

Received date: 2020-12-11
Accepted date: 2021-02-25
Online published: 2021-02-25
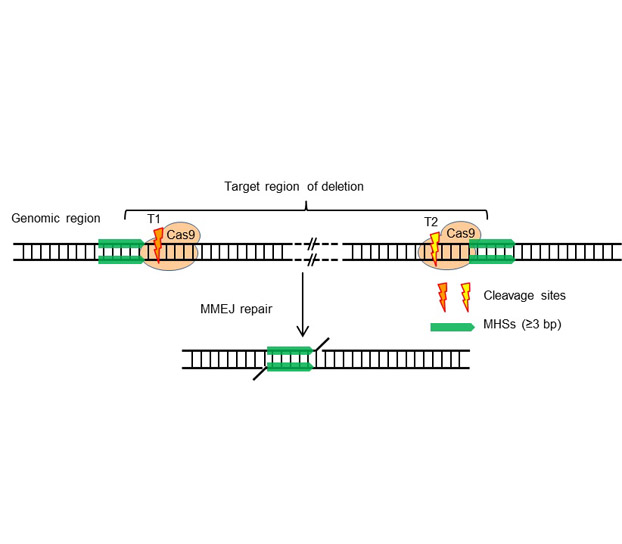
CRISPR/Cas9-based genome editing technology has been an important tool to study the gene function and genomic modification. Directed by a guide RNA, Cas9 protein can cleavage the genomic DNA at the target site, and produce mutations, including deletion, insertion, substitution and fragment deletion, by DNA double strand break (DSB) repair mechanism. In this protocol, we introduce the method to use CRISPR/Cas9 system to increase the efficiency of genomic DNA fragment deletion with microhomology-mediated end joining, especially the details in target design and detection of mutant plants.
Key words: genome editing; CRISPR/Cas9; MMEJ; microhomologous; fragment deletion
Xianrong Xie , Dongchang Zeng , Jiantao Tan , Qinlong Zhu , Yaoguang Liu . CRISPR-based DNA Fragment Deletion in Plants[J]. Chinese Bulletin of Botany, 2021 , 56(1) : 44 -49 . DOI: 10.11983/CBB20203
| [1] | 苏钺凯, 邱镜仁, 张晗, 宋振巧, 王建华 (2019). CRISPR/ Cas9系统在植物基因组编辑中技术改进与创新的研究进展. 植物学报 54, 385-395. |
| [2] | 曾栋昌, 马兴亮, 谢先荣, 祝钦泷, 刘耀光 (2018). 植物CRISPR/Cas9多基因编辑载体构建和突变分析的操作方法. 中国科学: 生命科学 48, 783-794. |
| [3] | Burgess DG, Xu J, Freeling M (2015). Advances in understanding cis regulation of the plant gene with an emphasis on comparative genomics. Curr Opin Plant Biol 27, 141-147. |
| [4] | Canver MC, Bauer DE, Dass A, Yien YY, Chung J, Masuda T, Maeda T, Paw BH, Orkin SH (2014). Characterization of genomic deletion efficiency mediated by clustered regularly interspaced palindromic repeats (CRIS- PR)/Cas9 nuclease system in mammalian cells. J Biol Chem 289, 21312-21324. |
| [5] | Liu WZ, Xie XR, Ma XL, Li J, Chen JH, Liu YG (2015). DSDecode: a web-based tool for decoding of sequencing chromatograms for genotyping of targeted mutations. Mol Plant 8, 1431-1433. |
| [6] | Ma XL, Zhang QY, Zhu QL, Liu W, Chen Y, Qiu R, Wang B, Yang ZF, Li HY, Lin YR, Xie YY, Shen RX, Chen SF, Wang Z, Chen YL, Guo JX, Chen LT, Zhao XC, Dong ZC, Liu YG (2015). A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol Plant 8, 1274-1284. |
| [7] | Ma XL, Zhu QL, Chen YL, Liu YG (2016). CRISPR/Cas9 platforms for genome editing in plants: developments and applications. Mol Plant 9, 961-974. |
| [8] | Manova V, Gruszka D (2015). DNA damage and repair in plants-from models to crops. Front Plant Sci 6, 885. |
| [9] | Owens DDG, Caulder A, Frontera V, Harman JR, Allan AJ, Bucakci A, Greder L, Codner GF, Hublitz P, McHugh PJ, Teboul L, de Bruijn MFTR (2019). Microhomologies are prevalent at Cas9-induced larger deletions. Nucleic Acids Res 47, 7402-7417. |
| [10] | Tan JT, Zhao YC, Wang B, Hao Y, Wang YX, Li YY, Luo WN, Zong WB, Li GS, Chen SF, Ma K, Xie XR, Chen LT, Liu YG, Zhu QL (2020). Efficient CRISPR/Cas9-based plant genomic fragment deletions by microhomology- mediated end joining. Plant Biotechnol J 18, 2161-2163. |
| [11] | Tuladhar R, Yeu Y, Piazza JT, Tan ZP, Clemenceau JR, Wu XF, Barrett Q, Herbert J, Mathews DH, Kim J, Hwang TH, Lum L (2019). CRISPR-Cas9-based mu- tagenesis frequently provokes on-target mRNA misregulation. Nat Commun 10, 4056. |
| [12] | Xie XR, Liu WZ, Dong G, Zhu QH, Liu YG (2020). MMEJ-KO: a web tool for designing paired CRISPR guide RNAs for microhomology-mediated end joining fragment deletion. Sci China Life Sci doi: 10.1007/s11427-020-1797-3. |
| [13] | Xie XR, Ma XL, Zhu QL, Zeng DC, Li GS, Liu YG (2017). CRISPR-GE: a convenient software toolkit for CRISPR- based genome editing. Mol Plant 10, 1246-1249. |
| [14] | Zhang Y, Massel K, Godwin ID, Gao CX (2018). Applications and potential of genome editing in crop improvement. Genome Biol 19, 210. |
/
| 〈 |
|
〉 |