branching: a new trick for a young dog.Dev Cell 15, 337-338. |
| 26 | Li Z, Czarnecki O, Chourey K, Yang J, Tuskan GA, Hurst GB, Pan CL, Chen JG (2014). Strigolactone-regulated proteins revealed by iTRAQ-based quantitative proteomics in Arabidopsis.J Proteome Res 13, 1359-1372. |
| 27 | Lin H, Wang RX, Qian Q, Yan MX, Meng XB, Fu ZM, Yan CY, Jiang B, Su Z, Li JY, Wang YH (2009). DWARF27, an iron-containing protein required for the biosynthesis of strigolactones, regulates rice tiller bud outgrowth.Plant Cell 21, 1512-1525. |
| 28 | Liu WZ, Wu C, Fu YP, Hu GC, Si HM, Zhu L, Luan WJ, He ZQ, Sun ZX (2009). Identification and characterization of HTD2: a novel gene negatively regulating tiller bud out- growth in rice.Planta 230, 649-658. |
| 29 | Mashiguchi K, Sasaki E, Shimada Y, Nagae M, Ueno K, Nakano T, Yoneyama K, Suzuki Y, Asami T (2009). Feedback-regulation of strigolactone biosynthetic genes and strigolactone-regulated genes in Arabidopsis.Biosci Biotechnol Biochem 73, 2460-2465. |
| 30 | Mayzlish-Gati E, De-Cuyper C, Goormachtig S, Beeck- man T, Vuylsteke M, Brewer PB, Beveridge CA, Yermi- yahu U, Kaplan Y, Enzer Y, Wininger S, Resnick N, Cohen M, Kapulnik Y, Koltai H (2012). Strigolactones are involved in root response to low phosphate conditions in Arabidopsis.Plant Physiol 160, 1329-1341. |
| 31 | Minakuchi K, Kameoka H, Yasuno N, Umehara M, Luo L, Kobayashi K, Hanada A, Ueno K, Asami T, Yamaguchi S, Kyozuka J (2010). FINE CULM1 (FC1) works down- stream of strigolactones to inhibit the outgrowth of axil- lary buds in rice.Plant Cell Physiol 51, 1127-1135. |
| 32 | Nakamura H, Xue YL, Miyakawa T, Hou F, Qin HM, Fukui K, Shi X, Ito E, Ito S, Park SH, Miyauchi Y, Asano A, Tot- suka N, Ueda T, Tanokura M, Asami T (2013). Molecular mechanism of strigolactone perception by DWARF14.Nat Commun 4, 2613. |
| 33 | Napoli C (1996). Highly branched phenotype of the petunia dad1-1 mutant is reversed by grafting.Plant Physiol 111, 27-37. |
| 34 | Nelson DC, Scaffidi A, Dun EA, Waters MT, Flematti GR, Dixon KW, Beveridge CA, Ghisalberti EL, Smith SM (2011). F-box protein MAX2 has dual roles in karrikin and strigolactone signaling in Arabidopsis thaliana. Proc Natl Acad Sci USA 108, 8897-8902. |
| 35 | Rani K, Zwanenburg B, Sugimoto Y, Yoneyama K, Bou- wmeester HJ (2008). Biosynthetic considerations could assist the structure elucidation of host plant produced rhizosphere signaling compounds (strigolactones) for arbuscular mycorrhizal fungi and parasitic plants.Plant Physiol Biochem 46, 617-626. |
| 36 | Sang DJ, Chen DQ, Liu GF, Liang Y, Huang LZ, Meng XB, Chu JF, Sun XH, Dong GJ, Yuan YD, Qian Q, Li JY, Wang YH (2014). Strigolactones regulate rice tiller angle by attenuating shoot gravitropism through inhibiting auxin biosynthesis.Proc Natl Acad Sci USA 111, 11199-11204. |
| 37 | Schwartz SH, Qin XQ, Loewen MC (2004). The biochemical characterization of two carotenoid cleavage enzymes from Arabidopsis indicates that a carotenoid-derived compound inhibits lateral branching.J Biol Chem 279, 46940-46945. |
| 38 | Seto Y, Yamaguchi S (2014). Strigolactone biosynthesis and perception.Curr Opin Plant Biol 21, 1-6. |
| 39 | Shen H, Zhu L, Bu QY, Huq E (2012). MAX2 affects multiple hormones to promote photomorphogenesis.Mol Plant 5, 750-762. |
| 40 | Shimizu-Sato S, Mori H (2001). Control of outgrowth and dormancy in axillary buds.Plant Physiol 127, 1405-1413. |
| 41 | Siame BA, Weerasuriya Y, Wood K, Ejeta G, Butler LG (1993). Isolation of strigol, a germination stimulant for Striga asiatica, from host plants.J Agric Food Chem 41, 1486-1491. |
| 42 | Simons JL, Napoli CA, Janssen BJ, Plummer KM, Snowden KC (2007). Analysis of the DECREASED APICAL DOMINANCE genes of petunia in the control of axillary branching.Plant Physiol 143, 697-706. |
| 43 | Smith SM, Li JY (2014). Signaling and responses to strigol- actones and karrikins.Curr Opin Plant Biol 21, 23-29. |
| 44 | Sorefan K, Booker J, Haurogné K, Goussot M, Bainbridge K, Foo E, Chatfield S, Ward S, Beveridge C, Rameau C, Leyser O (2003). MAX4 and RMS1 are orthologous di- oxygenase-like genes that regulate shoot branching in Arabidopsis and pea.Genes Dev 17, 1469-1474. |
| 45 | Stanga JP, Smith SM, Briggs WR, Nelson DC (2013). SUPPRESSOR OF MORE AXILLARY GROWTH2 1 controls seed germination and seedling development in Arabidopsis.Plant Physiol 163, 318-330. |
| 46 | Stirnberg P, Furner IJ, Ottoline-Leyser HM (2007). MAX2 participates in an SCF complex which acts locally at the node to suppress shoot branching.Plant J 50, 80-94. |
| 47 | Stirnberg P, van De Sande K, Leyser O (2002). MAX1 and MAX2 control shoot lateral branching in Arabidopsis.Development 129, 1131-1141. |
| 48 | Stirnberg P, Ward S, Leyser O (2010). Auxin and stri- golactones in shoot branching: intimately connected?Biochem Soc Trans 38, 717-722. |
| 49 | Umehara M, Hanada A, Yoshida S, Akiyama K, Arite T, Takeda-Kamiya N, Magome H, Kamiya Y, Shirasu K, Yoneyama K, Kyozuka J, Yamaguchi S (2008). Inhibition of shoot branching by new terpenoid plant hormones.Nature 455, 195-200. |
| 50 | Waldie T, McCulloch H, Leyser O (2014). Strigolactones and the control of plant development: lessons from shoot branching.Plant J 79, 607-622. |
| 51 | Wang Y, Sun SY, Zhu WJ, Jia KP, Yang HQ, Wang XL (2013). Strigolactone/MAX2-induced degradation of brass- inosteroid transcriptional effector BES1 regulates shoot branching.Dev Cell 27, 681-688. |
| 52 | Waters MT, Brewer PB, Bussell JD, Smith SM, Beveridge CA (2012). The Arabidopsis ortholog of rice DWARF27 acts upstream of MAX1 in the control of plant development by strigolactones.Plant Physiol 159, 1073-1085. |
| 53 | Waters MT, Smith SM, Nelson DC (2011). Smoke signals and seed dormancy: where next for MAX2?Plant Signal Behav 6, 1418-1422. |
| 54 | Wei LR, Xu JC, Li XB, Qian Q, Zhu LH (2006). Genetic analysis and mapping of the dominant dwarfing gene D-53 in rice.J Integr Plant Biol 48, 447-452. |
| 55 | Woo HR, Chung KM, Park JH, Oh SA, Ahn T, Hong SH, Jang SK, Nam HG (2001). ORE9, an F-box protein that regulates leaf senescence in Arabidopsis.Plant Cell 13, 1779-1790. |
| 56 | Yan HF, Saika H, Maekawa M, Takamure I, Tsutsumi N, Kyozuka J, Nakazono M (2007). Rice tillering dwarf mutant dwarf3 has increased leaf longevity during darkness-induced senescence or hydrogen peroxide- induced cell death.Genes Genetic Syst 82, 361-366. |
| 57 | Zhang YX, van Dijk ADJ, Scaffidi A, Flematti GR, Hofmann M, Charnikhova T, Verstappen F, Hepworth J, van der Krol S, Leyser O, Smith SM, Zwanenburg B, Al-Babili S, Ruyter-Spira C, Bouwmeester HJ (2014). Rice cytochrome P450 MAX1 homologs catalyze distinct steps in strigolactone biosynthesis.Nat Chem Biol 10, 1028-1033. |
| 58 | Zhao LH, Zhou XE, Wu ZS, Yi W, Xu Y, Li SL, Xu TH, Liu Y, Chen RZ, Kovach A, Kang YY, Hou L, He YZ, Xie C, Song WL, Zhong DF, Xu YC, Wang YH, Li JY, Zhang CH, Melcher K, Xu HE (2013). Crystal structures of two phytohormone signal-transducing α/β hydrolases: karrikin- signaling KAI2 and strigolactone-signaling DWARF14.Cell Res 23, 436-439. |
| 59 | Zhou F, Lin QB, Zhu LH, Ren YL, Zhou KN, Shabek N, Wu FQ, Mao HB, Dong W, Gan L, Ma WW, Gao H, Chen J, Yang C, Wang D, Tan JJ, Zhang X, Guo XP, Wang JL, Jiang L, Liu X, Chen WQ, Chu JF, Yan CY, Ueno K, Ito S, Asami T, Cheng ZJ, Wang J, Lei CL, Zhai HQ, Wu CY, Wang HY, Zheng N, Wan JM (2013). D14-SCFD3-de- pendent degradation of D53 regulates strigolactone signaling.Nature 504, 406-410. |
| 60 | Zou JH, Zhang SY, Zhang WP, Li G, Chen ZX, Zhai WX, Zhao XF, Pan XB, Xie Q, Zhu LH (2006). The rice HIGH-TILLERING DWARF1 encoding an ortholog of Ara- bidopsis MAX3 is required for negative regulation of the outgrowth of axillary buds.Plant J 48, 687-698. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||