

Chinese Bulletin of Botany ›› 2018, Vol. 53 ›› Issue (4): 391-440.DOI: 10.11983/CBB18177 cstr: 32102.14.CBB18177
• REVIEW BY EDITOR-IN-CHIEF • Next Articles
Chen Fan1, Qian Qian2, Wang Tai3, Dong Aiwu4, Qi Xiaoquan3, Zuo Jianru1, Yang Shuhua5, Lin Rongcheng3, Xiao Langtao6, Gu Hongya7, Chen Zhiduan3, Jiang Liwen8, Bai Yongfei3, Kong Hongzhi3, Chong Kang3
Online:2018-07-01
Published:2018-09-11
About author:† These authors contributed equally to this paper
Chen Fan, Qian Qian, Wang Tai, Dong Aiwu, Qi Xiaoquan, Zuo Jianru, Yang Shuhua, Lin Rongcheng, Xiao Langtao, Gu Hongya, Chen Zhiduan, Jiang Liwen, Bai Yongfei, Kong Hongzhi, Chong Kang. Research Advances in Plant Science in China in 2017[J]. Chinese Bulletin of Botany, 2018, 53(4): 391-440.
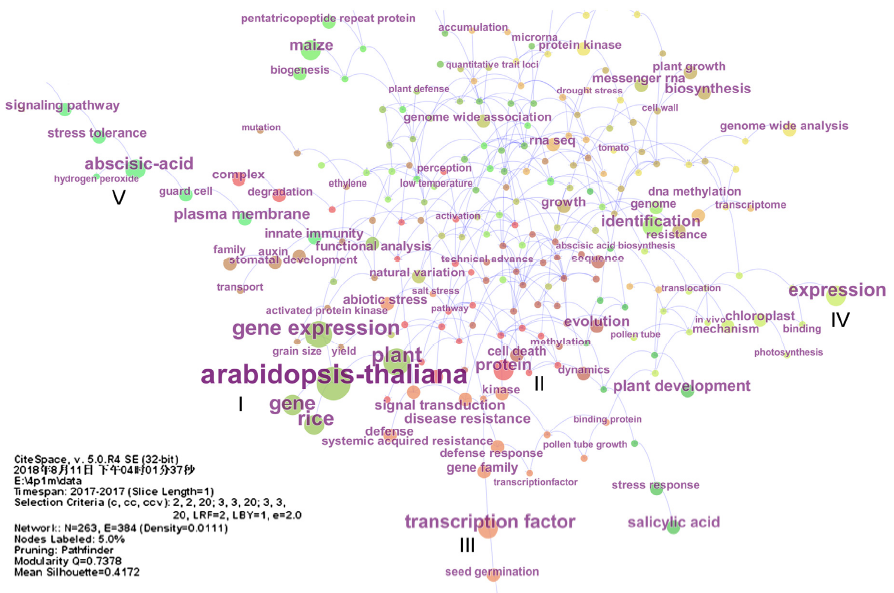
Figure 1 Research themes of Chinese scientists based on four plant science journals in 2017 (data sources: Web of Science)Keyword co-occurence cluster analysis on papers published in four plant science journals (The Plant Cell, The Plant Journal, Plant Physiology and Molecular Plant) of Chinese scientists was carried out by CiteSpace. The results showed that main hot topics studied by Chinese plant scientists included five aspects: (I) Gene and gene expression of model plant Arabidopsis and rice (dark green); (II) Degradation of plant (Arabidopsis and rice) protein complex, and transformation (dark brown); (III) Transcription factor gene family, plant resistance and signal transduction (brown); (IV) Expression of domain containing protein, photosynthesis of chloroplast inner envelope membrane, and translocation mechanism of plant tissue (Arabidopsis and tobacco etc.) (yellow green); (V) Signaling pathway of abscisic acid regulating stress tolerance, innate immunity of guard cell plasma membrane (green).
| The Plant Cell | Plant Physiology | The Plant Journal | Nature Plants | Molecular Plant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | |||||
| 美国 | 99 | 39.3 | 170 | 33.3 | 108 | 29.2 | 51 | 22.8 | 42 | 25.3 | ||||
| 中国 | 63 | 25 | 127 | 24.9 | 109 | 29.5 | 31 | 13.8 | 103 | 62 | ||||
| 德国 | 45 | 17.9 | 93 | 18.2 | 69 | 18.7 | 24 | 10.7 | 16 | 9.6 | ||||
| 法国 | 22 | 8.7 | 57 | 11.2 | 30 | 8.1 | 26 | 11.6 | 11 | 6.6 | ||||
Table 1 The number of plant science publications originating from four countries (China, America, Germany and France) in 2017, based on five plant science journals (data sources: Web of Science)
| The Plant Cell | Plant Physiology | The Plant Journal | Nature Plants | Molecular Plant | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | 文章数量 | 所占比例 (%) | |||||
| 美国 | 99 | 39.3 | 170 | 33.3 | 108 | 29.2 | 51 | 22.8 | 42 | 25.3 | ||||
| 中国 | 63 | 25 | 127 | 24.9 | 109 | 29.5 | 31 | 13.8 | 103 | 62 | ||||
| 德国 | 45 | 17.9 | 93 | 18.2 | 69 | 18.7 | 24 | 10.7 | 16 | 9.6 | ||||
| 法国 | 22 | 8.7 | 57 | 11.2 | 30 | 8.1 | 26 | 11.6 | 11 | 6.6 | ||||
| 1 | 毕国志, 周俭民 (2017). 厚积薄发: 我国植物-微生物互作研究取得突破. 植物学报 52, 685-688. |
| 2 | 马爱民, 漆小泉 (2018). 利用多组学手段解析番茄育种过程中代谢物变化的机制. 植物学报 53(5), doi: 10.11983/CBB1-8053. |
| 3 | 武维华, 朱玉贤, 种康 (2018). 从跟踪到引领——快速发展的中国植物科学. 科技导报 36, doi: 10.3981/j.issn.1000-7857.2018. |
| 4 | 习近平 (2017). 习近平致第十九届国际植物学大会的贺信. (来源: 新华社深圳7月24日电. |
| 5 | 许淑娟, 种康 (2018). “先驱”转录因子LEC1在早期胚胎重置春化状态的机制. 植物学报 53, 1-4. |
| 6 | An CP, Li L, Zhai QZ, You YR, Deng L, Wu FM, Chen R, Jiang HL, Wang H, Chen Q, Li CY (2017a). Mediator subunit MED25 links the jasmonate receptor to trans- criptionally active chromatin.Proc Natl Acad Sci USA 114, E8930-E8939. |
| 7 | An D, Chen JG, Gao YQ, Li X, Chao ZF, Chen ZR, Li QQ, Han ML, Wang YL, Wang YF, Chao DY (2017b). AtHKT1 drives adaptation of Arabidopsis thaliana to salinity by redu- cing floral sodium content. PLoS Genet 13, e1007086. |
| 8 | Bai XF, Huang Y, Hu Y, Liu HY, Zhang B, Smaczniak C, Hu G, Han ZM, Xing YZ (2017). Duplication of an upstream silencer of FZP increases grain yield in rice.Nat Plants 3, 885-893. |
| 9 | Cai HY, Zhao LH, Wang LL, Zhang M, Su ZX, Cheng Y, Zhao HM, Qin Y (2017a). ERECTA signaling controls Arabidopsis inflorescence architecture through chromatin- mediated activation of PRE1 expression. New Phytol 214, 1579-1596. |
| 10 | Cai SG, Chen G, Wang YY, Huang YQ, Marchant DB, Wang YZ, Yang Q, Dai F, Hills A, Franks PJ, Nevo E, Soltis DE, Soltis PS, Sessa E, Wolf PG, Xue DW, Zhang GP, Pogson BJ, Blatt MR, Chen ZH (2017b). Evo- lutionary conservation of ABA signaling for stomatal closure.Plant Physiol 174, 732-747. |
| 11 | Cao MJ, Zhang YL, Liu X, Huang H, Zhou XE, Wang WL, Zeng A, Zhao CZ, Si T, Du J, Wu WW, Wang FX, Xu HE, Zhu JK (2017). Combining chemical and genetic approa- ches to increase drought resistance in plants.Nat Com- mun 8, 1183. |
| 12 | Causier B, Schwarz-Sommer Z, Davies B (2010). Floral organ identity: 20 years of ABCs.Sem Cell Dev Biol 21, 73-79. |
| 13 | Chao LM, Liu YQ, Chen DY, Xue XY, Mao YB, Chen XY (2017). Arabidopsis transcription factors SPL1 and SPL12 confer plant thermotolerance at reproductive stage.Mol Plant 10, 735-748. |
| 14 | Chen H, Cao Y, Li Y, Xia Z, Xie J, Carr JP, Wu B, Fan Z, Zhou T (2017a). Identification of differentially regulated maize proteins conditioning Sugarcane mosaic virus syste- mic infection. New Phytol 215, 1156-1172. |
| 15 | Chen L, Yan Z, Xia Z, Cheng Y, Jiao Z, Sun B, Zhou T, Fan Z (2017b). A violaxanthin deepoxidase interacts with a vi- ral suppressor of RNA silencing to inhibit virus amplification.Plant Physiol 175, 1774-1794. |
| 16 | Chen ST, He NY, Chen JH, Guo FQ (2017c). Identification of core subunits of photosystem II as action sites of HSP21, which is activated by the GUN5-mediated retrograde path- way in Arabidopsis.Plant J 89, 1106-1118. |
| 17 | Chen XY, Zhang H, Sun HY, Luo HB, Zhao L, Dong ZB, Yan SS, Zhao C, Liu RY, Xu CY, Li S, Chen HB, Jin WW (2017d). IRREGULAR POLLEN EXINE1 is a novel factor in anther cuticle and pollen exine formation.Plant Physiol 173, 307-325. |
| 18 | Cheng CH, Wang ZJ, Ren ZY, Zhi LY, Yao B, Su C, Liu L, Li X (2017). SCFAtPP2-B11 modulates ABA signaling by facilitating SnRK2.3 degradation in Arabidopsis thaliana. PLoS Genet 13, e1006947. |
| 19 | Chu S, Wang J, Zhu Y, Liu S, Zhou X, Zhang H, Wang CE, Yang W, Tian Z, Cheng H, Yu D (2017). An R2R3-type MYB transcription factor, GmMYB29, regulates isoflavone biosynthesis in soybean.PLoS Genet 13, e1006770. |
| 20 | Cui Y, Zhao Q, Xie HT, Wong WS, Wang XF, Gao CJ, Ding Y, Tan YQ, Ueda T, Zhang Y, Jiang LW (2017). MON- ENSIN SENSITIVITY1 (MON1)/CALCIUM CAFFEINE ZINC SENSITIVITY1 (CCZ1)-mediated Rab7 activation regulates tapetal programmed cell death and pollen deve- lopment.Plant Physiol 173, 206-218. |
| 21 | Da QG, Wang P, Wang ML, Sun T, Jin HL, Liu B, Wang JF, Grimm B, Wang HB (2017). Thioredoxin and NADPH- dependent thioredoxin reductase C regulation of tetrapyr- role biosynthesis.Plant Physiol 175, 652-666. |
| 22 | Dai XZ, Bai YH, Zhao LH, Dou XY, Liu YH, Wang LL, Li Y, Li WM, Hui YN, Huang XY, Wang ZH, Qin Y (2017). H2A.Z represses gene expression by modulating promoter nucleosome structure and enhancer histone modifications in Arabidopsis.Mol Plant 10, 1274-1292. |
| 23 | Deng YW, Zhai KR, Xie Z, Yang DY, Zhu XD, Liu JZ, Wang X, Qin P, Yang YZ, Zhang GM, Li Q, Zhang JF, Wu SQ, Milazzo J, Mao BZ, Wang ET, Xie HA, Tharreau D, He ZH (2017). Epigenetic regulation of antagonistic receptors confers rice blast resistance with yield balance.Science 355, 962-965. |
| 24 | Ding J, Chen L, Ji C, Hugelius G, Li Y, Liu L, Qin S, Zhang B, Yang G, Li F, Fang K, Chen Y, Peng Y, Zhao X, He H, Smith P, Fang J, Yang Y (2017). Decadal soil carbon accumulation across Tibetan permafrost regions.Nat Geo- sci 10, 420-424. |
| 25 | Ding YL, Li H, Zhang XY, Xie Q, Gong ZZ, Yang SH (2015). OST1 kinase modulates freezing tolerance by enhancing ICE1 stability in Arabidopsis.Dev Cell 32, 278-289. |
| 26 | Dong CH, Agarwal M, Zhang YY, Xie Q, Zhu JK (2006). The negative regulator of plant cold responses, HOS1, is a RING E3 ligase that mediates the ubiquitination and degra- dation of ICE1.Proc Natl Acad Sci USA 103, 8281-8286. |
| 27 | Dong J, Ni W, Yu R, Deng XW, Chen HD, Wei N (2017a). Light-dependent degradation of PIF3 by SCFEBF1/2 pro- motes a photomorphogenic response in Arabidopsis.Curr Biol 27, 2420-2430. |
| 28 | Dong J, Pineros MA, Li X, Yang H, Liu Y, Murphy AS, Kochian LV, Liu D (2017b). An Arabidopsis ABC trans- porter mediates phosphate deficiency-induced remodeling of root architecture by modulating iron homeostasis in roots.Mol Plant 10, 244-259. |
| 29 | Du AP, Tian W, Wei MH, Yan W, He H, Zhou D, Huang X, Li SG, Ouyang XH (2017a). The DTH8-Hd1 module me- diates day length dependent regulation of rice flowering.Mol Plant 10, 948-961. |
| 30 | Du HL, Yu Y, Ma YF, Gao Q, Cao YH, Chen Z, Ma B, Qi M, Li Y, Zhao XF, Wang J, Liu KF, Qin P, Yang X, Zhu LH, Li SG, Liang CZ (2017b). Sequencing and de novo as- sembly of a near complete indica rice genome. Nat Commun 8, 15324. |
| 31 | Du M, Zhao J, Tzeng DTW, Liu Y, Deng L, Yang T, Zhai Q, Wu F, Huang Z, Zhou M, Wang Q, Chen Q, Zhong S, Li C, Li C (2017c). MYC2 orchestrates a hierarchical transcriptional cascade that regulates jasmonate-mediated plant immunity in tomato.Plant Cell 29,1883-1906. |
| 32 | Du YF, Liu L, Li MF, Fang S, Shen XM, Chu JF, Zhang ZX (2017d).UNBRANCHED3 regulates branching by modu- lating cytokinin biosynthesis and signaling in maize and rice. New Phytol 214, 721-733. |
| 33 | Duan CG, Wang X, Zhang L, Xiong X, Zhang Z, Tang K, Pan L, Hsu CC, Xu H, Tao WA, Zhang H, Zhu JK (2017a). A protein complex regulates RNA processing of intronic heterochromatin-containing genes in Arabidopsis.Proc Natl Acad Sci USA 114, E7377-E7384. |
| 34 | Duan NB, Bai Y, Sun HH, Wang N, Ma YM, Li MJ, Wang X, Jiao C, Legall N, Mao LY, Wan SB, Wang K, He TM, Feng SQ, Zhang ZY, Mao ZQ, Shen X, Chen XL, Jiang YM, Wu SJ, Yin CM, Ge SF, Yang L, Jiang SH, Xu HF, Liu JX, Wang DY, Qu CZ, Wang YC, Zuo WF, Xiang L, Liu C, Zhang DY, Gao Y, Xu YM, Xu KN, Chao T, Fazio G, Shu H, Zhong GY, Cheng LL, Fei ZJ, Chen XS (2017b). Genome re-sequencing reveals the history of apple and supports a two-stage model for fruit enlarge- ment.Nat Commun 8, 249. |
| 35 | Duan PG, Xu JS, Zeng D, Zhang BL, Geng MF, Zhang GZ, Huang K, Huang LJ, Xu R, Ge S, Qian Q, Li YH (2017c). Natural variation in the promoter of GSE5 contributes to grain size diversity in rice. Mol Plant 10, 685-694. |
| 36 | Editorial Office of Nature Plants (2017). A Chinese re- naissance.Nat Plants 3, 17006. |
| 37 | Fang L, Wang Q, Hu Y, Jia Y, Chen J, Liu B, Zhang Z, Guan X, Chen S, Zhou B, Mei G, Sun J, Pan Z, He S, Xiao S, Shi W, Gong W, Liu J, Ma J, Cai C, Zhu X, Guo W, Du X, Zhang T (2017). Genomic analyses in cotton identify signatures of selection and loci associated with fiber quality and yield traits.Nat Genet 49, 1089-1098. |
| 38 | Feng C, Xu MZ, Feng C, von Wettberg EJB, Kang M (2017a). The complete chloroplast genome of Primulina and two novel strategies for development of high poly- morphic loci for population genetic and phylogenetic studies. BMC Evol Biol 17, 224. |
| 39 | Feng QN, Song SJ, Yu SX, Wang JG, Li S, Zhang Y (2017b). Adaptor protein-3-dependent vacuolar trafficking involves a subpopulation of COPII and HOPS tethering proteins.Plant Physiol 174, 1609-1620. |
| 40 | Feng XJ, Li JR, Qi SL, Lin QF, Jin JB, Hua XJ (2016). Light affects salt stress-induced transcriptional memory of P5- CS1 in Arabidopsis.Proc Natl Acad Sci USA 113, E8335-E8343. |
| 41 | Feng Y, Xu P, Li BS, Li PP, Wen X, An FY, Gong Y, Xin Y, Zhu ZQ, Wang YC, Guo HW (2017c). Ethylene promotes root hair growth through coordinated EIN3/EIL1 and RHD6/RSL1 activity in Arabidopsis.Proc Natl Acad Sci USA 114, 13834-13839. |
| 42 | Gao H, Zhang YH, Wang WL, Zhao KK, Liu CM, Bai L, Li R, Guo Y (2017). Two membrane-anchored aspartic pro- teases contribute to pollen and ovule development.Plant Physiol 173, 219-239. |
| 43 | Ge ZX, Bergonci T, Zhao YL, Zou YJ, Du S, Liu MC, Luo XJ, Ruan H, García-Valencia LE, Zhong S, Hou SY, Huang QP, Lai LH, Moura DS, Gu HY, Dong J, Wu HM, Dresselhaus T, Xiao JY, Cheung AY, Qu LJ (2017). Arabidopsis pollen tube integrity and sperm release are regulated by RALF-mediated signaling.Science 358, 1596-1600. |
| 44 | Gu YZ, Li W, Jiang HW, Wang Y, Gao HH, Liu M, Chen QS, Lai YC, He CY (2017). Differential expression of a WRKY gene between wild and cultivated soybeans correlates to seed size. J Exp Bot 68, 2717-2729. |
| 45 | Guo C, Xu Y, Shi M, Lai Y, Wu X, Wang H, Zhu Z, Poethig RS, Wu G (2017a). Repression of miR156 by miR159 regulates the timing of the juvenile-to-adult transition in Arabidopsis.Plant Cell 29, 1293-1304. |
| 46 | Guo P, Li Z, Huang P, Li B, Fang S, Chu J, Guo H (2017b). A tripartite amplification loop involving the transcription factor WRKY75, salicylic acid, and reactive oxygen species accelerates leaf senescence.Plant Cell 29, 2854-2870. |
| 47 | Guo T, Mao X, Zhang H, Zhang Y, Fu M, Sun Z, Kuai P, Lou Y, Fang Y (2017c). Lamin-like proteins negatively regulate plant immunity through NAC WITH TRANSMEM- BRANE MOTIF1-LIKE9 and NONEXPRESSOR OF PR GENES1 in Arabidopsis thaliana. Mol Plant 10, 1334-1348. |
| 48 | He Y, Zhang H, Sun Z, Li J, Hong G, Zhu Q, Zhou X, Mac- Farlane S, Yan F, Chen J (2017). Jasmonic acid-media- ted defense suppresses brassinosteroid-mediated susceptibility to rice black streaked dwarf virus infection in rice. New Phytol 214, 388-399. |
| 49 | Hettenhausen C, Li J, Zhuang H, Sun H, Xu Y, Qi J, Zhang J, Lei Y, Qin Y, Sun G, Wang L, Baldwin IT, Wu J (2017). Stem parasitic plant Cuscuta australis(dodder) transfers herbivory-induced signals among plants. Proc Natl Acad Sci USA 114, E6703-E6709. |
| 50 | Hong HZ, Shen R, Zhang FT, Wen ZZ, Chang SW, Lin WF, Kranz SA, Luo YW, Kao SJ, Morel FMM, Shi DL (2017). The complex effects of ocean acidification on the promi- nent N2-fixing cyanobacterium Trichodesmium. Science 356, 527-531. |
| 51 | Hu JL, Yang HJ, Mu JY, Lu TC, Peng JL, Deng X, Kong ZS, Bao SL, Cao XF, Zuo JR (2017a). Nitric oxide reg- ulates protein methylation during stress responses in pla- nts.Mol Cell 67, 702-710. |
| 52 | Hu Q, Li Y, Wang H, Shen Y, Zhang C, Du G, Tang D, Cheng Z (2017b). Meiotic Chromosome Association 1 interacts with TOP3alpha and regulates meiotic recombination in rice.Plant Cell 29, 1697-1708. |
| 53 | Huang J, Gu L, Zhang Y, Yan T, Kong G, Kong L, Guo B, Qiu M, Wang Y, Jing M, Xing W, Ye W, Wu Z, Zhang Z, Zheng X, Gijzen M, Wang Y, Dong S (2017). An oomycete plant pathogen reprograms host pre-mRNA splicing to subvert immunity.Nat Commun 8, 2051. |
| 54 | Huo X, Wu S, Zhu ZF, Liu FX, Fu YC, Cai HW, Sun XY, Gu P, Xie DX, Tan LB, Sun CQ (2017). NOG1 increases grain production in rice.Nat Commun 8, 1497. |
| 55 | Jang SH, An GH, Li HY (2017). Rice leaf angle and grain size are affected by the OsBUL1 transcriptional activator complex.Plant Physiol 173, 688-702. |
| 56 | Jiang BC, Shi YT, Zhang XY, Xin XY, Qi LJ, Guo HW, Li JG, Yang SH (2017a). PIF3 is a negative regulator of the CBF pathway and freezing tolerance in Arabidopsis.Proc Natl Acad Sci USA 114, E6695-E6702. |
| 57 | Jiang L, Liu X, Xiong GS, Liu HH, Chen FL, Wang L, Meng XB, Liu GF, Yu H, Yuan YD, Yi W, Zhao LH, Ma HL, He YZ, Wu ZS, Melcher K, Qian Q, Xu HE, Wang YH, Li JY (2013). DWARF 53 acts as a repressor of strigolactone signaling in rice.Nature 504, 401-405. |
| 58 | Jiang YN, Wang WX, Xie QJ, Liu N, Liu LX, Wang DP, Zhang XW, Yang C, Chen XY, Tang DZ, Wang ET (2017b). Plants transfer lipids to sustain colonization by mutualistic mycorrhizal and parasitic fungi. Science 356, 1172-1175. |
| 59 | Jiang YX, Wang J, Xie YR, Chen NZ, Huang SJ (2017c). ADF10 shapes the overall organization of apical actin filaments by promoting their turnover and ordering in pollen tubes.J Cell Sci 130, 3988-4001. |
| 60 | Jiao YQ, Wang YH, Xue DW, Wang J, Yan MX, Liu GF, Dong GJ, Zeng DL, Zhu XD, Qian Q, Li JY (2010). Regulation of OsSPL14 by OsmiR156 defines ideal plant architecture in rice.Nat Genet 42, 541-544. |
| 61 | Kang E, Zheng M, Zhang Y, Yuan M, Yalovsky S, Zhu L, Fu Y (2017). The microtubule-associated protein MAP18 affects ROP2 GTPase activity during root hair growth.Plant Physiol 174, 202-222. |
| 62 | Lang Z, Wang Y, Tang K, Tang D, Datsenka T, Cheng J, Zhang Y, Handa AK, Zhu JK (2017). Critical roles of DNA demethylation in the activation of ripening-induced genes and inhibition of ripening-repressed genes in tomato fruit.Proc Natl Acad Sci USA 114, E4511-E4519. |
| 63 | Lei RH, Li XL, Ma ZB, Lv Y, Hu YR, Yu DQ (2017). Arabidopsis WRKY2 and WRKY34 transcription factors interact with VQ20 protein to modulate pollen development and function.Plant J 91, 962-976. |
| 64 | Leng YJ, Yang YL, Ren DY, Huang LC, Dai LP, Wang YQ, Chen L, Tu ZJ, Gao YH, Li XY, Zhu L, Hu J, Zhang GH, Gao ZY, Guo LB, Kong ZS, Lin YJ, Qian Q, Zeng DL (2017). A rice PECTATE LYASE-LIKE gene is required for plant growth and leaf senescence. Plant Physiol 174, 1151-1166. |
| 65 | Li F, Peng Y, Natali SM, Chen K, Han T, Yang G, Ding J, Zhang D, Wang G, Wang J, Yu J, Liu F, Yang Y (2017a). Warming effects on permafrost ecosystem carbon fluxes associated with plant nutrients.Ecology 98, 2851-2859. |
| 66 | Li GZ, Wu YF, Liu GY, Xiao XH, Wang PF, Gao T, Xu MJ, Han QX, Wang YH, Guo TC, Kang GZ (2017b). Large- scale proteomics combined with transgenic experiments demonstrates an important role of jasmonic acid in potas- sium deficiency response in wheat and rice.Mol Cell Pro- teomics 16, 1889-1905. |
| 67 | Li H, Ding YL, Shi YT, Zhang XY, Zhang SQ, Gong ZZ, Yang SH (2017c). MPK3-and MPK6-mediated ICE1 pho- sphorylation negatively regulates ICE1 stability and freez- ing tolerance in Arabidopsis.Dev Cell 43, 630-642. |
| 68 | Li H, Ye KY, Shi YT, Cheng JK, Zhang XY, Yang SH (2017d). BZR1 positively regulates freezing tolerance via CBF-dependent and CBF-independent pathways in Arabi- dopsis.Mol Plant 10, 545-559. |
| 69 | Li H, Yu M, Du XQ, Wang ZF, Wu WH, Quintero FJ, Jin XH, Li HD, Wang Y (2017e). NRT1.5/NPF7.3 functions as a proton-coupled H(+)/K(+) antiporter for K(+) loading into the xylem in Arabidopsis.Plant Cell 29, 2016-2026. |
| 70 | Li JH, Zhang Y, Song YZ, Zhang H, Fan JB, Li Q, Zhang DF, Xue YB (2017f). Electrostatic potentials of the S-locus F-box proteins contribute to the pollen S specificity in self- incompatibility in Petunia hybrid. Plant J 89, 45-57. |
| 71 | Li Q, Wang J, Ye J, Zheng X, Xiang X, Li C, Fu M, Wang Q, Zhang Z, Wu Y (2017g). The maize imprinted gene Floury3 encodes a PLATZ protein required for tRNA and 5S rRNA transcription through interaction with RNA Poly- merase III. Plant Cell 29, 2661-2675. |
| 72 | Li QQ, Zheng J, Li SZ, Huang GR, Skilling SJ, Wang LJ, Li L, Li MY, Yuan LX, Liu P (2017h). Transporter-mediated nuclear entry of jasmonoyl-isoleucine is essential for jas- monate signaling.Mol Plant 10, 695-708. |
| 73 | Li QT, Lu X, Song Q, Chen HW, Wei W, Tao JJ, Bian XH, Shen M, Ma B, Zhang WK, Bi YD, Li W, Lai YC, Lam SM, Shui G, Chen SY, Zhang JS (2017i). Selection for a zinc-finger protein contributes to seed oil increase during soybean domestication.Plant Physiol 173, 2208-2224. |
| 74 | Li SW, Dong HJ, Pei WK, Liu CN, Zhang S, Sun TT, Xue XH, Ren HY (2017j). LlFH1-mediated interaction between actin fringe and exocytic vesicles is involved in pollen tube tip growth.New Phytol 214, 745-761. |
| 75 | Li T, Xu Y, Zhang L, Ji Y, Tan D, Yuan H, Wang A (2017k). The jasmonate-activated transcription factor MdMYC2 regulates ETHYLENE RESPONSE FACTOR and ethylene biosynthetic genes to promote ethylene biosynthesis dur- ing apple fruit ripening.Plant Cell 29, 1316-1334. |
| 76 | Li W, Lacey RF, Ye Y, Lu J, Yeh KC, Xiao Y, Li L, Wen CK, Binder BM, Zhao Y (2017l). Triplin, a small molecule, reveals copper ion transport in ethylene signaling from ATX1 to RAN1.PLoS Genet 13, e1006703. |
| 77 | Li W, Xu F, Zheng S, Taube F, Bai Y (2017m). Patterns and thresholds of grazing-induced changes in community st- ructure and ecosystem functioning: species-level respon- ses and the critical role of species traits.J Appl Ecol 54, 963-975. |
| 78 | Li WT, Zhu ZW, Chern M, Yin JJ, Yang C, Ran L, Cheng MP, He M, Wang K, Wang J, Zhou XG, Zhu XB, Chen ZX, Wang JC, Zhao W, Ma BT, Qin P, Chen WL, Wang YP, Liu JL, Wang WM, Wu XJ, Li P, Wang JR, Zhu LH, Li SG, Chen XW (2017n). A natural allele of a transcription factor in rice confers broad-spectrum blast resistance.Cell 170, 114-126. |
| 79 | Li Y, Zhang X, Hu S, Liu H, Xu JR (2017o). PKA activity is essential for relieving the suppression of hyphal growth and appressorium formation by MoSfl1 in Magnaporthe oryzae. PLoS Genet 13, e1006954. |
| 80 | Li ZP, Wu SX, Chen JY, Wang XY, Gao J, Ren GD, Kuai BK (2017p). NYEs/SGRs-mediated chlorophyll degrada- tion is critical for detoxification during seed maturation in Arabidopsis.Plant J 92, 650-661. |
| 81 | Lian N, Liu XM, Wang XH, Zhou YY, Li H, Li JG, Mao TL (2017). COP1 mediates dark-specific degradation of mi- crotubule-associated protein WDL3 in regulating Arabi- dopsis hypocotyl elongation. Proc Natl Acad Sci USA 114, 12321-12326. |
| 82 | Lin F, Xu DQ, Jiang Y, Chen HD, Fan LM, Holm M, Deng XW (2017). Phosphorylation and negative regulation of CONSTITUTIVELY PHOTOMORPHOGENIC 1 by PINOID in Arabidopsis.Proc Natl Acad Sci USA 114, 6617-6622. |
| 83 | Ling JJ, Li J, Zhu DM, Deng XW (2017). Noncanonical role of Arabidopsis COP1/SPA complex in repressing BIN2- mediated PIF3 phosphorylation and degradation in darkness.Proc Natl Acad Sci USA 114, 3539-3544. |
| 84 | Litt A (2007). An evaluation of A-function: evidence from the APETALA1 and APETALA2 gene lineages. Int J Plant Sci 168, 73-91. |
| 85 | Liu J, Cheng XL, Liu P, Li DY, Chen T, Gu XF, Sun JQ (2017a). MicroRNA319-regulated TCPs interact with FBHs and PFT1 to activate CO transcription and control flower- ing time in Arabidopsis.PLoS Genet 13, e1006833. |
| 86 | Liu JF, Chen J, Zheng XM, Wu FQ, Lin QB, Heng YQ, Tian P, Cheng ZJ, Yu XW, Zhou KN, Zhang X, Guo XP, Wang JL, Wang HY, Wan JM (2017b). GW5 acts in the brassinosteroid signaling pathway to regulate grain width and weight in rice.Nat Plants 3, 17043. |
| 87 | Liu K, Yu Y, Dong A, Shen WH (2017c).SET DOMAIN GROUP701 encodes a H3K4-methytransferase and regu- lates multiple key processes of rice plant development. New Phytol 215, 609-623. |
| 88 | Liu LP, Jiang Y, Zhang XM, Wang X, Wang YB, Han YZ, Coupland G, Jin JB, Searle I, Fu YF, Chen FL (2017d). Two SUMO proteases SUMO PROTEASE RELATED TO FERTILITY1 and 2 are required for fertility in Arabidopsis.Plant Physiol 175, 1703-1719. |
| 89 | Liu M, Ba Z, Costa-Nunes P, Wei W, Li L, Kong F, Li Y, Chai J, Pontes O, Qi Y (2017e). IDN2 interacts with RPA and facilitates DNA double-strand break repair by homolo- gous recombination in Arabidopsis.Plant Cell 29, 589-599. |
| 90 | Liu N, Hake K, Wang W, Zhao T, Romeis T, Tang D (2017f). CALCIUM-DEPENDENT PROTEIN KINASE5 as- sociates with the truncated nlr protein TIR-NBS2 to contribute to exo70b1-mediated immunity.Plant Cell 29, 746-759. |
| 91 | Liu Q, Wang Q, Deng WX, Wang X, Piao MX, Cai DW, Li YX, Barshop WD, Yu XL, Zhou TT, Liu B, Oka Y, Wohlschlegel J, Zuo ZC, Lin CT (2017g). Molecular ba- sis for blue light-dependent phosphorylation of Arabidopsis cryptochrome 2.Nat Commun 8, 15234. |
| 92 | Liu XB, Liu YS, Huang P, Ma YS, Qing ZX, Tang Q, Cao HF, Cheng P, Zheng YJ, Yuan ZJ, Zhou Y, Liu JF, Tang ZS, Zhuo YX, Zhang YC, Yu LL, Huang JL, Yang P, Zeng JG (2017h). The genome of medicinal plant Mac- leaya cordata provides new insights into benzylisoquinoline alkaloids metabolism. Mol Plant 7, 975-989. |
| 93 | Liu XQ, Liu RL, Li Y, Shen X, Zhong SW, Shi H (2017i). EIN3 and PIF3 form an interdependent module that re- presses chloroplast development in buried seedlings.Plant Cell 29, 3051-3067. |
| 94 | Liu Y, Xie Y, Wang H, Ma X, Yao W, Wang H (2017j). Light and ethylene coordinately regulate the phosphate starva- tion response through transcriptional regulation of PHOS- PHATE STARVATION RESPONSE 1.Plant Cell 29, 2269-2284. |
| 95 | Liu ZY, Jia YX, Ding YL, Shi YT, Li Z, Guo Y, Gong ZZ, Yang SH (2017k). Plasma membrane CRPK1-mediated phosphorylation of 14-3-3 proteins induces their nuclear import to fine-tune CBF signaling during cold response.Mol Cell 66, 117-128. |
| 96 | Lu LM, Mao LF, Yang T, Ye JF, Liu B, Li HL, Sun M, Miller JT, Mathews S, Hu HH, Niu YT, Peng DX, Chen YH, Smith SA, Chen M, Xiang KL, Le CT, Dang VC, Lu AM, Soltis PS, Soltis DE, Li JH, Chen ZD (2017a). Evolutionary history of the angiosperm flora of China. Nature 554, 234-238. |
| 97 | Lu S, Zhao X, Hu Y, Liu S, Nan H, Li X, Fang C, Cao D, Shi X, Kong L, Su T, Zhang F, Li S, Wang Z, Yuan X, Cober ER, Weller JL, Liu B, Hou X, Tian Z, Kong F (2017b). Natural variation at the soybean J locus improves adaptation to the tropics and enhances yield. Nat Genet 49, 773-779. |
| 98 | Luo N, Yan A, Liu G, Guo JZ, Rong DY, Kanaoka MM, Xiao Z, Xu GS, Higashiyama T, Cui XP, Yang ZB (2017a). Exocytosis-coordinated mechanisms for tip grow- th underlie pollen tube growth guidance.Nat Commun 8, 1687. |
| 99 | Luo X, Hu QJ, Zhou PP, Zhang D, Wang Q, Abbott RJ, Liu JQ (2017b). Chasing ghosts: allopolyploid origin of Oxyria sinensis(Polygonaceae) from its only diploid congener and an unknown ancestor. Mol Ecol 26, 3037-3049. |
| 100 | Luo X, Xu N, Huang J, Gao F, Zou H, Boudsocq M, Coaker G, Liu J (2017c). A lectin receptor-like kinase mediates pattern-triggered salicylic acid signaling.Plant Physiol 174, 2501-2514. |
| 101 | Lv XQ, Jing YP, Xiao JW, Zhang YD, Zhu YF, Julian R, Lin JX (2017). Membrane microdomains and the cytoskeleton constrain AtHIR1 dynamics and facilitate the formation of an AtHIR1-associated immune complex.Plant J 90, 3-16. |
| 102 | Ma H, Chen J, Zhang Z, Ma L, Yang Z, Zhang Q, Li X, Xiao J, Wang S (2017a). MAPK kinase 10.2 promotes disease resistance and drought tolerance by activating different MAPKs in rice.Plant J 92, 557-570. |
| 103 | Ma L, Sang XC, Zhang T, Yu ZY, Li YF, Zhao FM, Wang ZW, Wang YT, Yu P, Wang N, Zhang CW, Ling YH, Yang ZL, He GH (2017b).ABNORMAL VASCULAR BUNDLES regulates cell proliferation and procambium cell establishment during aerial organ development in rice. New Phytol 213, 275-286. |
| 104 | Ma L, Xue N, Fu XY, Zhang H, Li G (2017c).Arabidopsis thaliana FAR-RED ELONGATED HYPOCOTYLS3 (FHY3) and FAR-RED-IMPAIRED RESPONSE1 (FAR1) modulate starch synthesis in response to light and sugar. New Phy- tol 213, 1682-1696. |
| 105 | Ma W, Wu F, Sheng P, Wang X, Zhang Z, Zhou K, Zhang H, Hu J, Lin Q, Cheng Z, Wang J, Zhu S, Zhang X, Guo X, Wang H, Wu C, Zhai H, Wan J (2017d). The LBD12-1 transcription factor suppresses apical meristem size by repressing Argonaute 10 expression.Plant Physiol 173, 801-811. |
| 106 | Ma ZC, Zhu L, Song TQ, Wang Y, Zhang Q, Xia YQ, Qiu M, Lin YC, Li HY, Kong L, Fang YF, Ye WW, Wang Y, Dong SM, Zheng XB, Tyler BM, Wang YC (2017e). A paralo- gous decoy protects Phytophthora sojae apoplastic effector PsXEG1 from a host inhibitor. Science 355, 710-714. |
| 107 | Mao Y, Liu Y, Chen D, Chen F, Fang X, Hong G, Wang L, Wang J, Chen X (2017). Jasmonate response decay and defense metabolite accumulation contributes to age-re- gulated dynamics of plant insect resistance.Nat Commun 8, 13925. |
| 108 | Meng WJ, Cheng ZJ, Sang YL, Zhang MM, Rong XF, Wang ZW, Tang YY, Zhang XS (2017). Type-B ARA- BIDOPSIS RESPONSE REGULATORs specify the shoot stem cell niche by dual regulation of WUSCHEL. Plant Cell 29, 1357-1372. |
| 109 | Miura K, Jin JB, Lee J, Yoo CY, Stirm V, Miura T, Ash- worth EN, Bressan RA, Yun DJ, Hasegawa PM (2007). SIZ1-mediated sumoylation of ICE1 controls CBF3/DRE- B1A expression and freezing tolerance in Arabidopsis.Plant Cell 19, 1403-1414. |
| 110 | Nan Q, Qian D, Niu Y, He YX, Tong SF, Niu ZM, Ma JC, Yang Y, An LZ, Wan DS, Xiang Y (2017). Plant actin-depolymerizing factors possess opposing biochemical properties arising from key amino acid changes throughout evolution.Plant Cell 29, 395-408. |
| 111 | Ni F, Wu L, Wang Q, Hong J, Qi Y, Zhou X (2017). Turnip yellow mosaic virus P69 interacts with and suppresses GLK transcription factors to cause pale-green symptoms in Arabidopsis.Mol Plant 10, 764-766. |
| 112 | Ouyang M, Li XY, Zhao S, Pu H, Shen JR, Adam Z, Clausen T, Zhang LX (2017). The crystal structure of Deg9 reveals a novel octameric-type HtrA protease.Nat Plant 3, 973-982. |
| 113 | Peng M, Shahzad R, Gul A, Subthain H, Shen S, Lei L, Zheng Z, Zhou J, Lu D, Wang S, Nishawy E, Liu X, Tohge T, Fernie AR, Luo J (2017). Differentially evolved glucosyltransferases determine natural variation of rice flavone accumulation and UV-tolerance.Nat Commun 8, 1975. |
| 114 | Qiao S, Sun S, Wang L, Wu Z, Li C, Li X, Wang T, Leng L, Tian W, Lu T, Wang X (2017). The RLA1/SMOS1 trans- cription factor functions with OsBZR1 to regulate brassinosteroid signaling and rice architecture.Plant Cell 29, 292-309. |
| 115 | Qin C, Li B, Fan Y, Zhang X, Yu Z, Ryabov E, Zhao M, Wang H, Shi N, Zhang P, Jackson S, Tör M, Cheng Q, Liu Y, Gallusci P, Hong Y (2017a). Roles of dicer-like proteins 2 and 4 in intra- and intercellular antiviral silenc- ing.Plant Physiol 174, 1067-1081. |
| 116 | Qin Z, Wu J, Geng S, Feng N, Chen F, Kong X, Song G, Chen K, Li A, Mao L, Wu L (2017b). Regulation of FT splicing by an endogenous cue in temperate grasses.Nat Commun 8, 14320. |
| 117 | Qin ZR, Bai YX, Wu L (2017c). Flowering on time: multilay- ered restrictions on FT in plants.Mol Plant 10, 1365-1367. |
| 118 | Qu LH, Zhou XM, Li XB, Li SS, Zhao J, Zhao P, Liu Y, Sun MX (2017a). The autonomous cell fate specification of basal cell lineage: the initial round of cell fate specification occurs at the two-celled proembryo stage.Plant J 91, 1051-1063. |
| 119 | Qu XL, Zhang RH, Zhang M, Diao M, Xue YB, Huang SJ (2017b). Organizational innovation of apical actin filaments drives rapid pollen tube growth and turning. Mol Plant 10, 930-947. |
| 120 | Ren HB, Dang X, Cai XZ, Yu PH, Li YJ, Zhang SS, Liu MH, Chen BQ, Lin DS (2017). Spatio-temporal orientation of microtubules controls conical cell shape in Arabidopsis thaliana petals. PLoS Genet 13, e1006851. |
| 121 | Shafiq S, Chen C, Yang J, Cheng L, Ma F, Widemann E, Sun Q (2017). DNA Topoisomerase 1 prevents R-loop accumulation to modulate auxin-regulated root develop- ment in rice.Mol Plant 10, 821-833. |
| 122 | Shao A, Ma WY, Zhao XQ, Hu MY, He X, Teng W, Li H, Tong YP (2017). The auxin biosynthetic TRYPTOPHAN AMINOTRANSFERASE RELATED TaTAR2.1-3A increas- es grain yield of wheat. Plant Physiol 174, 2274-2288 |
| 123 | Shen H, Jin DM, Shu JP, Zhou XL, Lei M, Wei R, Shang H, Wei HJ, Zhang R, Liu L, Gu YF, Zhang XC, Yan YH (2018). Large scale phylogenomic analysis resolves a backbone phylogeny in ferns.Giga Science 7, 1-11. |
| 124 | Shen Q, Bourdais G, Pan H, Robatzek S, Tang D (2017a). Arabidopsis glycosylphosphatidylinositol-anchored protein LLG1 associates with and modulates FLS2 to regulate innate immunity.Proc Natl Acad Sci USA 114, 5749-5754. |
| 125 | Shen RX, Wang L, Liu XP, Wu J, Jin WW, Zhao XC, Xie XR, Zhu QL, Tang HW, Li Q, Chen LT, Liu YG (2017b). Genomic structural variation-mediated allelic suppression causes hybrid male sterility in rice. Nat Commun 8, 1310. |
| 126 | Song XG, Lu ZF, Yu H, Shao GN, Xiong JS, Meng XB, Jing YH, Liu GF, Xiong GS, Duan JB, Yao XF, Liu CM, Li HQ, Wang YH, Li JY (2017). IPA1 functions as a downstream transcription factor repressed by D53 in strigolactone signaling in rice.Cell Res 27, 1128-1141. |
| 127 | Su C, Li Z, Cheng J, Li L, Zhong S, Liu L, Zheng Y, Zheng B (2017a). The protein phosphatase 4 and SMEK1 complex dephosphorylates HYL1 to promote miRNA biogenesis by antagonizing the MAPK cascade in Arabidopsis.Dev Cell 41, 527-539. |
| 128 | Su H, Cheng ZH, Huang JY, Lin J, Copenhaver GP, Ma H, Wang YX (2017b). Arabidopsis RAD51, RAD51C and XRCC3 proteins form a complex and facilitate RAD51 localization on chromosomes for meiotic recombination.PLoS Genet 13, e1006827. |
| 129 | Su Y, Wang S, Zhang F, Zheng H, Liu Y, Huang T, Ding Y (2017c). Phosphorylation of histone H2A at Serine 95: a plant-specific mark involved in flowering time regulation and H2A.Z deposition.Plant Cell 29, 2197-2213. |
| 130 | Sun C, Yan K, Han JT, Tao L, Lv MH, Shi T, He YX, Wierzba M, Tax FE, Li J (2017a). Scanning for new BRI1 mutations via TILLING analysis.Plant Physiol 174, 1881-1896. |
| 131 | Sun FM, Fan GY, Hu Q, Zhou YM, Guan M, Tong CB, Li JN, Du DZ, Qi CK, Jiang LC, Liu WQ, Huang SM, Chen WB, Yu JY, Mei DS, Meng JL, Zeng P, Shi JQ, Liu KD, Wang X, Wang XF, Long Y, Liang XM, Hu ZY, Huang GD, Dong CH, Zhang H, Li J, Zhang YL, Li LW, Shi CC, Wang JH, Lee SMY, Guan CY, Xu X, Liu SY, Liu X, Chalhoub B, Hua W, Wang HZ (2017b). The high-quality genome of Brassica napus cultivar ‘ZS11’ reveals the in- trogression history in semi-winter morphotype. Plant J 92, 452-468. |
| 132 | Sun WQ, Gao DW, Xiong Y, Tang XX, Xiao XF, Wang CR, Yu SB (2017c). Hairy Leaf 6, an AP2/ERF transcription factor, interacts with OsWOX3B and regulates trichome formation in rice.Mol Plant 10, 1417-1433. |
| 133 | Sun X, Li Y, He W, Ji C, Xia P, Wang Y, Du S, Li H, Raikhel N, Xiao J, Guo H (2017d). Pyrazinamide and derivatives block ethylene biosynthesis by inhibiting ACC oxidase.Nat Commun 8, 15758. |
| 134 | Sun Y, Ji K, Liang B, Du Y, Jiang L, Wang J, Kai W, Zhang Y, Zhai X, Chen P, Wang H, Leng P (2017e). Suppress- ing ABA uridine diphosphate glucosyltransferase (SlUGT- 75C1) alters fruit ripening and the stress response in tomato.Plant J 91, 574-589. |
| 135 | Sun Y, Wu Y, Yang CW, Sun S, Lin XY, Liu LX, Xu CM, Wendel JF, Gong L, Liu B (2017f). Segmental allote- traploidy generates extensive homoeologous expression rewiring and phenotypic diversity at the population level in rice.Mol Ecol 26, 5451-5466. |
| 136 | Tang X, Lowder LG, Zhang T, Malzahn AA, Zheng XL, `Voytas DF, Zhong ZH, Chen YY, Ren QR, Li Q, Kirkland ER, Zhang Y, Qi YP (2017a). A CRISPR-Cpf1 system for efficient genome editing and transcriptional repression in plants.Nat Plants 3, 17018. |
| 137 | Tang Y, Liu X, Liu X, Li Y, Wu K, Hou X (2017b). Arabidopsis NF-YCs mediate the light-controlled hypocotyl elongation via modulating histone acetylation.Mol Plant 10, 260-273. |
| 138 | Tang YY, Liu HH, Guo SY, Wang B, Li ZT, Chong K, Xu YY (2018). OsmiR396d miRNA affects gibberellin and brass- inosteroid signaling to regulate plant architecture.Plant Physiology 176, 946-959. |
| 139 | Tao Z, Shen LS, Gu XF, Wang YZ, Yu H, He YH (2017). Embryonic epigenetic reprogramming by a pioneer tran- scription factor in plants.Nature 551, 124-128. |
| 140 | Theißen G, Becker A, Di Rosa A, Kanno A, Kim JT, Münster T, Winter KU, Saedler H (2000). A short history of MADS-box genes in plants.Plant Mol Biol 42, 115-149. |
| 141 | Tieman D, Zhu GT, Resender M, Lin T, Nguyen C, Bies D, Rambla JL, Beltran KSO, Taylor M, Zhang B, Ikeda H, Liu ZY, Fisher J, Zemach I, Monforte A, Zamir D, Gra- nell A, Kirst M, Huang SW, Klee H (2017). A chemical genetic roadmap to improved tomato flavor.Science 355, 391-394. |
| 142 | Wang C, Wang G, Zhang C, Zhu P, Dai H, Yu N, He Z, Xu L, Wang E (2017a). OsCERK1-mediated chitin perception and immune signaling requires receptor-like cytoplasmic kinase 185 to activate an MAPK cascade in rice.Mol Plant 10, 619-633. |
| 143 | Wang DX, Yang CJ, Wang HJ, Wu ZH, Jiang JJ, Liu JJ, He ZN, Chang F, Ma H, Wang XL (2017b). BKI1 regulates plant architecture through coordinated inhibition of the brassinosteroid and ERECTA signaling pathways in Ara- bidopsis.Mol Plant 10, 297-308. |
| 144 | Wang H, Li Y, Pan J, Lou D, Hu Y, Yu D (2017c). The bHLH transcription factors MYC2, MYC3, and MYC4 are required for jasmonate-mediated inhibition of flowering in Arabidopsis.Mol Plant 10, 1461-1464. |
| 145 | Wang H, Prentice IC, Keenan TF, Davis TW, Wright IJ, Cornwell WK, Evans BJ, Peng C (2017d). Towards a universal model for carbon dioxide uptake by plants.Nat Plant 3, 734-741. |
| 146 | Wang HJ, Hsu YW, Guo CL, Jane WN, Wang H, Jiang L, Jauh GY (2017e). VPS36-dependent multivesicular bodies are critical for plasmamembrane protein turnover and vacuolar biogenesis.Plant Physiol 173, 566-581. |
| 147 | Wang J, Tian CH, Zhang C, Shi BH, Cao XW, Zhang TQ, Zhao Z, Wang JW, Jiao YL (2017f). Cytokinin signaling activates WUSCHEL expression during axillary meristem initiation. Plant Cell 29, 1373-1387. |
| 148 | Wang J, Wang R, Wang Y, Zhang L, Zhang L, Xu Y, Yao S (2017g). Short and Solid Culm/RFL/APO2 for culm deve- lopment in rice.Plant J 91, 85-96. |
| 149 | Wang J, Yu H, Xiong GS, Lu ZF, Jiao YQ, Meng XB, Liu GF, Chen XW, Wang YH, Li JY (2017h). Tissue-specific ubiquitination by IPA1 INTERACTING PROTEIN1 modu- lates IPA1 protein levels to regulate plant architecture in rice.Plant Cell 29, 697-707. |
| 150 | Wang JG, Feng C, Liu HH, Feng QN, Li S, Zhang Y (2017i). AP1G mediates vacuolar acidification during syner- gid-controlled pollen tube reception.Proc Natl Acad Sci USA 114, E4877-E4883. |
| 151 | Wang K, Tu WF, Liu C, Rao Y, Gao ZM, Yang CH (2017j). 9-cis-neoxanthin in light harvesting complexes of photosystem II regulates the binding of violaxanthin and xanthophyll cycle. Plant Physiol 174, 86-96. |
| 152 | Wang L, Ran L, Hou Y, Tian Q, Li C, Liu R, Fan D, Luo K (2017k). The transcription factor MYB115 contributes to the regulation of proanthocyanidin biosynthesis and enhances fungal resistance in poplar.New Phytol 215, 351-367. |
| 153 | Wang M, Tu L, Lin M, Lin Z, Wang P, Yang Q, Ye Z, Shen C, Li J, Zhang L, Zhou X, Nie X, Li Z, Guo K, Ma Y, Huang C, Jin S, Zhu L, Yang X, Min L, Yuan D, Zhang Q, Lindsey K, Zhang X (2017l). Asymmetric subgenome selection and cis-regulatory divergence during cotton do- mestication. Nat Genet 49, 579-587. |
| 154 | Wang SS, Lu JN, Song XF, Ren SC, You CJ, Xu J, Liu CM, Ma H, Chang F (2017m). Cytological and transcriptomic analyses reveal important roles of CLE19 in pollen exine formation.Plant Physiol 175, 1186-1202. |
| 155 | Wang WH, Li JY, Sun QQ, Yu XY, Zhang WW, Jia N, An CJ, Li YQ, Dong YN, Han FJ, Chang N, Liu XM, Zhu ZL, Yu Y, Fan SL, Yang MJ, Luo SZ, Gao HB, Feng Y (2017n). Structural insights into the coordination of plastid division by the ARC6-PDV2 complex.Nat Plant 3, 17011. |
| 156 | Wang X, Wang Q, Han YJ, Liu Q, Gu LF, Yang ZH, Su J, Liu BB, Zuo ZC, He WJ, Wang J, Liu B, Matsui M, Kim JI, Oka Y, Lin CT (2017o). A CRY-BIC negative-feedback circuitry regulating blue light sensitivity of Arabidopsis.Plant J 92, 426-436. |
| 157 | Wang X, Xu YT, Zhang SQ, Cao L, Huang Y, Cheng JF, Wu GZ, Tian SL, Chen CL, Liu Y, Yu HW, Yang XM, Lan H, Wang N, Wang L, Xu JD, Jiang XL, Xie ZZ, Tan ML, Larkin RM, Chen LL, Ma BG, Ruan YJ, Deng XX, Xu Q (2017p). Genomic analyses of primitive, wild and cultivated citrus provide insights into asexual reproduction.Nat Ge- net 49, 765-772. |
| 158 | Wang YJ, Mueller-Schaerer H, van Kleunen M, Cai AM, Zhang P, Yan R, Dong BC, Yu FH (2017q). Invasive alien plants benefit more from clonal integration in heterogeneous environments than natives.New Phytol 216, 1072-1078. |
| 159 | Wu F, Shi X, Lin X, Liu Y, Chong K, Theißen G, Meng Z (2017a). The ABCs of flower development: mutational analysis of AP1/FUL-like genes in rice provides evidence for a homeotic (A)-function in grasses. Plant J 89, 310-324. |
| 160 | Wu J, Yang R, Yang Z, Yao S, Zhao S, Wang Y, Li P, Song X, Jin L, Zhou T, Lan Y, Xie L, Zhou X, Chu C, Qi Y, Cao X, Li Y (2017b). ROS accumulation and antiviral defence control by microRNA528 in rice.Nat Plants 3, 16203. |
| 161 | Wu L, Liu D, Wu J, Zhang R, Qin Z, Li A, Fu D, Zhai W, Mao L (2013). Regulation of FLOWERING LOCUS T by a microRNA in Brachypodium distachyon. Plant Cell 25, 4363-4377. |
| 162 | Wu Q, Han TS, Chen X, Chen JF, Zou YP, Li ZW, Xu YC, Guo YL (2017c). Long-term balancing selection contri- butes to adaptation in Arabidopsis and its relatives.Ge- nome Biol 18, 217. |
| 163 | Wu WG, Liu XY, Wang MH, Meyer RS, Luo XJ, Ndjiondjop M, Tan LB, Zhang JW, Wu JZ, Cai HW, Sun CQ, Wang XK, Wing RA, Zhu ZF (2017d). A single-nucleotide polymorphism causes smaller grain size and loss of seed shattering during African rice domestication.Nat Plants 3, 17064. |
| 164 | Xia EH, Zhang HB, Sheng J, Li K, Zhang QJ, Kim CH, Zhang Y, Liu Y, Zhu T, Li W, Huang H, Tong Y, Nan H, Shi C, Shi C, Jiang JJ, Mao SY, Jiao JY, Zhang D, Zhao Y, Zhao YJ, Zhang LP, Liu YL, Liu BY, Yu Y, Shao SF, Ni DJ, Eichler E, Gao LZ (2017). The tea tree genome provides insights into tea flavor and independent evolution of caffeine biosynthesis.Mol Plant 10, 866-877. |
| 165 | Xiang KL, Zhao L, Erst AS, Yu SX, Jabbour F, Wang W (2017a). A molecular phylogeny of Dichocarpum(Ranun- culaceae): implications for eastern Asian biogeography. Mol Phylogenet Evol 107, 594-604. |
| 166 | Xiang Y, Sun X, Gao S, Qin F, Dai M (2017b). Deletion of an endoplasmic reticulum stress response element in a Zm- PP2C-A gene facilitates drought tolerance of maize seed- lings. Mol Plant 10, 456-469. |
| 167 | Xie JB, Yang XH, Song YP, Du QZ, Li Y, Chen JH, Zhang DQ (2017a). Adaptive evolution and functional innovation of Populus-specific recently evolved microRNAs. New Phy- tol 213, 206-219. |
| 168 | Xie YR, Liu Y, Wang H, Ma XJ, Wang BB, Wu GX, Wang HY (2017b). Phytochrome-interacting factors directly suppress MIR156 expression to enhance shade-avoidance syndrome in Arabidopsis.Nat Commun 8, 348. |
| 169 | Xiong Q, Ma B, Lu X, Huang YH, He SJ, Yang C, Yin CC, Zhao H, Zhou Y, Zhang WK, Wang WS, Li ZK, Chen SY, Zhang JS (2017). Ethylene-inhibited jasmonic acid bio- synthesis promotes mesocotylcoleoptile elongation of etio- lated rice seedlings.Plant Cell 29, 1053-1072. |
| 170 | Xu CX, Jiao C, Sun HH, Cai XF, Wang XL, Ge CH, Zheng Y, Liu WL, Sun XP, Xu YM, Deng J, Zhang ZH, Huang SW, Dai SJ, Mou BQ, Wang QX, Fei ZJ, Wang QH (2017a). Draft genome of spinach and transcriptome div- ersity of 120 Spinacia accessions.Nat Commun 8, 15275. |
| 171 | Xu DW, Shi JX, Rautengarten C, Yang L, Qian XL, Uzair M, Zhu L, Luo Q, An G, Waßmann F, Schreiber L, Heazlewood JL, Scheller HV, Hu JP, Zhang DB, Liang WQ (2017b).Defective Pollen Wall 2 (DPW2) encodes an acyl transferase required for rice pollen development. Plant Physiol 173, 240-255. |
| 172 | Xu L, Zhao H, Ruan W, Deng M, Wang F, Peng J, Luo J, Chen Z, Yi K (2017c). ABNORMAL INFLORESCENCE MERISTEM1 functions in salicylic acid biosynthesis to maintain proper reactive oxygen species levels for root meristem activity in rice.Plant Cell 29, 560-574. |
| 173 | Xu SH, He ZW, Guo ZX, Zhang Z, Wyckoff GJ, Greenberg A, Wu CI, Shi SH (2017d). Genome-wide convergence during evolution of mangroves from woody plants.Mol Biol Evol 34, 1008-1015. |
| 174 | Yan J, Wang P, Wang B, Hsu CC, Tang K, Zhang H, Hou YJ, Zhao Y, Wang Q, Zhao C, Zhu X, Tao WA, Li J, Zhu JK (2017a). The SnRK2 kinases modulate miRNA accu- mulation in Arabidopsis.PLoS Genet 13, e1006753. |
| 175 | Yan T, Chen M, Shen Q, Li L, Fu X, Pan Q, Tang Y, Shi P, Lv Z, Jiang W, Ma YN, Hao X, Sun X, Tang K (2017b). HOMEODOMAIN PROTEIN 1 is required for jasmonate- mediated glandular trichome initiation in Artemisia annua. New Phytol 213, 1145-1155. |
| 176 | Yang BJ, Lin WH, Fu FF, Xu ZH, Xue HW (2017a). Receptor-like protein ELT1 promotes brassinosteroid signaling through interacting with and suppressing the endocytosis-mediated degradation of receptor BRI1.Cell Res 27, 1182-1185. |
| 177 | Yang HX, Li P, Zhang AH, Wen XG, Zhang LX, Lu CM (2017b). Tetratricopeptide repeat protein Pyg7 is essential for photosystem I assembly by interacting with PsaC in Arabidopsis.Plant J 91, 950-961. |
| 178 | Yang J, Moeinzadeh M, Kuhl H, Helmuth J, Xiao P, Haas S, Liu GL, Zheng JL, Sun Z, Fan WJ, Deng GF, Wang HX, Hu FH, Zhao SS, Fernie A, Boerno S, Timmermann B, Zhang P, Vingron M (2017c). Haplotype-resolved sweet potate genome traces back its hexaploidization history.Nat Plants 3, 696-703. |
| 179 | Yang J, Xie X, Yang M, Dixon R, Wang YP (2017d). Modular electron-transport chains from eukaryotic organel- les function to support nitrogenase activity.Proc Natl Acad Sci USA 114, E2460-E2465. |
| 180 | Yang M, He ZW, Huang YL, Lu L, Yan YB, Hong L, Shen H, Liu Y, Guo Q, Jiang L, Zhang YW, Greenberg A, Zhou RC, Ge XJ, Wu CI, Shi SH (2017e). The emergence of the hyperinvasive vine, Mikania micrantha(Asteraceae), via admixture and founder events inferred from population transcriptomics. Mol Ecol 26, 3405-3423. |
| 181 | Yang MR, Li CX, Cai ZY, Hu YM, Nolan T, Yu FF, Yin YH, Xie Q, Tang GL, Wang XL (2017f). SINAT E3 ligases control the light-mediated stability of the brassinosteroid- activated transcription factor BES1 in Arabidopsis.Dev Cell 41, 47-58. |
| 182 | Yang N, Xu XW, Wang RR, Peng WL, Cai LC, Song JM, Li WQ, Luo X, Niu LY, Wang YB, Jin M, Chen L, Luo JY, Deng M, Wang L, Pan QC, Liu F, Jackson D, Yang XH, Chen LL, Yan JB (2017g). Contributions of Zea mays sub- species mexicana haplotypes to modern maize. Nat Com- mun 8, 1874. |
| 183 | Ye J, Wang X, Hu T, Zhang F, Wang B, Li C, Yang T, Li H, Lu Y, Giovannoni JJ, Zhang Y, Ye Z (2017a). An InDel in the promoter of Al-ACTIVATED MALATE TRANSPOR- TER9 selected during tomato domestication determines fruit malate contents and aluminum tolerance.Plant Cell 29, 2249-2268. |
| 184 | Ye YJ, Zhou LJ, Liu X, Liu H, Li DQ, Cao MJ, Chen HF, Xu L, Zhu JK, Zhao Y (2017b). A novel chemical inhibitor of ABA signaling targets all ABA receptors.Plant Physiol 173, 2356-2369. |
| 185 | Yu CW, Tai R, Wang SC, Yang P, Luo M, Yang S, Cheng K, Wang WC, Cheng YS, Wu K (2017a). HISTONE DEA- CETYLASE6 acts in concert with histone methyltransferases SUVH4, SUVH5, and SUVH6 to regulate transposon silencing.Plant Cell 29, 1970-1983. |
| 186 | Yu JP, Han JJ, Kim YJ, Song M, Yang Z, He Y, Fu RF, Luo ZJ, Hu JP, Liang WQ, Zhang DB (2017b). Two rice receptor-like kinases maintain male fertility under changing temperatures.Proc Natl Acad Sci USA 114, 12327-12332. |
| 187 | Yu LH, Wu J, Zhang ZS, Miao ZQ, Zhao PX, Wang Z, Xiang CB (2017c). Arabidopsis MADS-box transcription factor AGL21 acts as environmental surveillance of seed germination by regulating ABI5 expression.Mol Plant 10, 834-845. |
| 188 | Yuan LB, Dai YS, Xie LJ, Yu LJ, Zhou Y, Lai YX, Yang YC, Xu L, Chen QF, Xiao S (2017). Jasmonate regulates plant responses to postsubmergence reoxygenation through transcriptional activation of antioxidant synthesis.Plant Physiol 173, 1864-1880. |
| 189 | Zeng J, Dong ZC, Wu HJ, Tian ZX, Zhao Z (2017). Redox regulation of plant stem cell fate.EMBO J 36, 2844-2855. |
| 190 | Zhang A, Li N, Gong L, Gou XW, Wang B, Deng X, Li CP, Dong QL, Zhang HK, Liu B (2017a). Global analysis of gene expression in response to whole-chromosome aneu- ploidy in hexaploid wheat.Plant Physiol 175, 828-847. |
| 191 | Zhang B, Tan X, Wang S, Chen M, Chen S, Ren T, Xia J, Bai Y, Huang J, Han X (2017b). Asymmetric sensitivity of ecosystem carbon and water processes in response to precipitation change in a semi-arid steppe.Funct Ecol 31, 1301-1311. |
| 192 | Zhang B, Zhang L, Li F, Zhang D, Liu X, Wang H, Xu Z, Chu C, Zhou Y (2017c). Control of secondary cell wall patterning involves xylan deacetylation by a GDSL ester- ase.Nat Plants 3, 17017. |
| 193 | Zhang D, Li W, Xia EH, Zhang QJ, Liu Y, Zhang Y, Tong Y, Zhao Y, Niu YC, Xu JH, Gao LZ (2017d). The medicinal herb Panax notoginseng genome provides insights into ginsenoside biosynthesis and genome evolution. Mol Plant 6, 903-907. |
| 194 | Zhang D, Li YH, Zhang XY, Zha P, Lin RC (2017e). The SWI2/SNF2 chromatin-remodeling ATPase BRAHMA re- gulates chlorophyll biosynthesis in Arabidopsis.Mol Plant 10, 155-167. |
| 195 | Zhang F, Tang D, Shen Y, Xue Z, Shi W, Ren L, Du G, Li Y, Cheng Z (2017f). The F-Box protein ZYGO1 mediates bouquet formation to promote homologous pairing, synap- sis, and recombination in rice meiosis.Plant Cell 29, 2597-2609. |
| 196 | Zhang GQ, Liu KW, Li Z, Lohaus R, Hsiao YY, Niu SC, Wang JY, Lin YC, Xu Q, Chen LJ, Yoshida K, Fujiwara S, Wang ZW, Zhang YQ, Mitsuda N, Wang M, Liu GH, Peroraro L, Huang HX, Xiao XJ, Lin M, Wu XY, Wu WL, Chen YY, Chang SB, Sakamoto S, Takagi M, Yagi M, Zeng SJ, Shen CY, Yeh CM, Luo YB, Tsai WC, Van de Peer Y, Liu ZJ (2017g). The Apostasia genome and the evolution of orchids.Nature 549, 379-383. |
| 197 | Zhang H, Zhang JS, Yan J, Gou F, Mao YF, Tang GL, Botella JR, Zhu JK (2017h). Short tandem target mimic rice lines uncover functions of miRNAs in regulating important agronomic traits.Proc Natl Acad Sci USA 114, 5277-5282. |
| 198 | Zhang J, Huang QP, Zhong S, Bleckmann A, Huang JY, Guo XY, Lin Q, Gu HY, Dong J, Dresselhaus T, Qu LJ (2017i). Sperm cells are passive cargo of the pollen tube in plant fertilization.Nat Plant 3, 17079 |
| 199 | Zhang J, Ma JF, Liu DS, Qin S, Sun S, Zhao JD, Sui SF (2017j). Structure of phycobilisome from the red alga Griffithsia pacifica. Nature 551, 57-63. |
| 200 | Zhang J, Wei B, Yuan R, Wang J, Ding M, Chen Z, Yu H, Qin G (2017k). The Arabidopsis RING-Type E3 ligase TEAR1 controls leaf development by targeting the TIE1 transcriptional repressor for degradation.Plant Cell 29, 243-259. |
| 201 | Zhang JP, Yu Y, Feng YZ, Zhou YF, Zhang F, Yang YW, Lei MQ, Zhang YC, Chen YQ (2017l). MiR408 regulates grain yield and photosynthesis via a phytocyanin protein.Plant Physiol 175, 1175-1185. |
| 202 | Zhang K, Xu W, Wang C, Yi X, Zhang W, Su Z (2017m). Differential deposition of H2A.Z in combination with histone modifications within related genes in Oryza sativa callus and seedling. Plant J 89, 264-277. |
| 203 | Zhang L, Yu H, Ma B, Liu GF, Wang JJ, Wang JM, Gao RC, Li JJ, Liu JY, Xu J, Zhang YY, Li Q, Huang XH, Xu JL, Li JM, Qian Q, Han B, He ZH, Li JY (2017n). A natural tandem array alleviates epigenetic repression of IPA1 and leads to superior yielding rice.Nat Commun 8, 14789. |
| 204 | Zhang L, Zhou XM, Chen DK, Schuettpelz E, Knapp R, Lu NT, Luong TT, Dang MT, Duan YF, He H, Gao XF, Zhang LB (2017o). A global phylogeny of the fern genus Tectaria(Tectariaceae: Polypodiales) based on plastid and nuclear markers identifies major evolutionary lineages and suggests repeated evolution of free venation from anastomosing venation. Mol Phylogenet Evol 114, 295-333. |
| 205 | Zhang LJ, Li XX, Ma B, Gao Q, Du HL, Han YH, Li Y, Cao YH, Qi M, Zhu YX, Lu HW, Ma MC, Liu LL, Zhou JP, Nan CH, Qin YJ, Wang J, Cui L, Qiao ZJ (2017p). The tartary buckwheat genome provides insights into rutin biosynthesis and abiotic stress tolerance.Mol Plant 10, 1224-1237. |
| 206 | Zhang SD, Jin JJ, Chen SY, Chase M, Soltis D, Li HT, Yang JB, Li DZ, Yi TS (2017q). Diversification of Ros- aceae since the Late Cretaceous based on plastid phylogenomics.New Phytol 214, 1355-1367. |
| 207 | Zhang SD, Jin JJ, Chen SY, Chase MW, Soltis DE, Li HT, Yang JB, Li DZ, Yi TS (2017r). Diversification of Ros- aceae since the Late Cretaceous based on plastid phylogenomics.New Phytol 214, 1355-1367. |
| 208 | Zhang SS, Yang HX, Ding L, Song ZT, Ma H, Chang F, Liu JX (2017s). Tissue-specific transcriptomics reveals an important role of the unfolded protein response in main- taining fertility upon heat stress in Arabidopsis.Plant Cell 29, 1007-1023. |
| 209 | Zhang T, Li YF, Ma L, Sang XC, Ling YH, Wang YT, Yu P, Zhuang H, Huang JY, Wang N, Zhao FM, Zhang CW, Yang ZL, Fang LK, He GH (2017t). LATERAL FLORET 1 induced the three-florets spikelet in rice.Proc Natl Acad Sci USA 114, 9984-9989. |
| 210 | Zhang TQ, Lian H, Zhou CM, Xu L, Jiao YL, Wang JW (2017u). A two-step model for de novo activation of WUS- CHEL during plant shoot regeneration. Plant Cell 29, 1073-1087. |
| 211 | Zhang XX, Liu WJ, Nagae TT, Takeuchi H, Zhang HQ, Han ZF, Higashiyama T, Chai JJ (2017v). Structural basis for receptor recognition of pollen tube attraction peptides.Nat Commun 8, 1331. |
| 212 | Zhang XY, Huai JL, Shang FF, Xu G, Tang WJ, Jing YJ, Lin RC (2017w). A PIF1/PIF3-HY5-BBX23 transcription factor cascade affects photomorphogenesis.Plant Physiol 174, 2487-2500. |
| 213 | Zhang ZG, Cui X, Wang YW, Wu JX, Gu XF, Lu TG (2017x). The RNA editing factor WSP1 is essential for chloroplast development in rice.Mol Plant 10, 86-98. |
| 214 | Zhang ZS, Liu MJ, Scheibe R, Selinski J, Zhang LT, Yang C, Meng XL, Gao HY (2017y). Contribution of the alternative respiratory pathway to PSII photoprotection in C3 and C4 plants.Mol Plant 10, 131-142. |
| 215 | Zhang ZY, Li JH, Li F, Liu HH, Yang WS, Chong K, Xu YY (2017z). OsMAPK3 phosphorylates OsbHLH002/OsICE1 and inhibits its ubiquitination to activate OsTPP1 and enhances rice chilling tolerance.Dev Cell 43, 731-743. |
| 216 | Zhang ZY, Li JJ, Pan YH, Li JL, Zhou L, Shi HL, Zeng YW, Guo HF, Yang SM, Zheng WW, Yu JP, Sun XM, Li GL, Ding YL, Ma L, Shen SQ, Dai LY, Zhang HL, Yang SH, Guo Y, Li ZC (2017aa). Natural variation inCTB4a enhan- ces rice adaptation to cold habitats. Nat Commun 8, 14788. |
| 217 | Zhao C, Piao S, Wang X, Huang Y, Ciais P, Elliott J, Huang M, Janssens IA, Li T, Lian X, Liu Y, Mueller C, Peng S, Wang T, Zeng Z, Penuelas J (2017a). Plausible rice yield losses under future climate warming.Nat Plant 3, 16202. |
| 218 | Zhao CZ, Wang PC, Si T, Hsu CC, Wang L, Zayed O, Yu ZP, Zhu YF, Dong J, Tao WA, Zhu JK (2017b). MAP kinase cascades regulate the cold response by modulating ICE1 protein stability.Dev Cell 43, 618-629. |
| 219 | Zhao F, Zheng YF, Zeng T, Sun R, Yang JY, Li Y, Ren DT, Ma H, Xu ZH, Bai SN (2017c). Phosphorylation of SPO- ROCYTELESS/NOZZLE by the MPK3/6 kinase is required for anther development. Plant Physiol 173, 2265-2277. |
| 220 | Zhao GY, Zou C, Li K, Wang K, Li TB, Gao LF, Zhang XX, Wang HJ, Yang ZJ, Liu X, Jiang WK, Mao L, Kong XY, Jiao YN, Jia JZ (2017d). The Aegilops tauschii genome reveals multiple impacts of transposons. Nat Plants 3, 946-955. |
| 221 | Zhao L, Cheng DM, Huang XH, Chen M, Dall’Osto L, Xing JL, Gao LY, Li LY, Wang YL, Bassi R, Peng LW, Wang YC, Rochaix JD, Huang F (2017e). A light harvesting complex-like protein in maintenance of photosynthetic components in chlamydomonas.Plant Physiol 174, 2419-2433. |
| 222 | Zheng Y, Cui X, Su L, Fang S, Chu J, Gong Q, Yang J, Zhu Z (2017). Jasmonate inhibits COP1 activity to suppress hypocotyl elongation and promote cotyledon opening in etiolated Arabidopsis seedlings.Plant J 90, 1144-1155. |
| 223 | Zhou F, Wang CY, Gutensohn M, Jiang L, Zhang P, Zhang DB, Dudareva N, Lu S (2017a). A recruiting protein of geranylgeranyl diphosphate synthase controls metabolic flux toward chlorophyll biosynthesis in rice.Proc Natl Acad Sci USA 114, 6866-6871. |
| 224 | Zhou S, Jiang W, Long F, Cheng S, Yang W, Zhao Y, Zhou DX (2017b). Rice homeodomain protein WOX11 recruits a histone acetyltransferase complex to establish programs of cell proliferation of crown root meristem.Plant Cell 29, 1088-1104. |
| 225 | Zhou W, Barrett S, Li HD, Wu ZK, Wang XJ, Wang H, Li DZ (2017c). Phylogeographic insights on the evolutionary breakdown of heterostyly.New Phytol 214, 1368-1380. |
| 226 | Zhou W, Lu QT, Li QW, Wang L, Ding SH, Zhang AH, Wen XG, Zhang LX, Lu CM (2017d). PPR-SMR protein SOT1 has RNA endonuclease activity.Proc Natl Acad Sci USA 114, E1554-E1563. |
| 227 | Zhu J, Hu H, Tao S, Chi X, Li P, Jiang L, Ji C, Zhu J, Tang Z, Pan Y, Birdsey RA, He X, Fang J (2017a). Carbon stocks and changes of dead organic matter in China’s forests.Nat Commun 8, 151. |
| 228 | Zhu JG, Nan Q, Qin T, Qian D, Mao TL, Yuan SJ, Wu XR, Niu Y, Bai QF, An LZ, Xiang Y (2017b). Higher-ordered actin structures remodeled by Arabidopsis ACTIN-DE- POLYMERIZING FACTOR5 are important for pollen germination and pollen tube growth.Mol Plant 10, 1065-1081. |
| 229 | Zhu M, Jiang L, Bai B, Zhao W, Chen X, Li J, Liu Y, Chen Z, Wang B, Wang C, Wu Q, Shen Q, Dinesh-Kumar SP, Tao X (2017c). The intracellular immune receptor Sw-5b confers broad-spectrum resistance to spoviruses through recognition of a conserved 21-amino acid viral effector epitope.Plant Cell 29, 2214-2232. |
| 230 | Zhu X, Qi T, Yang Q, He F, Tan C, Ma W, Voegele RT, Kang Z, Guo J (2017d). Host-induced gene silencing of the MAPKK gene PsFUZ7 confers stable resistance to wheat stripe rust. Plant Physiol 175, 1853-1863. |
| 231 | Zhu Y, Wang B, Tang K, Hsu CC, Xie S, Du H, Yang Y, Tao WA, Zhu JK (2017e). An Arabidopsis nucleoporin NUP85 modulates plant responses to ABA and salt stress.PLoS Genet 13, e1007124. |
| 232 | Zhu YF, Yu YM, Cheng K, Ouyang YD, Wang J, Gong L, Zhang QH, Li XH, Xiao JH, Zhang QF (2017f). Processes underlying a reproductive barrier in indica-japonica rice hybrids revealed by transcriptome analysis.Plant Physiol 174, 1683-1696. |
| 233 | Zhu Y, Rong L, Luo Q, Wang B, Zhou N, Yang Y, Zhang C, Feng H, Zheng L, Shen WH, Ma J, Dong A (2017g). The histone chaperone NRP1 interacts with WEREWOLF to activate GLABRA2 in Arabidopsis root hair development.Plant Cell 29, 260-276. |
| 234 | Zhuang XH, Chung KP, Cui Y, Lin WL, Gao CJ, Kang BH, Jiang LW (2017). ATG9 regulates autophagosome pro- gression from the endoplasmic reticulum in Arabidopsis.Proc Natl Acad Sci USA 114, E426-E435. |
| 235 | Zou YP, Hou XH, Wu Q, Chen JF, Li ZW, Han TS, Niu XM, Yang L, Xu YC, Zhang J, Zhang FM, Tan DY, Tian ZX, Gu HY, Guo YL (2017). Adaptation ofArabidopsis thaliana to the Yangtze River basin. Genome Biol 18, 239. |
| 236 | Zuo YJ, Wen J, Zhou SL (2017). Intercontinental and in- tracontinental biogeography of the eastern Asian-Eastern North American disjunctPanax(the ginseng genus, Araliaceae), emphasizing its diversification processes in eastern Asia. Mol Phylogenet Evol 117, 60-74. |
| [1] | Peng-Bin Han Kang-Di PEI Bin He Shu-Li XIAO Cindy Q.TANG. Community composition and characteristics of Tsuga dumosa forests in China [J]. Chin J Plant Ecol, 2025, 49(植被): 1-0. |
| [2] | Bin He Peng-Bin Han Shu-Li XIAO Kang-Di PEI Cindy Q.TANG. Community types and characteristics of Keteleeria davidiana forests [J]. Chin J Plant Ecol, 2025, 49(植被): 1-0. |
| [3] | Menglu Wei, Jinyue Li, Yuchun Rao. Practice of Cultivating Innovative Talents in the Field of Plant Science Based on "Promoting Innovation through Competition" [J]. Chinese Bulletin of Botany, 2025, 60(4): 1-0. |
| [4] | Tian Zhiqi, Su Yang. China’s implementation model of international environmental conventions and its application to the Convention on Biological Diversity—Achieving the Kunming-Montreal Global Biodiversity Framework targets and the role of national parks [J]. Biodiv Sci, 2025, 33(3): 24593-. |
| [5] | Zhou Zhihua, Jin Xiaohua, Luo Ying, Li Diqiang, Yue Jianbing, Liu Fang, He Tuo, Li Xi, Dong Hui, Luo Peng. Analyses and suggestions on mechanisms of forestry and grassland administrations in China to achieve targets of Kunming-Montreal Global Biodiversity Framework [J]. Biodiv Sci, 2025, 33(3): 24487-. |
| [6] | Hongya Gu, Fan Chen, Rongcheng Lin, Xiaoquan Qi, Shuhua Yang, Zhiduan Chen, Xuewei Chen, Zhaojun Ding, Langtao Xiao, Jianru Zuo, Liwen Jiang, Yongfei Bai, Kang Chong, Lei Wang. Achievements and Advances of Plant Sciences Research in China in 2024 [J]. Chinese Bulletin of Botany, 2025, 60(2): 151-171. |
| [7] | Wei Yi, Yi Ai, Meng Wu, Liming Tian, Tserang Donko Mipam. Soil archaeal community responses to different grazing intensities in the alpine meadows of the Qinghai-Tibetan Plateau [J]. Biodiv Sci, 2025, 33(1): 24339-. |
| [8] | Bangze Li, Shuren Zhang. An updated species checklist and taxonomic synopsis of Cyperaceae in China [J]. Biodiv Sci, 2024, 32(7): 24106-. |
| [9] | Zonggang Hu. China and the United States once planned joint compilation of the Flora of China after the victory in the China’s Resistance War Against Japan [J]. Biodiv Sci, 2024, 32(6): 24220-. |
| [10] | Qiyu Kuang, Liang Hu. Species diversity and geographical distribution of marine benthic shell-bearing mollusks around Donghai Island and Naozhou Island, Guangdong Province [J]. Biodiv Sci, 2024, 32(5): 24065-. |
| [11] | Yujie Chi, Xintian Zhang, Zhixuan Tian, Chengshuai Guan, Xinzhi Gu, Zhihui Liu, Zhanbin Wang, Jinjie Wang. Species diversity of powdery mildew fungi (Erysiphaceae) and their host plants in Northeast Asia [J]. Biodiv Sci, 2024, 32(4): 23443-. |
| [12] | PAN Yuan-Fang, PAN Liang-Hao, QIU Si-Ting, QIU Guang-Long, SU Zhi-Nan, SHI Xiao-Fang, FAN Hang-Qing. Variations in tree height among mangroves and their environmental adaptive mechanisms in China’s coastal areas [J]. Chin J Plant Ecol, 2024, 48(4): 483-495. |
| [13] | Di Lin, Shuanglin Chen, Que Du, Wenlong Song, Gu Rao, Shuzhen Yan. Investigation of species diversity of myxomycetes in Dabie Mountains [J]. Biodiv Sci, 2024, 32(2): 23242-. |
| [14] | Fan Chen, Hongya Gu, Xiaoquan Qi, Rongcheng Lin, Qian Qian, Langtao Xiao, Shuhua Yang, Jianru Zuo, Yongfei Bai, Zhiduan Chen, Zhaojun Ding, Xiaojing Wang, Liwen Jiang, Kang Chong, Lei Wang. Achievements and Advances of Plant Sciences Research in China in 2023 [J]. Chinese Bulletin of Botany, 2024, 59(2): 171-187. |
| [15] | Nuriye Muhetaier, Xiuying Zhang, Subinuer Eli, Houhun Li. Annual report of new taxa for Chinese Lepidoptera in 2023 [J]. Biodiv Sci, 2024, 32(11): 24428-. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||