

Chinese Bulletin of Botany ›› 2017, Vol. 52 ›› Issue (2): 179-187.DOI: 10.11983/CBB16041
Previous Articles Next Articles
Wang Qian, Sun Wenjing, Bao Ying*( )
)
Received:2016-03-08
Accepted:2016-08-08
Online:2017-03-01
Published:2017-04-05
Contact:
Bao Ying
About author:# Co-first authors
Wang Qian, Sun Wenjing, Bao Ying. Evolutionary Pattern of the GBSS Gene Family in Plants[J]. Chinese Bulletin of Botany, 2017, 52(2): 179-187.
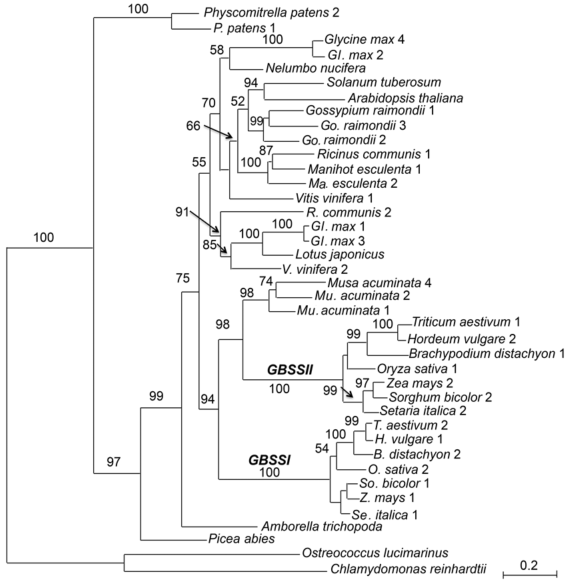
Figure 1 The maximum likelihood tree of the granule-bound starch synthase gene family based on amino acid sequences of 22 plant speciesNumbers above the branches indicate bootstrap values above 50%. Numbers following species names indicate different gene loci as listed in Table 1.
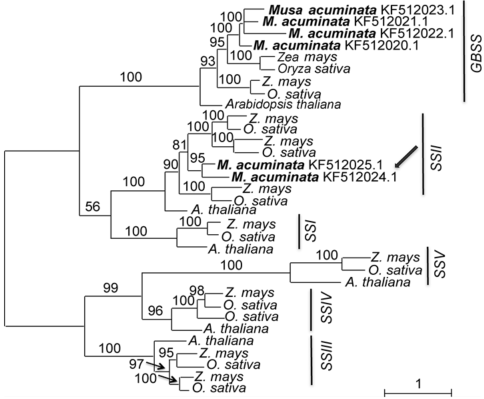
Figure 2 Maximum likelihood tree of 4 species based on amino acid homologous sequences of the starch synthase genesNumbers above the branches indicate bootstrap values above 50%. Two genes of Musa acuminata indicated by arrow should be SSII rather than GBSS.
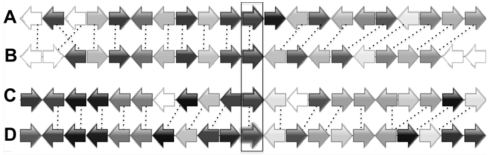
Figure 3 Synteny alignment of the chromosome regions with 4 GBSS homologous genes in Glycine max(A) The chromosome region includes locus GM10G31540; (B) The chromosome region includes locus GM20G36040; (C) The chromosome region includes locus GM16G02110; (D) The chromosome region includes locus GM07G05580. Arrows indicate the direction of genes’ transcription. Homologous genes are shown in same colors. Four GBSS genes are in the frame.
| [1] | 包颖, 杜家潇, 景翔, 徐思 (2015). 药用野生稻叶中淀粉合成酶基因家族的序列分化和特异表达. 植物学报50, 683-690. |
| [2] | Ahuja G, Jaiswal S, Hucl P, Chibbar RN (2014). Wheat genome specific granule-bound starch synthase I differentially influence grain starch synthesis.Carbohydr Polym 114, 87-94. |
| [3] | Alison MS (2012). Starch in the Arabidopsis plant.Starch 64, 421-434. |
| [4] | Ball S, Colleoni C, Cenci U, Raj JN, Tirtiaux C (2011). The evolution of glycogen and starch metabolism in eukaryotes gives molecular clues to understand the establishment of plastid endosymbiosis.J Exp Bot 62, 1775-1801. |
| [5] | Baranov Iu O, Slishchuk HI, Volkova NE, SyvolapIu M (2014). Bioinformatic analysis of maize granule-bound starch synthase gene.Tsitol Genet 48, 18-23. |
| [6] | Criscuolo A (2011). morePhyML: improving the phylogenetic tree space exploration with PhyML 3.Mol Phylogenet Evol 61, 944-948. |
| [7] | Deschamps P, Moreau H, Worden AZ, Dauvillee D, Ball SG (2008). Early gene duplication within chloroplastida and its correspondence with relocation of starch metabolism to chloroplasts.Genetics 178, 2373-2387. |
| [8] | Dian W, Jiang H, Chen Q, Liu F, Wu P (2003). Cloning and characterization of the granule-bound starch synthase II gene in rice: gene expression is regulated by the nitrogen level, sugar and circadian rhythm.Planta 218, 261-268. |
| [9] | Fasahat P, Rahman S, Ratnam W (2014). Genetic controls on starch amylose content in wheat and rice grains.J Genet 93, 279-292. |
| [10] | Fulton DC, Edwards A, Pilling E, Robinson HL, Fahy B, Seale R, Kato L, Donald AM, Geigenberger P, Martin C, Smith AM (2002). Role of granule-bound starch synthase in determination of amylopectin structure and starch granule morphology in potato.J Biol Chem 277, 10834-10841. |
| [11] | Guzman C, Alvarez JB (2015). Wheat waxy proteins: polymorphism, molecular characterization and effects on starch properties.Theor Appl Genet 9, 1049-1060. |
| [12] | Hirose T, Hashida Y, Aoki N, Okamura M, Yonekura M, Ohto C, Terao T, Ohsugi R (2014). Analysis of gene- disruption mutants of a sucrose phosphate synthase gene in rice,OsSPS1, shows the importance of sucrose synthesis in pollen germination. Plant Sci 225, 102-106. |
| [13] | Hoai TT, Matsusaka H, Toyosawa Y, Suu TD, Satoh H, Kumamaru T (2014). Influence of single-nucleotide poly- morphisms in the gene encoding granule-bound starch synthase I on amylose content in Vietnamese rice cultivars.Breed Sci 64, 142-148. |
| [14] | Jeon JS, Ryoo N, Hahn TR, Walia H, Nakamura Y (2010). Starch biosynthesis in cereal endosperm.Plant Physiol Biochem 48, 383-392. |
| [15] | Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007). Clustal W and Clustal X version 2.0.Bioinformatics 23, 2947-2948. |
| [16] | Miao H, Sun P, Liu W, Xu B, Jin Z (2014). Identification of genes encoding granule-bound starch synthase involved in amylose metabolism in banana fruit.PLoS One 9, e88077. |
| [17] | Ohdan T, Francisco PB, Sawada TJ, Hirose T, Terao T, Satoh H, Nakamura Y (2005). Expression profiling of genes involved in starch synthesis in sink and source organs of rice.J Exp Bot 56, 3229-3244. |
| [18] | Orzechowski S (2008). Starch metabolism in leaves.Acta Biochim Pol 55, 435-445. |
| [19] | Patron NJ, Smith AM, Fahy BF, Hylton CM, Naldrett MJ, Rossnagel BG, Denyer K (2002). The altered pattern of amylose accumulation in the endosperm of low-amylose barley cultivars is attributable to a single mutant allele of granule-bound starch synthase I with a deletion in the 5'-non-coding region.Plant Physiol 130, 190-198. |
| [20] | Tsai CY (1974). The function of the waxy locus in starch synthesis in maize endosperm.Biochem Genet 11, 83-96. |
| [21] | Vrinten PL, Nakamura T (2000). Wheat granule-bound starch synthase I and II are encoded by separate genes that are expressed in different tissues.Plant Physiol 122, 255-264. |
| [22] | Waterhouse AM, Procter JB, Martin DM, Clamp M, Barton GJ (2009). Jalview Version 2—a multiple sequence alignment editor and analysis workbench.Bioinformatics 25, 1189-1191. |
| [23] | Yan HB, Pan XX, Jiang HW, Wu GJ (2009). Comparison of the starch synthesis genes between maize and rice: copies, chromosome location and expression divergence.Theor Appl Genet 119, 815-825. |
| [24] | Zhu L, Gu M, Meng X, Cheung SC, Yu H, Huang J, Sun Y, Shi Y, Liu Q (2012). High-amylose rice improves indices of animal health in normal and diabetic rats.Plant Biotechnol J 10, 353-362. |
| [1] | Zhao-Quan CHEN Ming-Hui WANG Zi-Han Hu xuedong Lang Yun-Qiong HE LIU Wan-De. Community assembly mechanism of seedling in a monsoon evergreen broad-leaved forest in Pu’er, Yunnan Province, China [J]. Chin J Plant Ecol, 2024, 48(预发表): 0-0. |
| [2] | Congcong Liu Nianpeng He Ying LI Jiahui Zhang Pu Yan Ruomeng Wang Wang RuiLi. Current and future trends of plant functional traits in macro-ecology [J]. Chin J Plant Ecol, 2024, 48(预发表): 0-0. |
| [3] | Jialong Ren, Yongzhen Wang, Yilin Feng, Qihan Yan, chang Qin, Jing Fang, Weidong Xin, Jiliang Liu. Beetle data set collected using pitfall trapping in the Gobi Desert of the Hexi Corridor [J]. Biodiv Sci, 2024, 32(2): 23375-. |
| [4] | Hai Ren, Tuo He, Shifeng Wen, Hui Dong. The main features of the world-class national botanical garden with Chinese characteristics [J]. Biodiv Sci, 2023, 31(9): 23192-. |
| [5] | YUAN Ya-Ni, ZHOU Zhe, CHEN Bin-Zhou, GUO Yao-Xin, YUE Ming. Differential ecological strategies in functional traits among coexisting tree species in a Quercus aliena var. acuteserrata forest [J]. Chin J Plant Ecol, 2023, 47(9): 1270-1277. |
| [6] | ZHAO Yan-Chao, CHEN Li-Tong. Soil nutrients modulate response of aboveground biomass to warming in alpine grassland on the Qingzang Plateau [J]. Chin J Plant Ecol, 2023, 47(8): 1071-1081. |
| [7] | SUN Jia-Hui, SHI Hai-Lan, CHEN Ke-Yu, JI Bao-Ming, ZHANG Jing. Research advances on trade-off relationships of plant fine root functional traits [J]. Chin J Plant Ecol, 2023, 47(8): 1055-1070. |
| [8] | ZHAO Meng-Juan, JIN Guang-Ze, LIU Zhi-Li. Vertical variations in leaf functional traits of three typical ferns in mixed broadleaved- Korean pine forest [J]. Chin J Plant Ecol, 2023, 47(8): 1131-1143. |
| [9] | Yasu Cao, Min Fan, Yu Peng, Jiaxun Xin, Nanyi Peng. Effects of landscape pattern dynamics on plant species and functional diversity in Hunshandak Sandland [J]. Biodiv Sci, 2023, 31(8): 23048-. |
| [10] | Chunling Wu, Zhuhui Luo, Yide Li, Han Xu, Dexiang Chen, Qiong Ding. Foliar endophytic bacterial communities of woody Fabaceae and Lauraceae plants in tropical mountain rainforests: Understanding species and functional diversity and their driving factors [J]. Biodiv Sci, 2023, 31(8): 23146-. |
| [11] | Shengwen Chen, Haibao Ren, Guangrong Tong, Ningning Wang, Wenchao Lan, Jianhua Xue, Xiangcheng Mi. Spatial patterns in woody species diversity in the Qianjiangyuan National Park [J]. Biodiv Sci, 2023, 31(7): 22587-. |
| [12] | Fayang Li, Yingyu Li, Wenni Jiang, Shuguang Liu, Chao Huo, Qiaoqi Sun, Hongfei Zou. How forest fires affect bird diversity over time in boreal forest interiors and edges in the Greater Khingan Mountains [J]. Biodiv Sci, 2023, 31(7): 22665-. |
| [13] | DAI Jing-Zhong, BAI Yu-Ting, WEI Zhi-Jun, ZHANG Chu, XIN Xiao-Ping, YAN Yu-Chun, YAN Rui-Rui. Dynamic response of functional traits to fertilization in Leymus chinensis [J]. Chin J Plant Ecol, 2023, 47(7): 943-953. |
| [14] | LÜ Zi-Li, LIU Bin, CHANG Feng, MA Zi-Jing, CAO Qiu-Mei. Relationship between plant functional diversity and ecosystem multifunctionality in Bayanbulak alpine meadow along an altitude gradient [J]. Chin J Plant Ecol, 2023, 47(6): 822-832. |
| [15] | Xiting Yu, Xuehui Huang. New Insights Into the Origin of Modern Maize-hybridization of Two Teosintes [J]. Chinese Bulletin of Botany, 2023, 58(6): 857-860. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||