

SlyFRG5 methylation level activated by Fol-milR1 mediates immune response against tomato wilt disease
School of Life Sciences, Zhejiang Normal University, Jinhua 321004
Received date: 2025-07-21
Revised date: 2025-08-19
Online published: 2025-09-05
INTRODUCTION: Tomato wilt disease, caused by Fusarium oxysporum f. sp. lycopersici (Fol), is a soil borne fungal disease that causes significant economic losses to global tomato agricultural production. The molecular mechanism of pathogen sRNA cross-border regulation of host DNA methylation mediated tomato resistance to wilt disease is still unclear. Here, we illustrate that SlyFRG5 methylation level activated by Fol-milR1 mediates immune response against tomato wilt disease.
RATIONALE: The latest research shows that sRNA derived from pathogens can serve as novel effectors for cross-border participation in host-pathogen interactions. Our previous research documented that Fol-milR1 was transferred horizontally into host cells as an effector. Fol-milR1 repressed the expression of a resistant gene SlyFRG4 in tomato, as well as binded to SlyAGO4a leading to interfere with the host immune system to achieve successful infection. Therefore, this research further determines whether transmission of?Fol-milR1 into tomato causes DNA methylation in the tomato genome and elucidate other aspects of the molecular mechanism involving the action of Fol-milR1.
RESULTS: The results indicate that SlyAGO4a negatively participates in the resistance to tomato wilt disease. The methylation level of Solyc02g081370 (SlyFRG5) was identified to be directly associated with Fol-milR1 using whole genome methylation sequencing and molecular biology methods. SlyFRG5 loss-of-function alleles created using CRISPR/Cas9 in resistant cultivar tomato Motelle exhibited enhanced disease susceptibility to Fol, further supporting the idea that SlyFRG4 is essential for tomato wilt disease resistance. SlyFRG5 contributed wilt disease resistance by regulating the accumulation of host ROS in tomato.
CONCLUSION: Combining previous research, we concluded that,during the infection Fol, Fol-milR1 was transferred into host cells to hijack SlyAGO4a which leads to methylation a Fusarium disease resistant gene SlyFRG5. This study analyzed how the Fol-milR1-SlyAGO4a-SlyFRG5 functional module mediated tomato resistance to wilt disease, which will provide a new ideas for exploring the cultivation and quality improvement of tomato resistance to Fusarium disease.
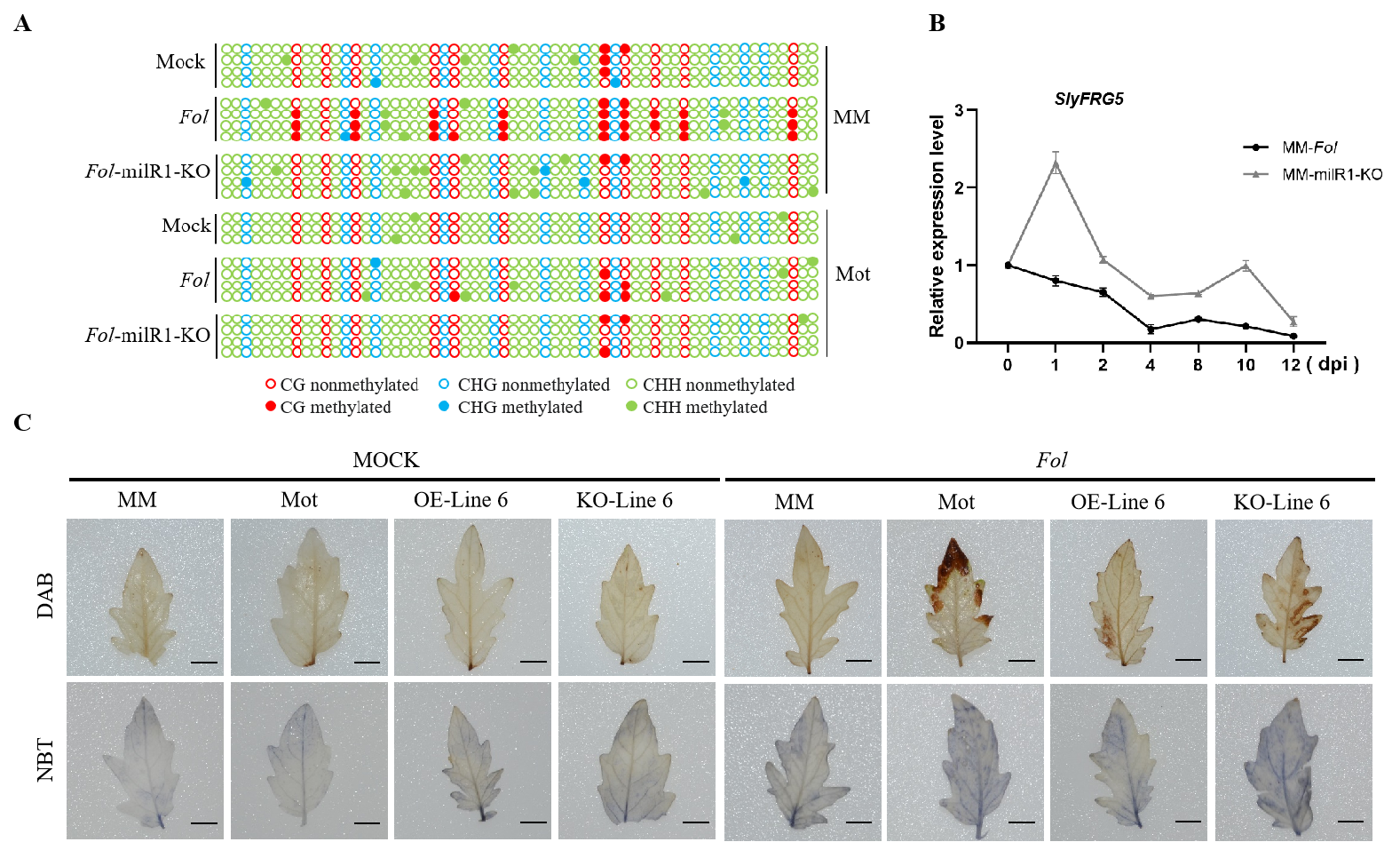
The Fol-milR1-SlyAGO4a-SlyFRG5 functional module mediated tomato resistance to wilt disease. (A) Distribution of methylation type in the CDS (Partial region) region of SlyFRG5; (B) The transcription level of SlyFRG5 at different time points after pathogen infection in MM; (C) Leaf DAB (H2O2) and NBT (•O₂⁻) staining (bar=1 cm).
YU Ru , LI Pan-Ting , XIAO Ying , YE Hong-Xia , CHEN Shu-Man , OU Yang-ShouJiang . SlyFRG5 methylation level activated by Fol-milR1 mediates immune response against tomato wilt disease[J]. Chinese Bulletin of Botany, 0 : 1 -0 . DOI: 10.11983/CBB25129
/
| 〈 |
|
〉 |