

Chinese Bulletin of Botany ›› 2020, Vol. 55 ›› Issue (2): 137-146.DOI: 10.11983/CBB19109
• EXPERIMENTAL COMMUNICATIONS • Previous Articles Next Articles
Yang Zhang,Huajie Liu,Ruili Xue,Haixia Li,Hua Li( )
)
Received:2019-06-14
Accepted:2019-11-05
Online:2020-03-01
Published:2020-02-12
Contact:
Hua Li
Yang Zhang,Huajie Liu,Ruili Xue,Haixia Li,Hua Li. Cloning of Wheat TaLCD Gene and Its Regulation on Osmotic Stress[J]. Chinese Bulletin of Botany, 2020, 55(2): 137-146.
| Gene name | Primer sequences | |
|---|---|---|
| Forward (5'-3') | Reverse (5'-3') | |
| TaDCD | GAGGAAGGACGGAAGCCATAT | TCAGGGTCATCGCAAACAGAG |
| TaLCD | TCCATTACGCCTACGGAGCAG | CAAGCCGGACCTTACGACCA |
| ABF2 | ATCAGAAGGGATAGGGAAGAGTAAT | TTGGTCTGCCGTGAATATCTGT |
| HAB1 | GTGTTCTCGCCATGTCTAGGTC | CTATTTCGCAGACTTCTTGGTTG |
| HAB2 | GCAGAAGTCCTTATTGCGAGTC | CTCAGAAGTTGCCACCTCCATA |
| ABI1 | TGACGGCTGTGAAGAGAGTA | CCATCTCACACGCTTCTTCA |
| ABI2 | ATTCAGACCATTCACTGACCCTC | GCTCCGTCGCCAGAACAAG |
| NCED3 | TGGCTTCTTTCACGGCAAC | ACAACAATGGCGGGAGAGTTT |
| COR47 | GAGGTTACGGATCGTGGAT | GAGCTGTTGGATCGGTGA |
| RAB18 | ATGGCGTCTTACCAGAACCGTCCA | TACCCTTGGCCACCTGATCC |
| RD29A | GTTACTGATCCCACCAAAGAAGA | GGAGACTCATCAGTCACTTCCA |
| RD22 | AGGGCTGTTTCCACTGAGG | CACCACAGATTTATCGTCAGACA |
| Actin2 | TACCCGATGGGCAAGTCA | TGCTCATACGGTCAGCGATA |
Table 1 Primers used for qRT-PCR analysis
| Gene name | Primer sequences | |
|---|---|---|
| Forward (5'-3') | Reverse (5'-3') | |
| TaDCD | GAGGAAGGACGGAAGCCATAT | TCAGGGTCATCGCAAACAGAG |
| TaLCD | TCCATTACGCCTACGGAGCAG | CAAGCCGGACCTTACGACCA |
| ABF2 | ATCAGAAGGGATAGGGAAGAGTAAT | TTGGTCTGCCGTGAATATCTGT |
| HAB1 | GTGTTCTCGCCATGTCTAGGTC | CTATTTCGCAGACTTCTTGGTTG |
| HAB2 | GCAGAAGTCCTTATTGCGAGTC | CTCAGAAGTTGCCACCTCCATA |
| ABI1 | TGACGGCTGTGAAGAGAGTA | CCATCTCACACGCTTCTTCA |
| ABI2 | ATTCAGACCATTCACTGACCCTC | GCTCCGTCGCCAGAACAAG |
| NCED3 | TGGCTTCTTTCACGGCAAC | ACAACAATGGCGGGAGAGTTT |
| COR47 | GAGGTTACGGATCGTGGAT | GAGCTGTTGGATCGGTGA |
| RAB18 | ATGGCGTCTTACCAGAACCGTCCA | TACCCTTGGCCACCTGATCC |
| RD29A | GTTACTGATCCCACCAAAGAAGA | GGAGACTCATCAGTCACTTCCA |
| RD22 | AGGGCTGTTTCCACTGAGG | CACCACAGATTTATCGTCAGACA |
| Actin2 | TACCCGATGGGCAAGTCA | TGCTCATACGGTCAGCGATA |
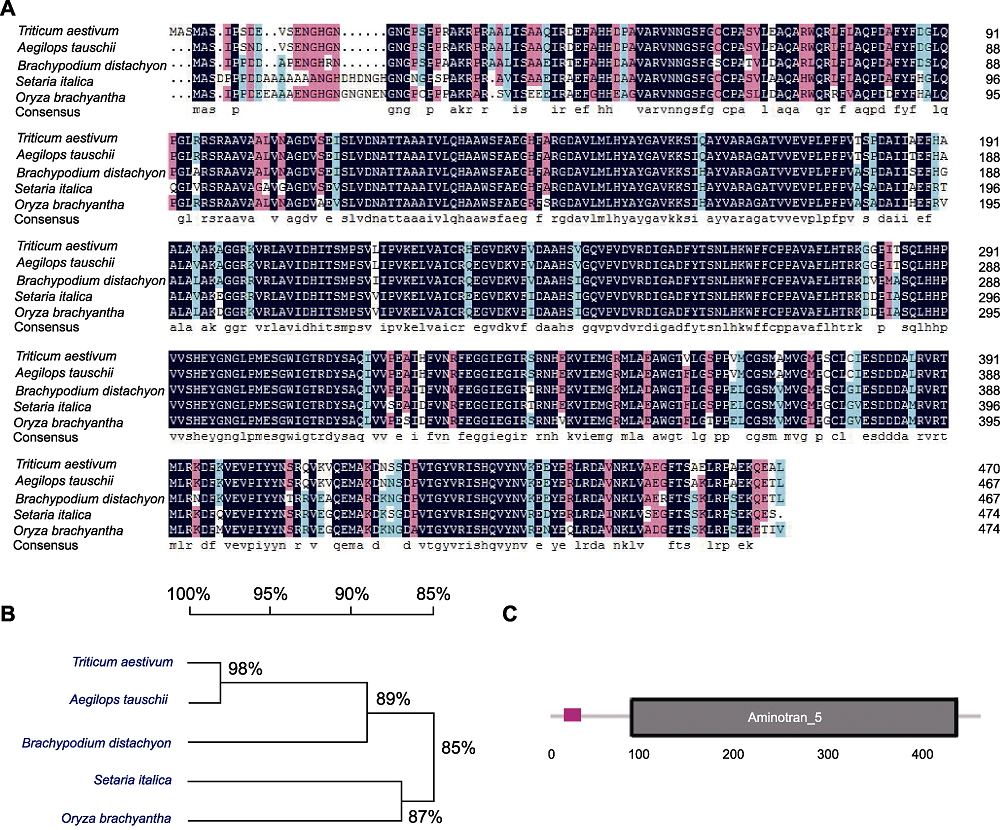
Figure 1 Sequence analysis of TaLCD (A) Comparison of the derived amino acid sequences of TaLCD in Triticum aestivum with other 4 species (Aegilops tauschii, Brachypodium distachyon, Setaria italica, and Oryza brachyantha); (B) Phylogenetic analysis of LCD in five species; (C) Schematic structures of TaLCD, black box indicates Aminotran_5 domain.
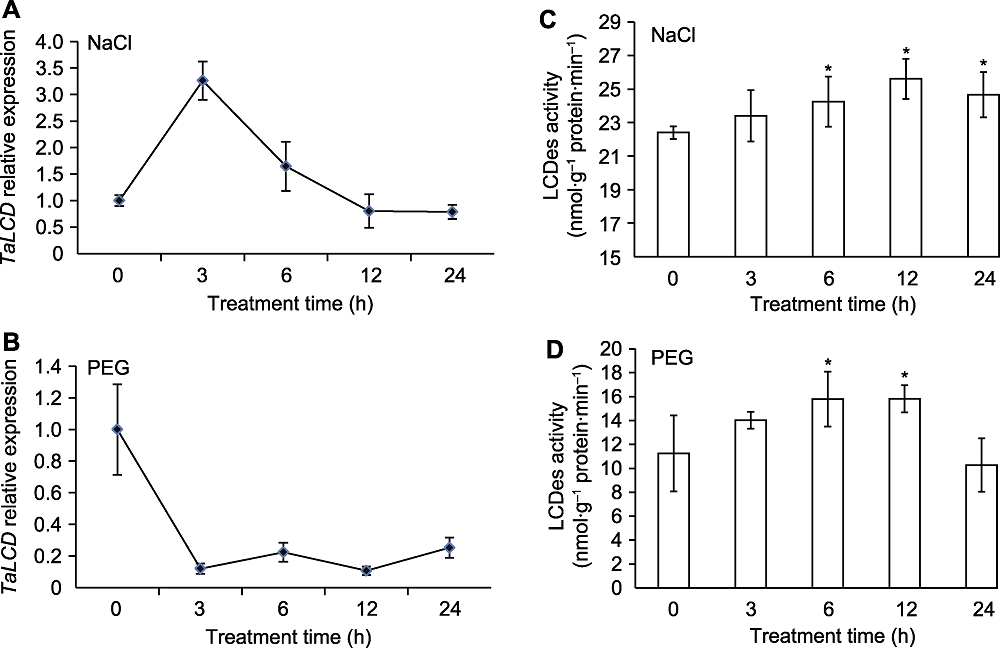
Figure 2 The expression of TaLCD and enzyme activity of TaLCD in wheat under NaCl and PEG treatments (A) The expression of TaLCD under NaCl treatment; (B) The expression of TaLCD under PEG treatment; (C) The enzyme activity of TaLCD under NaCl treatment; (D) The enzyme activity of TaLCD under PEG treatment. * indicate significant differences (P<0.05).
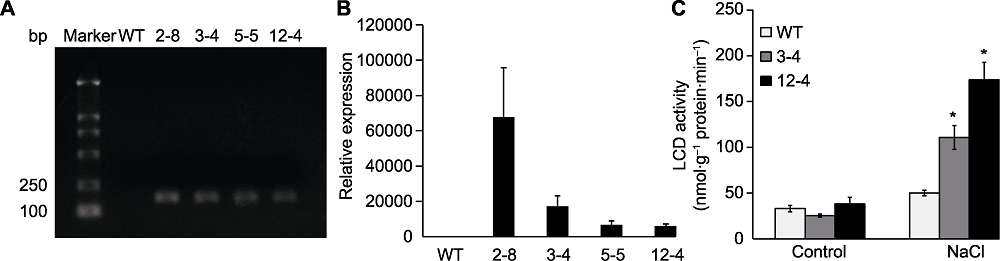
Figure 3 Expression of TaLCD and enzyme activity of TaLCD in transgenic Arabidopsis (A) RT-PCR; (B) qRT-PCR; (C) Enzyme activity of TaLCD under NaCl treatment. Marker: DNA marker; WT: Wild type; 2-8, 3-4, 5-5, and 12-4: TaLCD transgenic lines; * indicate significant differences (P<0.05).
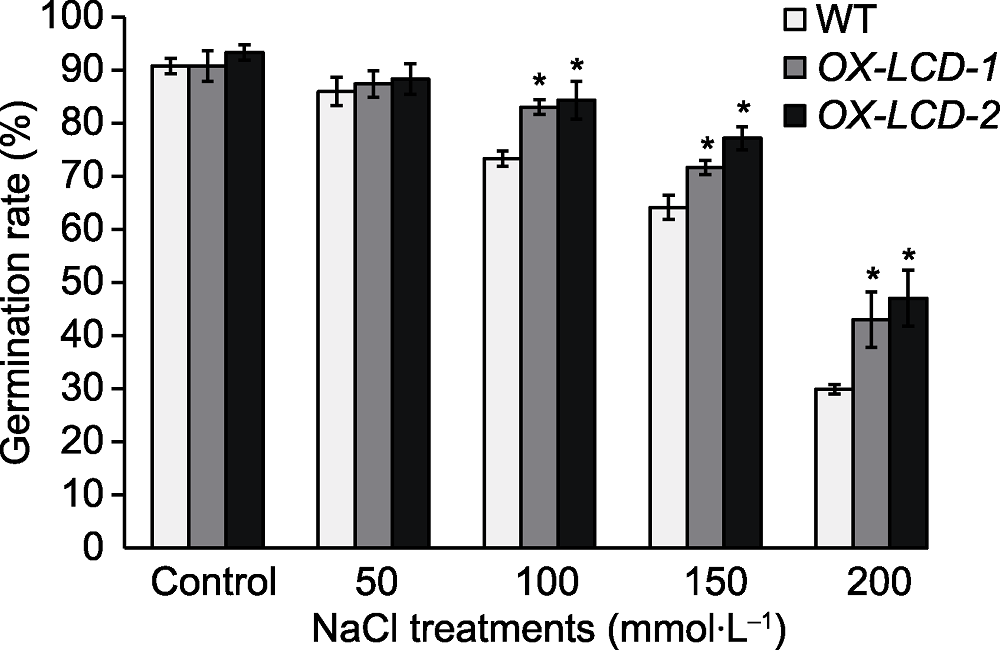
Figure 4 Seed germination rate of Arabidopsis lines expressing TaLCD under different concentrations of NaCl treatment WT: Wild type; OX-LCD-1 and OX-LCD-2: Transgenic lines; * indicate significant differences (P<0.05).
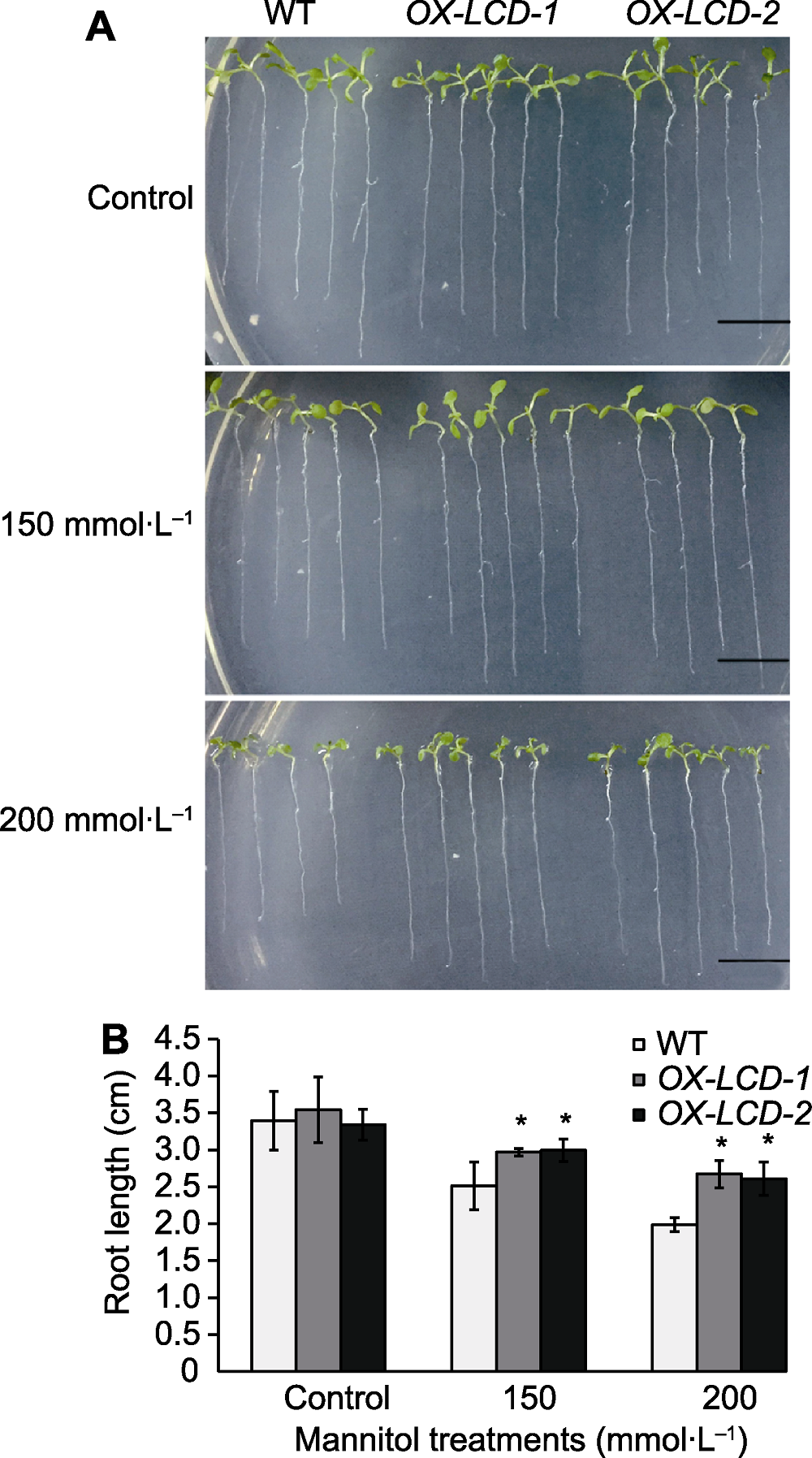
Figure 5 Root growth of Arabidopsis lines expressing TaLCD under different concentrations of mannitol treatment (A) Phenotype of root growth (Bars=1 cm); (B) Root elongation measurements. WT: Wild type; OX-LCD-1 and OX-LCD-2: Transgenic lines; * indicate significant differences (P<0.05).
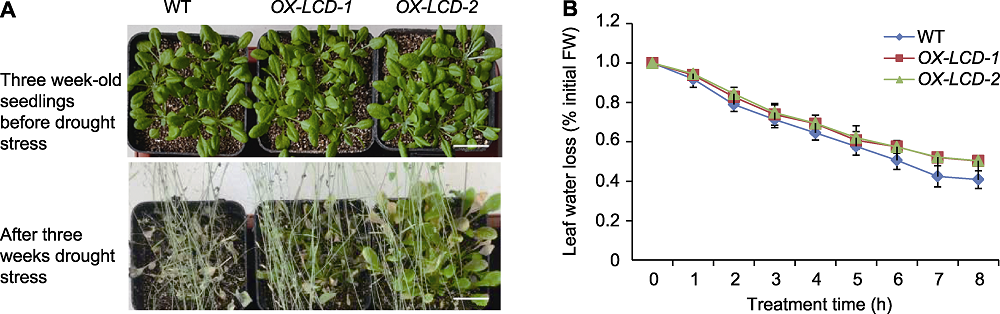
Figure 6 Drought resistance of Arabidopsis lines expressing TaLCD (A) Phenotype of three week-old wild type (WT), OX-LCD-1 and OX-LCD-2 plants after drought treatment (Bars=2 cm); (B) Leaf water loss
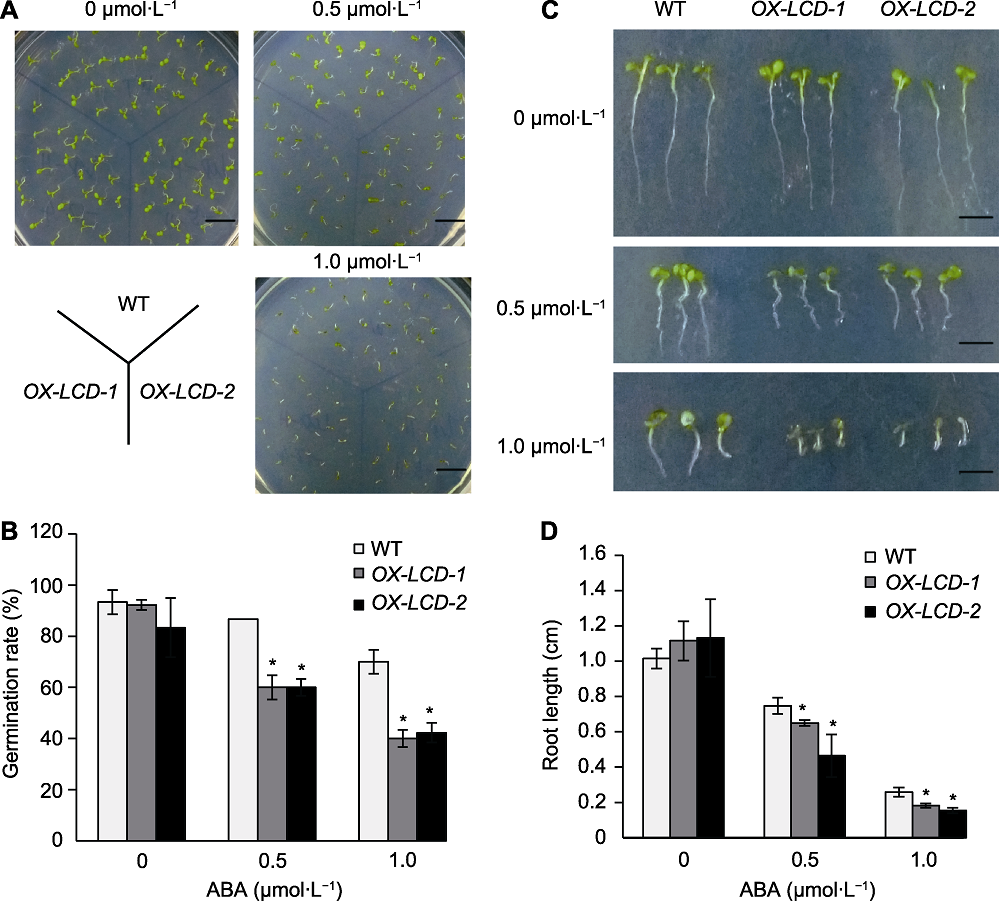
Figure 7 Seed germination and growth of Arabidopsis lines expressing TaLCD under different concentrations of ABA treatment (A) Phenotype on the 10th day (Bars=1 cm); (B) Seed germination rate after 4 d incubation; (C) Phenotype of root growth on the 10th day (Bars=0.4 cm); (D) Root length on the 10th day. WT: Wild type; ABA: Abscisic acid; * indicate significant differences (P<0.05).
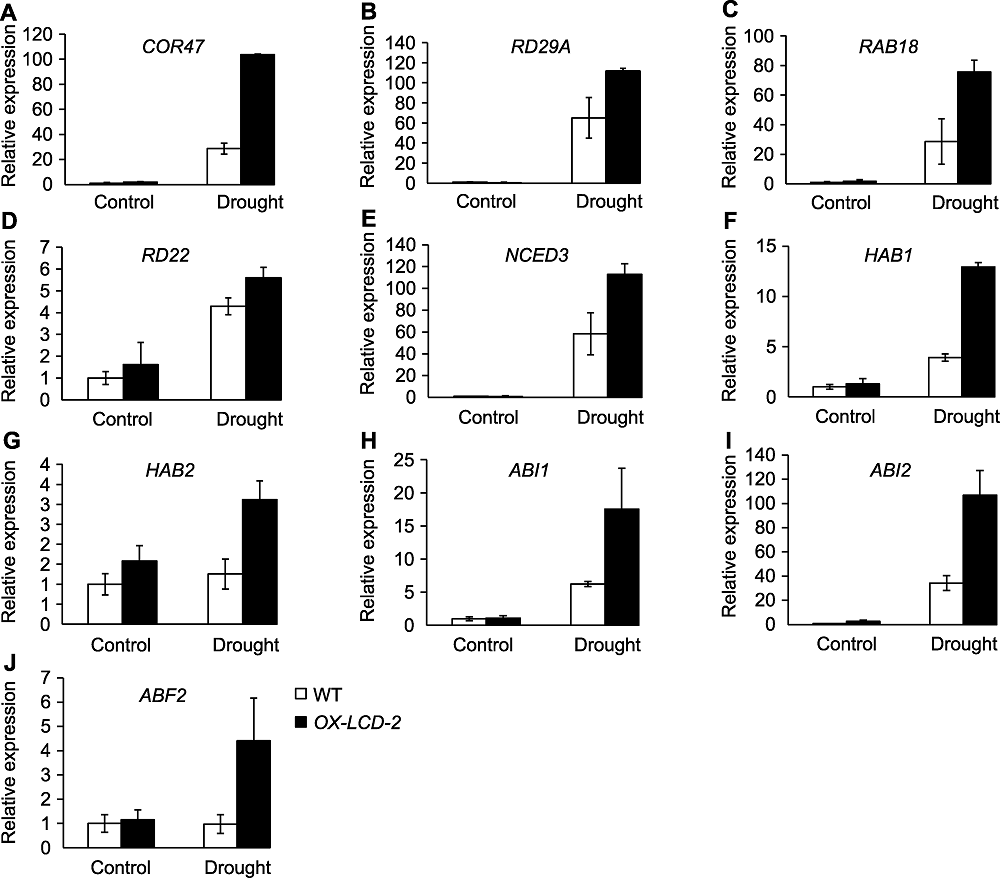
Figure 8 The expression of stress response genes and ABA signaling related genes in Arabidopsis lines expressing TaLCD under drought stress (A)-(D) Stress response genes; (E)-(J) ABA signaling related genes. WT: Wild type; OX-LCD-2: Transgenic lines
| [1] | 侯智慧, 刘菁, 侯丽霞, 李希东, 刘新 ( 2011). H2S可能作为H2O2的下游信号介导茉莉酸诱导的蚕豆气孔关闭. 植物学报 46, 396-406. |
| [2] | 刘菁, 侯智慧, 赵方贵, 刘新 ( 2011). H2S介导ABA诱导蚕豆气孔运动的生理机制研究. 西北植物学报 31, 298-304. |
| [3] | 单长卷, 周岩 ( 2011). 外源硫化氢对水分胁迫下玉米种子萌发和生长的影响. 广东农业科学 38(20), 28-30. |
| [4] | Chen J, Wang WH, Wu FH, You CY, Liu TW, Dong XJ, He JX, Zheng HL ( 2013). Hydrogen sulfide alleviates aluminum toxicity in barley seedlings. Plant Soil 362, 301-318. |
| [5] | Christou A, Manganaris GA, Papadopoulos I, Fotopoulos V ( 2013). Hydrogen sulfide induces systemic tolerance to salinity and non-ionic osmotic stress in strawberry plants through modification of reactive species biosynthesis and transcriptional regulation of multiple defence pathways. J Exp Bot 64, 1953-1966. |
| [6] | Dooley FD, Nair SP, Ward PD ( 2013). Increased growth and germination success in plants following hydrogen sulfide administration. PLoS One 8, e62048. |
| [7] | Fang HH, Liu ZQ, Long YP, Liang YL, Jin ZP, Zhang LP, Liu DM, Li H, Zhai JX, Pei YX ( 2017). The Ca 2+/CaM2- binding transcription factor TGA3 elevates LCD expression and H2S production to bolster Cr 6+ tolerance in Arabidopsis. Plant J 91, 1038-1050. |
| [8] | García-Mata C, Lamattina L ( 2010). Hydrogen sulphide, a novel gasotransmitter involved in guard cell signaling. New Phytol 188, 977-984. |
| [9] | Guo HM, Xiao TY, Zhou H, Xie YJ, Shen WB ( 2016). Hydrogen sulfide: a versatile regulator of environmental stress in plants. Acta Physiol Plant 38, 16. |
| [10] | Jia HL, Hu YF, Fan TT, Li JS ( 2015). Hydrogen sulfide modulates actin-dependent auxin transport via regulating ABPs results in changing of root development in Arabidopsis. Sci Rep 5, 8251. |
| [11] | Jin ZP, Shen JJ, Qiao ZJ, Yang GD, Wang R, Pei YX ( 2011). Hydrogen sulfide improves drought resistance in Arabidopsis thaliana. Biochem Biophys Res Commun 414, 481-486. |
| [12] | Jin ZP, Xue SW, Luo YA, Tian BH, Fang HH, Li H, Pei YX ( 2013). Hydrogen sulfide interacting with abscisic acid in stomatal regulation responses to drought stress in Arabidopsis. Plant Physiol Biochem 62, 41-46. |
| [13] | Letunic I, Doerks T, Bork P ( 2015). SMART: recent updates, new developments and status in 2015. Nucleic Acids Res 43, D257-D260. |
| [14] | Li CJ, Liu ZJ, Zhang QR, Wang RZ, Xiao LT, Ma H, Chong K, Xu YY ( 2012). SKP1 is involved in abscisic acid signaling to regulate seed germination, stomatal opening and root growth in Arabidopsis thaliana. Plant Cell Environ 35, 952-965. |
| [15] | Li H, Gao MQ, Xue RL, Wang D, Zhao HJ ( 2015). Effect of hydrogen sulfide on D1 protein in wheat under drought stress. Acta Physiol Plant 37, 225. |
| [16] | Li H, Li M, Wei XL, Zhang X, Xue RL, Zhao YD, Zhao HJ ( 2017). Transcriptome analysis of drought-responsive genes regulated by hydrogen sulfide in wheat ( Triticum aestivum L.) leaves. Mol Genet Genomics 292, 1091-1110. |
| [17] | Ma DY, Ding HN, Wang CY, Qin HX, Han QX, Hou JF, Lu HF, Xie YX, Guo TC ( 2016). Alleviation of drought stress by hydrogen sulfide is partially related to the abscisic acid signaling pathway in wheat. PLoS One 11, e0163082. |
| [18] | Papenbrock J, Riemenschneider A, Kamp A, Schulz- Vogt HN, Schmidt A ( 2007). Characterization of cysteine-degrading and H2S-releasing enzymes of higher plants—from the field to the test tube and back. Plant Biol 9, 582-588. |
| [19] | Shen JJ, Qiao ZJ, Xing TJ, Zhang LP, Liang YL, Jin ZP, Yang GD, Pei YX ( 2012). Cadmium toxicity is alleviated by AtLCD and AtDCD in Escherichia coli. J Appl Microbiol 113, 1130-1138. |
| [20] | Shi HT, Ye TT, Chan ZL ( 2013). Exogenous application of hydrogen sulfide donor sodium hydrosulfide enhanced multiple abiotic stress tolerance in bermudagrass (Cynodon dactylon(L). Pers.). Plant Physiol Biochem 71, 226-234. |
| [21] | Shi HT, Ye TT, Han N, Bian HW, Liu XD, Chan ZL ( 2015). Hydrogen sulfide regulates abiotic stress tolerance and biotic stress resistance in Arabidopsis. J Integr Plant Biol 57, 628-640. |
| [22] | Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S ( 2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28, 2731-2739. |
| [23] | Wang YQ, Li L, Cui WT, Xu S, Shen WB, Wang R ( 2012). Hydrogen sulfide enhances alfalfa ( Medicago sativa) tolerance against salinity during seed germination by nitric oxide pathway. Plant Soil 351, 107-119. |
| [24] | Xie YJ, Lai DW, Mao Y, Zhang W, Shen WB, Guan RZ ( 2013). Molecular cloning, characterization, and expression analysis of a novel gene encoding L-cysteine desulfhydrase from Brassica napus. Mol Biotechnol 54, 737-746. |
| [25] | Zhang H, Hu LY, Hu KD, He YD, Wang SH, Luo JP ( 2008). Hydrogen sulfide promotes wheat seed germination and alleviates oxidative damage against copper stress. J Integr Plant Biol 50, 1518-1529. |
| [26] | Zhang H, Hu LY, Li P, Hu KD, Jiang CX, Luo JP ( 2010a). Hydrogen sulfide alleviated chromium toxicity in wheat. Biol Plantarum 54, 743-747. |
| [27] | Zhang H, Hu SL, Zhang ZJ, Hu LY, Jiang CX, Wei ZJ, Liu J, Wang HL, Jiang ST ( 2011). Hydrogen sulfide acts as a regulator of flower senescence in plants. Postharvest Biol Technol 60, 251-257. |
| [28] | Zhang H, Jiao H, Jiang CX, Wang SH, Wei ZJ, Luo JP, Jones RL ( 2010b). Hydrogen sulfide protects soybean seedlings against drought-induced oxidative stress. Acta Physiol Plant 32, 849-857. |
| [1] | Zhiyuan Dong, Linlin Chen, Naipeng Zhang, Li Chen, Debin Sun, Yanmei Ni, Baoquan Li. Response of fish diversity to hydrological connectivity of typical tidal creek system in the Yellow River Delta based on environmental DNA metabarcoding [J]. Biodiv Sci, 2023, 31(7): 23073-. |
| [2] | Chao Xing, Yi Lin, Zhiqiang Zhou, Lianjun Zhao, Shiwei Jiang, Zhenzhen Lin, Jiliang Xu, Xiangjiang Zhan. The establishment of terrestrial vertebrate genetic resource bank and species identification based on DNA barcoding in Wanglang National Nature Reserve [J]. Biodiv Sci, 2023, 31(7): 22661-. |
| [3] | Zhenjie Zhan, Chao Zhang, Minhao Chen, Jiadong Wang, Aihua Fu, Yuwei Fan, Xiaofeng Luan. DNA metabarcoding-based winter diet analysis of Eurasian otter (Lutra lutra) in the northern Greater Khingan Mountains [J]. Biodiv Sci, 2023, 31(6): 22586-. |
| [4] | Buqing Peng, Ling Tao, Jing Li, Ronghui Fan, Shunde Chen, Changkun Fu, Qiong Wang, Keyi Tang. DNA metabarcoding dietary analysis of six sympatric small mammals at the Laojunshan National Nature Reserve, Sichuan Province [J]. Biodiv Sci, 2023, 31(4): 22474-. |
| [5] | CHEN Xue-Chun, LIU Hong, ZHU Shao-Qi, SUN Ming-Yao, YU Zhen-Rong, WANG Qing-Gang. Intraspecific variations in plant functional traits of four common herbaceous species under different abandoned years and their relevant driving factors in Lijiang River Basin, China [J]. Chin J Plant Ecol, 2023, 47(4): 559-570. |
| [6] | ZHOU Bo-Rui, LIAO Meng-Na, LI Kai, XU De-Yu, CHEN Hai-Yan, NI Jian, CAO Xian-Yong, KONG Zhao-Chen, XU Qing-Hai, ZHANG Yun, Ulrike HERZSCHUH, CAI Yong-Li, CHEN Bi-Shan, CHEN Jing-An, CHEN Ling-Kang, CHENG Bo, GAO Yang, $\boxed{\hbox{HUANG Ci-Xuan}}$ , HUANG Xiao-Zhong, LI Sheng-Feng, LI Wen-Yi, LIU Kam-Biu, LIU Guang-Xiu, LIU Ping-Mei, LIU Xing-Qi, MA Chun-Mei, SONG Chang-Qing, SUN Xiang-Jun, TANG Ling-Yu, WANG Man-Hua, WANG Yong-Bo, $\boxed{\hbox{XIA Yu-Mei}}$ , XU Jia-Sheng, YAN Shun, YANG Xiang-Dong, YAO Yi-Feng, YE Chuan-Yong, ZHANG Zhi-Yong, ZHAO Zeng-You, ZHENG Zhuo, ZHU Cheng. A fossil pollen dataset of China [J]. Chin J Plant Ecol, 2023, 47(10): 1453-1463. |
| [7] | Mei Shen, Ningning Guo, Zunlan Luo, Xiaochen Guo, Guang Sun, Nengwen Xiao. Explore the distribution and influencing factors of fish in major rivers in Beijing with eDNA metabarcoding technology [J]. Biodiv Sci, 2022, 30(7): 22240-. |
| [8] | Xiaotong Liu, Yijia Tian, Hanwen Liu, Cuiying Liang, Siwei Jiang, Wenju Liang, Xiaoke Zhang. Seasonal variation in cropland soil nematode community composition in the lower reaches of Liaohe Plain [J]. Biodiv Sci, 2022, 30(12): 22222-. |
| [9] | CHEN Hai-Yan, XU De-Yu, LIAO Meng-Na, LI Kai, NI Jian, CAO Xian-Yong, CHENG Bo, HAO Xiu-Dong, KONG Zhao-Chen, LI Sheng-Feng, LI Xiao-Qiang, LIU Guang-Xiu, LIU Ping-Mei, LIU Xing-Qi, SUN Xiang-Jun, TANG Ling-Yu, WEI Hai-Cheng, XU Qing-Hai, YAN Shun, YANG Xiang-Dong, YANG Zhen-Jing, YU Ge, ZHANG Yun, ZHANG Zhi-Yong, ZHAO Ke-Liang, ZHENG Zhuo, Ulrike HERZSCHUH. A modern pollen dataset of China [J]. Chin J Plant Ecol, 2021, 45(7): 799-808. |
| [10] | Yichao Li, Yongsheng Chen, Denis Sandanov, Ao Luo, Tong Lü, Xiangyan Su, Yunpeng Liu, Qinggang Wang, Viktor Chepinoga, Sergey Dudov, Wei Wang, Zhiheng Wang. Patterns and environmental drivers of Ranunculaceae species richness and phylogenetic diversity across eastern Eurasia [J]. Biodiv Sci, 2021, 29(5): 561-574. |
| [11] | Yawen Zhang, Shan Liang, Guoyun Xu, Wuxia Guo, Shulin Deng. Genome-wide Identification and Analysis of CONSTANS-like Gene Family in Nicotiana tabacum [J]. Chinese Bulletin of Botany, 2021, 56(1): 33-43. |
| [12] | Xuehua Liu, Yuke Zhang, Xiangyu Zhao, Xiangbo He, Qiong Cai, Yun Zhu, Baisuo He, Qiang Jiu. Introduction to the wildlife camera-trapping database of the middle Qinling Mountains [J]. Biodiv Sci, 2020, 28(9): 1075-1080. |
| [13] | Yaqiong Wan, Jiaqi Li, Xingwen Yang, Sheng Li, Haigen Xu. Progress of the China mammal diversity observation network (China BON-Mammal) based on camera-trapping [J]. Biodiv Sci, 2020, 28(9): 1115-1124. |
| [14] | Zhanhui Xu, Shiyao Liu, Ying Zhao, Wenqin Tu, Zhaofeng Chang, Entao Zhang, Jing Guo, Di Zheng, Jun Geng, Gaoying Gu, Chunpeng Guo, Lulu Guo, Jing Wang, Chunyang Xu, Chuan Peng, Teng Yang, Mengqi Cui, Weicheng Sun, Jiantan Zhang, Haotian Liu, Chaoqun Ba, Heqi Wang, Jingchao Jia, Jinzhou Wu, Cui Xiao, Keping Ma. Evaluation of the identification ability of eight commonly used plant identification application softwares in China [J]. Biodiv Sci, 2020, 28(4): 524-533. |
| [15] | Ke Wang,Mingjun Zhao,Jinhe Su,Liu Yang,Hong Deng,Yonghui Wang,Haijun Wu,Yi Li,Hongmei Wu,Xiaodan Wei,Tiezheng Wei,Lei Cai,Yijian Yao. The use of Checklist of Fungi in China database in the red list assessment of macrofungi in China [J]. Biodiv Sci, 2020, 28(1): 74-98. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||