

Chinese Bulletin of Botany ›› 2017, Vol. 52 ›› Issue (1): 30-42.DOI: 10.11983/CBB16107 cstr: 32102.14.CBB16107
Special Issue: 水稻生物学专辑 (2017年52卷1期)
Previous Articles Next Articles
Xiaohui Zhang, Xiaomin Feng, Shaoyang Lin*
Received:2016-05-11
Accepted:2016-06-15
Online:2017-01-01
Published:2017-01-23
Contact:
Lin Shaoyang
About author:# Co-first authors
Xiaohui Zhang, Xiaomin Feng, Shaoyang Lin. Scanning for Pi Loci and Rebuilding an Improved Genome of Elite Rice Variety Kongyu 131[J]. Chinese Bulletin of Botany, 2017, 52(1): 30-42.
| Year | Artifical inoculation | Natural infection | |||||
|---|---|---|---|---|---|---|---|
| Seedling blast | Leaf blast | Panicle blast | Seedling blast | Leaf blast | Panicle blast | ||
| 1995 | 8 | 8 | 9 | 6 | 5 | 9 | |
| 1996 | 9 | 9 | 9 | 9 | 9 | 9 | |
| 1998 | unknown | 9 | 9 | unknown | 9 | 9 | |
| 1999 | unknown | 8 | 9 | unknown | 8 | 9 | |
| 2002 | unknown | 9 | 9 | unknown | 8 | 9 | |
| 2003 | unknown | 8 | 9 | unknown | 7 | 9 | |
| 2004 | unknown | 7 | 9 | unknown | 7 | 7 | |
| 2005 | 9 | 9 | 9 | 9 | 9 | 9 | |
Table 1 The results of rice cultivar Kongyu 131’s susceptibility to rice blast with score 1-9 (Scale by IRRI Standard Evaluation System for rice)
| Year | Artifical inoculation | Natural infection | |||||
|---|---|---|---|---|---|---|---|
| Seedling blast | Leaf blast | Panicle blast | Seedling blast | Leaf blast | Panicle blast | ||
| 1995 | 8 | 8 | 9 | 6 | 5 | 9 | |
| 1996 | 9 | 9 | 9 | 9 | 9 | 9 | |
| 1998 | unknown | 9 | 9 | unknown | 9 | 9 | |
| 1999 | unknown | 8 | 9 | unknown | 8 | 9 | |
| 2002 | unknown | 9 | 9 | unknown | 8 | 9 | |
| 2003 | unknown | 8 | 9 | unknown | 7 | 9 | |
| 2004 | unknown | 7 | 9 | unknown | 7 | 7 | |
| 2005 | 9 | 9 | 9 | 9 | 9 | 9 | |
| Markers | Chr. | Position (IRGSP1.0) | Forward primer (5′-3′) | Reverse primer (5′-3′) | |
|---|---|---|---|---|---|
| SNP1 | 11 | 24107429 | CTTACAACCGAATTGCTTATCAC | TATAGGATAAGAGTCGAAAGCATC | |
| SNP2 | 11 | 24552815 | CCTCAACAGGATCCAGATTCAATAC | CAAGATCAATTGAGTTAGACAAC | |
| SNP3 | 11 | 24717067 | CCAGACATAATGCTTAAAGTAG | CATAGTCGCAGTTCATCTTATG | |
| SNP4 | 11 | 24747927 | GACATATTGTATGCAATCTACTCG | TTCACAGAAGCTAGTATAAGAG | |
| SNP5 | 11 | 25104609 | AAGTAATATACGCTACTATCTTG | CGACGTGATGGGTAAGAACAG | |
Table 2 Molecular markers SNP1-SNP5
| Markers | Chr. | Position (IRGSP1.0) | Forward primer (5′-3′) | Reverse primer (5′-3′) | |
|---|---|---|---|---|---|
| SNP1 | 11 | 24107429 | CTTACAACCGAATTGCTTATCAC | TATAGGATAAGAGTCGAAAGCATC | |
| SNP2 | 11 | 24552815 | CCTCAACAGGATCCAGATTCAATAC | CAAGATCAATTGAGTTAGACAAC | |
| SNP3 | 11 | 24717067 | CCAGACATAATGCTTAAAGTAG | CATAGTCGCAGTTCATCTTATG | |
| SNP4 | 11 | 24747927 | GACATATTGTATGCAATCTACTCG | TTCACAGAAGCTAGTATAAGAG | |
| SNP5 | 11 | 25104609 | AAGTAATATACGCTACTATCTTG | CGACGTGATGGGTAAGAACAG | |
| Pi gene | Chr. | GenBank ID | BLAST results (with CDS) | Be contained or lacked in Kongyu 131 |
|---|---|---|---|---|
| Pib | 2 | AB013448.1 | High sequence homology | Uncertain |
| Pita/Pi4a | 12 | AF207842.1 | 3 SNP sites | Uncertain |
| Pi2/Piz5 | 6 | DQ352453.1 | 53 SNP sites | Lacked |
| Pi9 | 6 | DQ285630 | Sequence deletion | Lacked |
| Pizt | 6 | DQ352040 | High sequence homology | Uncertain |
| Pi36 | 8 | DQ900896.1 | Multi-SNP sites | Lacked |
| Pi37 | 1 | DQ923494.1 | 4 SNP sites | Uncertain |
| Pi5-1/Pi3/Pii | 9 | EU869185.1 | 29 SNP sites | Lacked |
| Pit | 1 | AB379815.1 | 6 SNP sites | Uncertain |
| Pish | 1 | Os01g0782100 | No difference | Contained |
| Pb1 | 11 | AB570371.1 | Structural variation | Lacked |
| Pia | 11 | AB604622 and AB604627 | High sequence homology | Uncertain |
| pi-21 | 4 | AB430853.1 | 1 SNP site | Uncertain |
| Pi-d2 | 6 | FJ915121.1 | 6 SNP sites | Uncertain |
| Pi-d3 | 6 | FJ773285 | Terminator codon mutation | Lacked |
| Pi25 | 6 | HM448480 | 18 SNP sites | Lacked |
| Pik | 11 | HM048900.1 | 2 SNP sites | Uncertain |
| Pikh/Pi-54 | 11 | AY914077(Pikh)/HE589445 (Pi54) | Structural variation | Lacked |
| Pi1 | 11 | HQ606329cds | 11 SNP sites | Lacked |
| Pik-m | 11 | AB510262.1 | Structural variation | Lacked |
| Pik-p | 11 | HM035360 | Multi-SNP sites | Lacked |
| Pi56t | 9 | None | Multi-SNP sites | Lacked |
Table 3 The results of the whole genome scanning of rice cultivar Kongyu 131 for Pi genes
| Pi gene | Chr. | GenBank ID | BLAST results (with CDS) | Be contained or lacked in Kongyu 131 |
|---|---|---|---|---|
| Pib | 2 | AB013448.1 | High sequence homology | Uncertain |
| Pita/Pi4a | 12 | AF207842.1 | 3 SNP sites | Uncertain |
| Pi2/Piz5 | 6 | DQ352453.1 | 53 SNP sites | Lacked |
| Pi9 | 6 | DQ285630 | Sequence deletion | Lacked |
| Pizt | 6 | DQ352040 | High sequence homology | Uncertain |
| Pi36 | 8 | DQ900896.1 | Multi-SNP sites | Lacked |
| Pi37 | 1 | DQ923494.1 | 4 SNP sites | Uncertain |
| Pi5-1/Pi3/Pii | 9 | EU869185.1 | 29 SNP sites | Lacked |
| Pit | 1 | AB379815.1 | 6 SNP sites | Uncertain |
| Pish | 1 | Os01g0782100 | No difference | Contained |
| Pb1 | 11 | AB570371.1 | Structural variation | Lacked |
| Pia | 11 | AB604622 and AB604627 | High sequence homology | Uncertain |
| pi-21 | 4 | AB430853.1 | 1 SNP site | Uncertain |
| Pi-d2 | 6 | FJ915121.1 | 6 SNP sites | Uncertain |
| Pi-d3 | 6 | FJ773285 | Terminator codon mutation | Lacked |
| Pi25 | 6 | HM448480 | 18 SNP sites | Lacked |
| Pik | 11 | HM048900.1 | 2 SNP sites | Uncertain |
| Pikh/Pi-54 | 11 | AY914077(Pikh)/HE589445 (Pi54) | Structural variation | Lacked |
| Pi1 | 11 | HQ606329cds | 11 SNP sites | Lacked |
| Pik-m | 11 | AB510262.1 | Structural variation | Lacked |
| Pik-p | 11 | HM035360 | Multi-SNP sites | Lacked |
| Pi56t | 9 | None | Multi-SNP sites | Lacked |
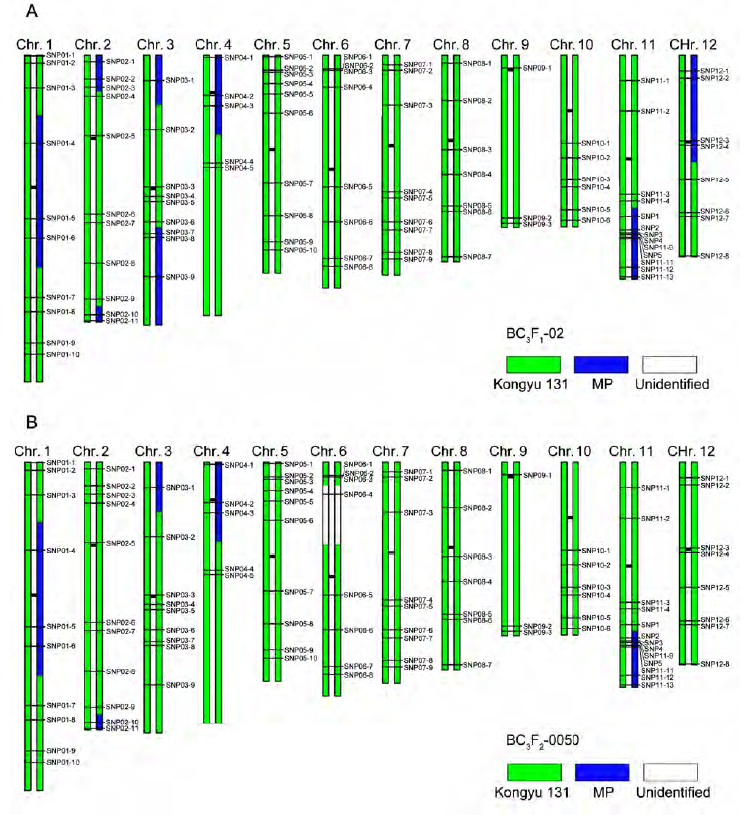
Figure 3 Graphical genotype of the selected individuals of rice^(A) Graphical genotype of BC3F1-02; (B) Graphical genotype of BC3F2-0050. stands for the fragment derived from Kongyu 131, for the fragment derived from MP and for the unidentified loci.
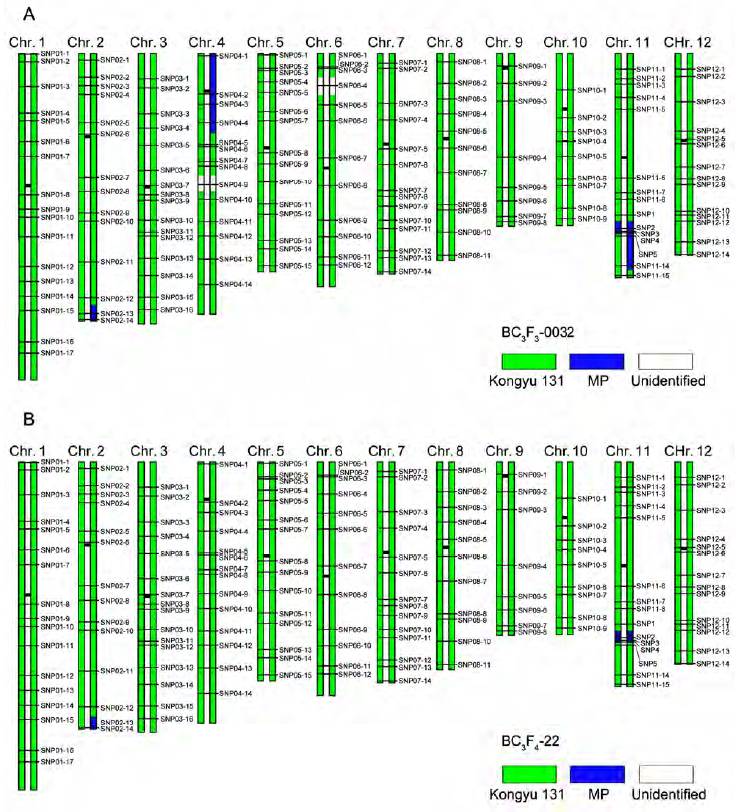
Figure 4 Graphical genotype of the selected individuals of rice^(A) Graphical genotype of BC3F3-0032; (B) Graphical genotype of BC3F4-22. stands for the fragment derived from Kongyu 131, for the fragment derived from MP and for the unidentified loci.
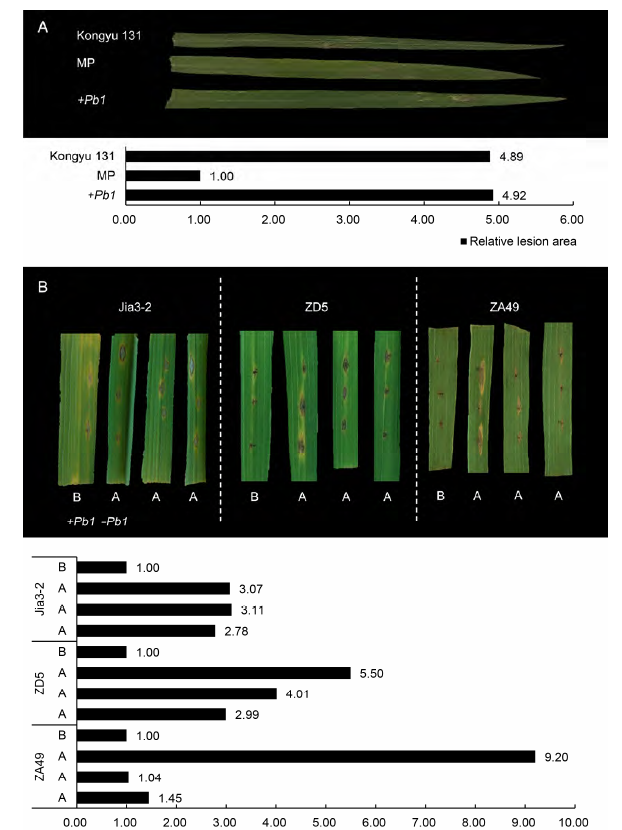
Figure 5 The results of inoculating the improved line of rice^(A) The phenotypic results after inoculation on rice seedlings (5 leaves) and the bar chart of relative lesion area; (B) The phenotypic results after inoculation on ±Pb1 near-isogenic lines in rice adult stage (11 leaves) and the bar chart of relative lesion area.
| [1] | 董志峰, 马荣才, 彭于发, 管华诗 (2001). 转基因植物中外源非目的基因片段的生物安全研究进展. 植物学报 43, 661-672. |
| [2] | 姜玉英, 曾娟, 陆明红, 刘杰 (2013). 2013年全国主要粮食作物重大病虫害发生趋势预报. 植物保护 39, 1-4. |
| [3] | 姜玉英, 曾娟, 陆明红, 刘杰 (2014). 2014年全国主要粮食作物重大病虫害发生趋势预报. 植物保护 40, 1-4. |
| [4] | 姜玉英, 曾娟, 陆明红, 刘杰 (2015). 2015年全国三大谷类作物重大病虫害发生趋势预报. 植物保护 41, 1-4. |
| [5] | 宋成艳, 王桂玲, 辛爱华, 丛万彪 (2007). 黑龙江省水稻品种空育131稻瘟病菌生理小种种类及发病原因分析. 黑龙江农业科学 (1), 41-42. |
| [6] | 孙向东 (2005). 黑龙江省粮食主产区主要作物品种种植情况分析. 黑龙江农业科学 (2), 12-14. |
| [7] | Bryan GT, Wu KS, Farrall L, Jia Y, Hershey HP, Mc- Adams SA, Faulk KN, Donaldson GK, Tarchini R, Valent B (2000). A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta.Plant Cell 12, 2033-2045. |
| [8] | Chen J, Shi Y, Liu W, Chai R, Fu Y, Zhuang J, Wu J (2011). A Pid3 allele from rice cultivar Gumei2 confers resistance to Magnaporthe oryzae.J Genetics Genomics 38, 209-216. |
| [9] | Chen X, Shang J, Chen D, Lei C, Zou Y, Zhai W, Liu G, Xu J, Ling Z, Cao G, Ma B, Wang Y, Zhao X, Li S, Zhu L (2006). A B-lectin receptor kinase gene conferring rice blast resistance.Plant J 46, 794-804. |
| [10] | Fukuoka S, Okuno K (2001). QTL analysis and mapping of pi21, a recessive gene for field resistance to rice blast in Japanese upland rice.Theor Appl Genet 103, 185-190. |
| [11] | Hayashi N, Inoue H, Kato T, Funao T, Shirota M, Shimizu T, Kanamori H, Yamane H, Hayano-Saito Y, Matsumoto T, Yano M, Takatsuji H (2010). Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS- LRR protein and was generated by acquiring a promoter through local genome duplication.Plant J 64, 498-510. |
| [12] | Hua L, Wu J, Chen C, Wu W, He X, Lin F, Wang L, Ashikawa I, Matsumoto T, Wang L, Pan Q (2012). The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast.Theor Appl Genet 125, 1047-1055. |
| [13] | Inoue H, Hayashi N, Matsushita A, Liu X, Nakayama A, Sugano S, Jiang CJ, Takatsuji H (2013). Blast resistance of CC-NB-LRR protein Pb1 is mediated by WRKY- 45 through protein-protein interaction.Proc Natl Acad Sci USA 110, 9577-9582. |
| [14] | Lin F, Chen S, Que Z, Wang L, Liu X, Pan Q (2007). The blast resistance gene Pi37 encodes a nucleotide binding site-leucine-rich repeat protein and is a member of a resistance gene cluster on rice chromosome 1.Genetics 177, 1871-1880. |
| [15] | Okuyama Y, Kanzaki H, Abe A, Yoshida K, Tamiru M, Saitoh H, Fujibe T, Matsumura H, Shenton M, Galam DC, Undan J, Ito A, Sone T, Terauchi R (2011). A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS-LRR protein genes.Plant J 66, 467-479. |
| [16] | Peleman JD, van der Voort JR (2003). Breeding by design.Trends Plant Sci 8, 330-334. |
| [17] | Qu S, Liu G, Zhou B, Bellizzi M, Zeng L, Dai L, Han B, Wang GL (2006). The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics 172, 1901-1914. |
| [18] | Shang J, Tao Y, Chen X, Zou Y, Lei C, Wang J, Li X, Zhao X, Zhang M, Lu Z, Xu J, Cheng Z, Wan J, Zhu L (2009). Identification of a new rice blast resistance gene, Pid3, by genomewide comparison of paired nucleotide-binding site-leucine-rich repeat genes and their pseudogene alleles between the two sequenced rice genomes.Genetics 182, 1303-1311. |
| [19] | Takahashi A, Hayashi N, Miyao A, Hirochika H (2010). Unique features of the rice blast resistance Pish locus revealed by large scale retrotransposon-tagging. BMC Plant Biol 10,175. |
| [20] | Tanaka T, Antonio BA, Kikuchi S, Matsumoto T, Nagamura Y, Numa H, Sakai H, Wu J, Itoh T, Sasaki T, Aono R, Fujii Y, Habara T, Harada E, Kanno M, Kawahara Y, Kawashima H, Kubooka H, Matsuya A, Nakaoka H, Saichi N, Sanbonmatsu R, Sato Y, Shinso Y, Suzuki M, Takeda J, Tanino M, Todokoro F, Yamaguchi K, Yamamoto N, Yamasaki C, Imanishi T, Okido T, Tada M, Ikeo K, Tateno Y, Gojobori T, Lin YC, Wei FJ, Hsing Y, Zhao Q, Han B, Kramer MR, McCombie RW, Lonsdale D, O'Donovan CC, Whitfield EJ, Apweiler R, Koyanagi KO, Khurana JP, Raghuvanshi S, Singh NK, Tyagi AK, Haberer G, Fujisawa M, Hosokawa S, Ito Y, Ikawa H, Shibata M, Yamamoto M, Bruskiewich RM, Hoen DR, Bureau TE, Namiki N, Ohyanagi H, Sakai Y, Nobushima S, Sakata K, Barrero RA, Sato Y, Souvorov A, Smith-White B, Tatusova T, An S, An G, OOta S, Fuks G, Messing J, Christie KR, Lieberherr D, Kim H, Zuccolo A, Wing RA, Nobuta K, Green PJ, Lu C, Meyers BC, Chaparro C, Piegu B, Panaud O, Echeverria M (2008). The rice annotation project database (RAP-DB): 2008 update.Nucleic Acids Res 36, 1028-1033. |
| [21] | von Bubnoff A (2008). Next-generation sequencing: the race is on.Cell 132, 721-723. |
| [22] | Wang ZX, Yano M, Yamanouchi U, Iwamoto M, Monna L, Hayasaka H, Katayose Y, Sasaki T (1999). The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes.Plant J 19, 55-64. |
| [23] | Zhai C, Lin F, Dong Z, He X, Yuan B, Zeng X, Wang L, Pan Q (2011). The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication.New Phytol 189, 321-334. |
| [24] | Zhou B, Qu S, Liu G, Dolan M, Sakai H, Lu G, Bellizzi M, Wang GL (2006). The eight amino-acid differences within three Leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea.Mol Plant-Microbe Interact 19, 1216-1228. |
| No related articles found! |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||