

Chinese Bulletin of Botany ›› 2020, Vol. 55 ›› Issue (2): 177-181.DOI: 10.11983/CBB19238
• INVITED PROTOCOL • Previous Articles Next Articles
Yingjun Yu1,2,Hang Xu1,2,Lei Wang1,2,*( )
)
Received:2019-12-13
Accepted:2020-01-23
Online:2020-03-01
Published:2020-02-12
Contact:
Lei Wang
Yingjun Yu,Hang Xu,Lei Wang. A Non-invasive Method for Measuring and Analyzing Circadian Phenotype in Living Plants[J]. Chinese Bulletin of Botany, 2020, 55(2): 177-181.
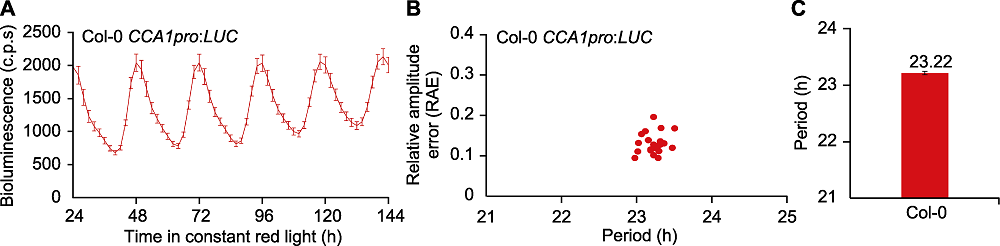
Figure 1 Circadian phenotype analysis of Col-0 wild type Arabidopsis thaliana CCA1pro:LUC under continuous red light (A) Trace plot of bioluminescence in Col-0 wild type Arabidopsis thaliana with CCA1pro:LUC reporter under continuous red light conditions, from LL24 to LL144 (the abbreviation of c.p.s represents the counts of photons per seedling). (B) Scatter plot of circadian period (x-axis) and relative amplitude error (RAE) (y-axis) of Col-0; According to the results, the periods are more concentrated and relatively consistent, and the relative amplitude errors are all lower than 0.5, which indicates that the result is reliable (generally, when the RAE is greater than 0.5, the circadian oscillation intensity is relatively low, which is not suitable for calculating the period). (C) Estimated circadian period of CCA1pro:LUC in Col-0; According to the calculation, circadian period length of Col-0 is 23.22 hours under continuous red light. Calculation was performed using FFT-NLLS method. Data are presented as means±SE (n=21).
| [1] | 魏华, 王岩, 刘宝辉, 王雷 ( 2018). 植物生物钟及其调控生长发育的研究进展. 植物学报 53, 456-467. |
| [2] | 谢启光, 徐小冬 ( 2015). 植物生物钟与关键农艺性状调控. 生命科学 27, 1336-1344. |
| [3] | Agosti RD, Jouve L, Greppin H ( 1997). Computer-assisted measurements of plant growth with linear variable differential transformer (LVDT) sensors. Arch Sci 50, 233-244. |
| [4] | Alabadi D, Oyama T, Yanovsky MJ, Harmon FG, Más P, Kay SA ( 2001). Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880-883. |
| [5] | Covington MF, Harmer SL ( 2007). The circadian clock regulates auxin signaling and responses in Arabidopsis. PLoS Biol 5, e222. |
| [6] | Dodd AN, Salathia N, Hall A, Kévei E, Tóth R, Nagy F, Hibberd JM, Millar AJ, Webb AAR ( 2005). Plant circadian clocks increase photosynthesis, growth, survival, and competitive advantage. Science 309, 630-633. |
| [7] | Dowson-Day MJ, Millar AJ ( 1999). Circadian dysfunction causes aberrant hypocotyl elongation patterns in Arabidopsis. Plant J 17, 63-71. |
| [8] | Doyle MR, Davis SJ, Bastow RM, McWatters HG, Kozma-Bognár L, Nagy F, Millar AJ, Amasino RM ( 2002). The ELF4 gene controls circadian rhythms and flowering time in Arabidopsis thaliana. Nature 419, 74-77. |
| [9] | Edwards KD, Millar AJ ( 2007). Analysis of circadian leaf movement rhythms in Arabidopsis thaliana. Methods Mol Biol 362, 103-113. |
| [10] | Kim J, Somers DE ( 2010). Rapid assessment of gene function in the circadian clock using artificial microRNA in Arabidopsis mesophyll protoplasts. Plant Physiol 154, 611-621. |
| [11] | Li B, Wang Y, Zhang YY, Tian WW, Chong K, Jang JC, Wang L ( 2019). PRR5, 7 and 9 positively modulate TOR signaling-mediated root cell proliferation by repressing TANDEM ZINC FINGER 1 in Arabidopsis. Nucleic Acids Res 47, 5001-5015. |
| [12] | Más P, Alabadi D, Yanovsky MJ, Oyama T, Kay SA ( 2003). Dual role of TOC1 in the control of circadian and photomorphogenic responses in Arabidopsis. Plant Cell 15, 223-236. |
| [13] | McClung CR ( 2006). Plant circadian rhythms. Plant Cell 18, 792-803. |
| [14] | Millar AJ, Carre IA, Strayer CA, Chua NH, Kay SA ( 1995). Circadian clock mutants in Arabidopsis identified by luciferase imaging. Science 267, 1161-1163. |
| [15] | Müller LM, von Korff M, Davis SJ ( 2014). Connections bet- ween circadian clocks and carbon metabolism reveal species-specific effects on growth control. J Exp Bot 65, 2915-2923. |
| [16] | Muller NA, Jimenez-Gomez JM ( 2016). Analysis of circadian leaf movements. In: Duque P, ed. Environmental Responses in Plants. Methods in Molecular Biology, Vol. 1398. New York: Humana Press. pp. 71-79. |
| [17] | Nakamichi N, Ito S, Oyama T, Yamashino T, Kondo T, Mizuno T ( 2004). Characterization of plant circadian rhythms by employing Arabidopsis cultured cells with bioluminescence reporters. Plant Cell Physiol 45, 57-67. |
| [18] | Niwa Y, Yamashino T, Mizuno T ( 2009). The circadian clock regulates the photoperiodic response of hypocotyl elongation through a coincidence mechanism in Arabidopsis thaliana. Plant Cell Physiol 50, 838-854. |
| [19] | Ronald J, Davis SJ ( 2017). Making the clock tick: the transcriptional landscape of the plant circadian clock. F1000Res 6, 951. |
| [20] | Salomé PA, Michael TP, Kearns EV, Fett-Neto AG, Sharrock RA, McClung CR ( 2002). The out of phase 1 mutant defines a role for PHYB in circadian phase control in Arabidopsis. Plant Physiol 129, 1674-1685. |
| [21] | Sanchez A, Shin J, Davis SJ ( 2011). Abiotic stress and the plant circadian clock. Plant Signal Behav 6, 223-231. |
| [22] | Somers DE, Webb AA, Pearson M, Kay SA ( 1998). The short-period mutant, toc1-1, alters circadian clock regulation of multiple outputs throughout development in Arabidopsis thaliana. Development 125, 485-494. |
| [23] | Uehlein N, Kaldenhoff R ( 2008). Aquaporins and plant leaf movements. Ann Bot 101, 1-4. |
| [24] | Wang L, Kim J, Somers DE ( 2013). Transcriptional corepressor TOPLESS complexes with pseudoresponse regulator proteins and histone deacetylases to regulate circadian transcription. Proc Natl Acad Sci USA 110, 761-766. |
| [25] | Yanovsky MJ, Izaguirre M, Wagmaister JA, Gatz C, Jackson SD, Thomas B, Casal JJ ( 2000). Phytochrome A resets the circadian clock and delays tuber formation under long days in potato. Plant J 23, 223-232. |
| [26] | Zhang YY, Bo CP, Wang L ( 2019). Novel crosstalks between circadian clock and jasmonic acid pathway finely coordinate the tradeoff among plant growth, senescence and defense. Int J Mol Sci 20, 5254. |
| [27] | Zhang YY, Wang Y, Wei H, Li N, Tian WW, Chong K, Wang L ( 2018). Circadian evening complex represses jasmonate-induced leaf senescence in Arabidopsis. Mol Plant 11, 326-337. |
| [28] | Zielinski T, Moore AM, Troup E, Halliday KJ, Millar AJ ( 2014). Strengths and limitations of period estimation methods for circadian data. PLoS One 9, e96462. |
| [1] | Zhang Yingchuan, Wu Xiaomingyu, Tao Baolong, Chen Li, Lu Haiqin, Zhao Lun, Wen Jing, Yi Bin, Tu Jinxing, Fu Tingdong, Shen Jinxiong. Bna-miR43 Mediates the Response of Drought Tolerance in Brassica napus [J]. Chinese Bulletin of Botany, 2023, 58(5): 701-711. |
| [2] | Shuyao Zhou, Jianming Li, Juan Mao. AtGH3.17-mediated Regulation of Auxin and Brassinosteroid Response in Arabidopsis thaliana [J]. Chinese Bulletin of Botany, 2023, 58(3): 373-384. |
| [3] | Nan Wu, Lei Qin, Kan Cui, Haiou Li, Zhongsong Liu, Shitou Xia. Cloning of Brassica napus EXA1 Gene and Its Regulation on Plant Disease Resistance [J]. Chinese Bulletin of Botany, 2023, 58(3): 385-393. |
| [4] | Gang Wang, Ertao Wang. The Broad-spectrum Innate Resistance Against Clubroot Disease Conferred by WeiTsing is Mechanistically Revealed [J]. Chinese Bulletin of Botany, 2023, 58(3): 356-358. |
| [5] | Ming-Yi Bai, Jinrong Peng, Xiangdong Fu. Coordinated Regulation of Gibberellin and Brassinosteroid Signalings Drives Toward a Sustainable “Green Revolution” by Breeding the New Generation of High-yield Wheat [J]. Chinese Bulletin of Botany, 2023, 58(2): 194-198. |
| [6] | Shanwu Lü, Changwei Zhang, Xilin Hou, Shulin Deng. Research Progress of TuMV Resistance for Brassica rapa [J]. Chinese Bulletin of Botany, 2023, 58(2): 335-346. |
| [7] | Rui Luo, Ya Chen, Hanma Zhang. Research progress on whole-genome resequencing in Brassica [J]. Biodiv Sci, 2023, 31(10): 23237-. |
| [8] | Wenjing Zhang, Xiaomeng Yang, Kan Gao, Xinyi Wei, Xuetong Shi, Ruixuan Wang, Fengxia Wu, Juqing Kang. Analysis on the Evolution and Transcription Activation Activity of ABI4 in Brassicaceae [J]. Chinese Bulletin of Botany, 2021, 56(6): 676-686. |
| [9] | Min Song,Yao Zhang,Liying Wang,Xiangyong Peng. Genome-wide Identification and Phylogenetic Analysis of Zinc Finger Homeodomain Family Genes in Brassica napus [J]. Chinese Bulletin of Botany, 2019, 54(6): 699-710. |
| [10] | Lulu Li,Wenchao Yin,Mei Niu,Wenjing Meng,Xiaoxing Zhang,Hongning Tong. Functional Analysis of Brassinosteroids in Salt Stress Responses in Rice [J]. Chinese Bulletin of Botany, 2019, 54(2): 185-193. |
| [11] | Haiwei Shuai, Yongjie Meng, Feng Chen, Wenguan Zhou, Xiaofeng Luo, Wenyu Yang, Kai Shu. Phytohormone-mediated Plant Shade Responses [J]. Chinese Bulletin of Botany, 2018, 53(1): 139-148. |
| [12] | Gao Huhu, Zhang Yunxiao, Hu Shengwu, Guo Yuan. Genome-wide Survey and Phylogenetic Analysis of MADS-box Gene Family in Brassica napus [J]. Chinese Bulletin of Botany, 2017, 52(6): 699-712. |
| [13] | Liu Kaige, Qi Shuanghui, Duan Shaowei, Li Dong, Jin Changyu, Gao Chenhao, Liu Mingxun Chen Xuanxia. Functional Analysis of Brassica napus BnTTG1-1 Gene [J]. Chinese Bulletin of Botany, 2017, 52(6): 713-722. |
| [14] | Jiangmin Xu, Hongzhen Jiang, Han Lin, Miaomiao Huang, Qiaoli Fu, Dali Zeng, Yuchun Rao. EARLY SENESCENCE 1 Participates in the Expression Regulation of Circadian Clock Genes and Response to Stress in Rice [J]. Chinese Bulletin of Botany, 2016, 51(6): 743-756. |
| [15] | Jia Ledong, Li Shimeng, Xu Daixiang, Qu Cunmin, Li Jiana, Wang Rui. Bioinformatics Analysis of BnMYB80 Genes in Brassica napus [J]. Chinese Bulletin of Botany, 2016, 51(5): 620-630. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||