

Chinese Bulletin of Botany ›› 2019, Vol. 54 ›› Issue (4): 503-508.DOI: 10.11983/CBB19087
• TECHNIQUES AND METHODS • Previous Articles Next Articles
Xinjie Cheng1,Hengxiu Yu1,Zhukuan Cheng2,*( )
)
Received:2019-05-12
Accepted:2019-06-14
Online:2019-07-10
Published:2019-07-01
Contact:
Zhukuan Cheng
Xinjie Cheng,Hengxiu Yu,Zhukuan Cheng. Protocols for Analyzing Rice Meiotic Chromosomes[J]. Chinese Bulletin of Botany, 2019, 54(4): 503-508.
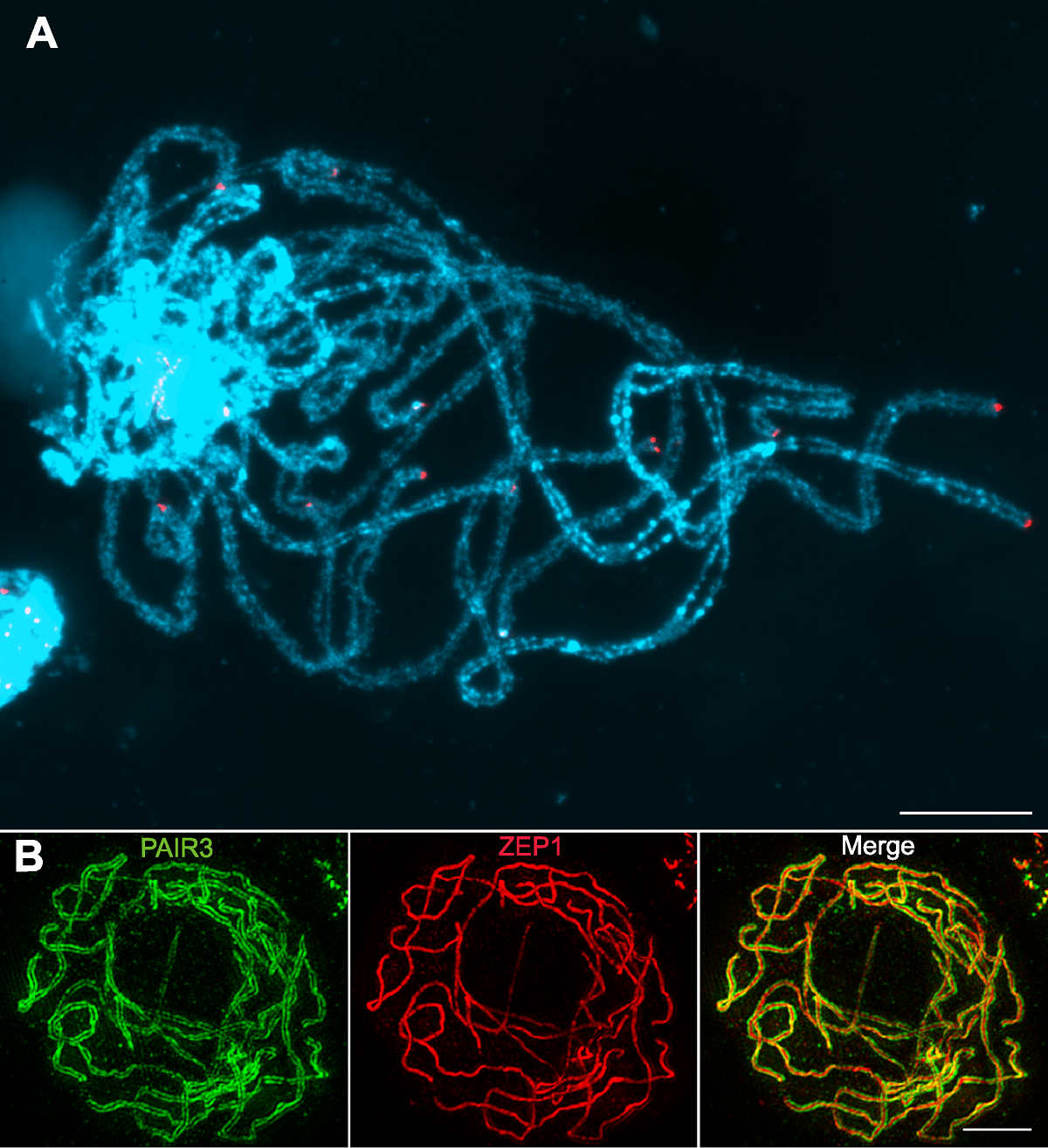
Figure 1 Characterization of rice meiotic pachytene chromosomes (A) Fluorescent in situ hybridization using rice pachytene chromosomes probed with the telomeric sequence pAtT4 (red); Chromosomes stained with DAPI (blue) (Images taken with a regular fluorescence microscope); (B) Immunodetection of PAIR3 (green) and ZEP1 (red) in rice pachytene chromosomes (Images taken with a super resolution microscope). Bars=5 µm
| [1] | Arumuganathan K, Earle ED ( 1991). Estimation of nuclear DNA content of plants by flow cytometry. Plant Mol Biol Rep 9, 229-241. |
| [2] | Che L, Tang D, Wang K, Wang M, Zhu K, Yu H, Gu M, Cheng Z ( 2011). OsAM1 is required for leptotene transition in rice. Cell Res 21, 654-665. |
| [3] | Cheng Z, Buell CR, Wing RA, Gu M, Jiang J ( 2001 a). Toward a cytological characterization of the rice genome. Genome Res 11, 2133-2141. |
| [4] | Cheng Z, Dong F, Langdon T, Ouyang S, Buell CR, Gu M, Blattner FR, Jiang J ( 2002). Functional rice centromeres are marked by a satellite repeat and a centromere-specific retrotransposon. Plant Cell 14, 1691-1704. |
| [5] | Cheng Z, Stupar RM, Gu M, Jiang J ( 2001 b). A tandemly repeated DNA sequence is associated with both knob-like heterochromatin and a highly decondensed structure in the meiotic pachytene chromosomes of rice. Chromosoma 110, 24-31. |
| [6] | Goff SA ( 1999). Rice as a model for cereal genomics. Curr Opin Plant Biol 2, 86-89. |
| [7] | Ji JH, Tang D, Shen Y, Xue ZH, Wang HJ, Shi WQ, Zhang C, Du GJ, Li YF, Cheng ZK ( 2016). P31 comet, a member of the synaptonemal complex, participates in meiotic DSB formation in rice . Proc Natl Acad Sci USA 113, 10577-10582. |
| [8] | Kurata N, Omura T, Iwata N ( 1981). Studies on centromere, chromomere and nucleolus in pachytene nuclei of rice, Oryza sativa, microsporocytes. Cytologia 46, 791-800. |
| [9] | Li X, Chao D, Wu Y, Huang X, Chen K, Cui L, Su L, Ye W, Chen H, Chen H, Dong N, Guo T, Shi M, Feng Q, Zhang P, Han B, Shan J, Gao J, Lin H ( 2015). Natural alleles of a proteasome α2 subunit gene contribute to thermotolerance and adaptation of African rice. Nat Genet 47, 827-833. |
| [10] | Li YF, Qin BX, Shen Y, Zhang FF, Liu CZ, You HL, Du GJ, Tang D, Cheng ZK ( 2018). HEIP1 regulates crossover formation during meiosis in rice. Proc Natl Acad Sci USA 115, 10810-10815. |
| [11] | Luo Q, Li YF, Shen Y, Cheng ZK ( 2014). Ten years of gene discovery for meiotic event control in rice. J Genet Genomics 41, 125-137. |
| [12] | Ma Y, Dai X, Xu Y, Luo W, Zheng X, Zeng D, Pan Y, Lin X, Liu H, Zhang D, Xiao J, Guo X, Xu S, Niu Y, Jin J, Zhang H, Xu X, Li L, Wang W, Qian Q, Ge S, Chong K ( 2015). COLD1 confers chilling tolerance in rice. Cell 160, 1209-1221. |
| [13] | Miao CB, Tang D, Zhang HG, Wang M, Li YF, Tang SZ, Yu HX, Gu MH, Cheng ZK ( 2013). CENTRAL REGION COMPONENT 1, a novel synaptonemal complex component, is essential for meiotic recombination initiation in rice. Plant Cell 25, 2998-3009. |
| [14] | Nonomura K, Morohoshi A, Nakano M, Eiguchi M, Miyao A, Hirochika H, Kurata N ( 2007). A germ cell specific gene of the ARGONAUTE family is essential for the progression of premeiotic mitosis and meiosis during sporogenesis in rice. Plant Cell 19, 2583-2594. |
| [15] | Pawlowski WP, Golubovskaya IN, Timofejeva L, Meeley RB, Sheridan WF, Cande WZ ( 2004). Coordination of meiotic recombination, pairing, and synapsis by PHS1. Science 303, 89-92. |
| [16] | Ren LJ, Tang D, Zhao TT, Zhang FF, Liu CZ, Xue ZH, Shi WQ, Du GJ, Shen Y, Li YF, Cheng ZK ( 2018). OsSPL regulates meiotic fate acquisition in rice. New Phytol 218, 789-803. |
| [17] | Tang X, Bao W, Zhang W, Cheng Z ( 2007). Identification of chromosomes from multiple rice genomes using a universal molecular cytogenetic marker system. J Integr Plant Biol 49, 953-960. |
| [18] | Wang K, Tang D, Wang M, Lu J, Yu H, Liu J, Qian B, Gong Z, Wang X, Chen J, Gu M, Cheng Z ( 2009). MER3 is required for normal meiotic crossover formation, but not for presynaptic alignment in rice. J Cell Sci 122, 2055-2063. |
| [19] | Wang M, Wang K, Tang D, Wei C, Li M, Shen Y, Chi Z, Gu M, Cheng Z ( 2010). The central element protein ZEP1 of the synaptonemal complex regulates the number of crossovers during meiosis in rice. Plant Cell 22, 417-430. |
| [20] | Wang Y, Xiong G, Hu J, Jiang L, Yu H, Xu J, Fang Y, Zeng L, Xu E, Xu J, Ye W, Meng X, Liu R, Chen H, Jing Y, Wang Y, Zhu X, Li J, Qian Q ( 2015). Copy number variation at the GL7 locus contributes to grain size diversity in rice. Nat Genet 47, 944-948. |
| [21] | Wu HK ( 1967). Note on preparing of pachytene chromosomes by double mordant. Sci Agric 15, 40-44. |
| [22] | Yu H, Wang M, Tang D, Wang K, Chen F, Gong Z, Gu M, Cheng Z ( 2010). OsSPO11-1 is essential for both homologous chromosome pairing and crossover formation in rice. Chromosoma 119, 625-636. |
| [23] | Zhang F, Tang D, Shen Y, Xue ZH, Shi WQ, Ren LJ, Du GJ, Li Y, Cheng ZK ( 2017). The F-box protein ZYGO1 mediates bouquet formation to promote homologous pairing, synapsis, and recombination in rice meiosis. Plant Cell 29, 2597-2609. |
| [24] | Zhao TT, Ren LJ, Chen XJ, Yu HX, Liu CJ, Shen Y, Shi WQ, Tang D, Du GJ, Li YF, Ma BJ, Cheng ZK ( 2018). The OsRR24/LEPTO1 type-B response regulator is essential for the organization of leptotene chromosomes in rice meiosis. Plant Cell 30, 3024-3037. |
| [1] | Bao Zhu, Jiangzhe Zhao, Kewei Zhang, Peng Huang. Oryza sativa CYTOKININ OXIDASE 9(OsCKX9)Is Involved in Regulating the Rice Lamina Joint Development and Leaf Angle [J]. Chinese Bulletin of Botany, 2024, 59(1): 0-0. |
| [2] | Yanli Fang, Chuanyu Tian, Ruyi Su, Yapei Liu, Chunlian Wang, Xifeng Chen, Wei Guo, Zhiyuan Ji. Mining and Preliminary Mapping of Rice Resistance Genes against Bacterial Leaf Streak [J]. Chinese Bulletin of Botany, 2024, 59(1): 0-0. |
| [3] | Cailian Liu, Qing Xu, Linlong Wang, Yankuo Xing, Jiahao Song, Baian Lin, Bin Kang, Min Liu. Nekton diversity, density, and community structure of spring and autumn in coastal waters of eastern Fujian Province [J]. Biodiv Sci, 2023, 31(7): 22635-. |
| [4] | Zhiyuan Dong, Linlin Chen, Naipeng Zhang, Li Chen, Debin Sun, Yanmei Ni, Baoquan Li. Response of fish diversity to hydrological connectivity of typical tidal creek system in the Yellow River Delta based on environmental DNA metabarcoding [J]. Biodiv Sci, 2023, 31(7): 23073-. |
| [5] | Tian Chuanyu, Fang Yanli, Shen Qing, Wang Hongjie, Chen Xifeng, Guo Wei, Zhao Kaijun, Wang Chunlian, Ji Zhiyuan. Genotypic Diversity and Pathogenisity of Xanthomonas oryzae pv. oryzae Isolated from Southern China in 2019-2021 [J]. Chinese Bulletin of Botany, 2023, 58(5): 743-749. |
| [6] | Dai Ruohui, Qian Xinyu, Sun Jinglei, Lu Tao, Jia Qiwei, Lu Tianqi, Lu Mei, Rao Yuchun. Research Progress on the Mechanisms of Leaf Color Regulation and Related Genes in Rice [J]. Chinese Bulletin of Botany, 2023, 58(5): 799-812. |
| [7] | Shusen Fu, Puqing Song, Yuan Li, Yuanyuan Li, Ran Zhang, Hushun Zhang, Rui Wang, Longshan Lin. Trophic levels and trophic niches of fish from the Bering Sea and Chukchi Sea [J]. Biodiv Sci, 2023, 31(4): 22521-. |
| [8] | Shang Sun, Yingying Hu, Yangshuo Han, Chao Xue, Zhiyun Gong. Double-stranded Labelled Oligo-FISH in Rice Chromosomes [J]. Chinese Bulletin of Botany, 2023, 58(3): 433-439. |
| [9] | Jiayi Jin, Yiting Luo, Huimin Yang, Tao Lu, Hanfei Ye, Jiyi Xie, Kexin Wang, Qianyu Chen, Yuan Fang, Yuexing Wang, Yuchun Rao. QTL Mapping and Expression Analysis on Candidate Genes Related to Chlorophyll Content in Rice [J]. Chinese Bulletin of Botany, 2023, 58(3): 394-403. |
| [10] | Yuping Yan, Xiaoqi Yu, Deyong Ren, Qian Qian. Genetic Mechanisms and Breeding Utilization of Grain Number Per Panicle in Rice [J]. Chinese Bulletin of Botany, 2023, 58(3): 359-372. |
| [11] | Yuqiang Liu, Jianmin Wan. The Host Controls the Protein Level of Insect Effectors to Balance Immunity and Growth [J]. Chinese Bulletin of Botany, 2023, 58(3): 353-355. |
| [12] | Jingjing Zhao, Haibin Jia, Tien Ming Lee. Market status and the sustainable utilization strategy of wild earthworm (earth dragon) for medicinal use [J]. Biodiv Sci, 2023, 31(3): 22478-. |
| [13] | Wei Zhang, Dongdong Zhai, Fei Xiong, Hongyan Liu, Yuanyuan Chen, Ying Wang, Chuansong Liao, Xinbin Duan, Huiwu Tian, Huatang Deng, Daqing Chen. Community structure and functional diversity of fishes in the Three Gorges Reservoir [J]. Biodiv Sci, 2023, 31(2): 22136-. |
| [14] | Qi Wang, Yunzhe Wu, Xueying Liu, Lili Sun, Hong Liao, Xiangdong Fu. The Rice Receptor-like Kinases Function as Key Regulators of Plant Development and Adaptation to the Environment [J]. Chinese Bulletin of Botany, 2023, 58(2): 199-213. |
| [15] | Fanfan Zhang, Xinying Xing, Wenqing Shi, Yi Shen, Zhukuan Cheng. Protocals for Oligonucleotide Fluorescence in situ Hybridization in Plants [J]. Chinese Bulletin of Botany, 2023, 58(2): 274-284. |
| Viewed | ||||||
|
Full text |
|
|||||
|
Abstract |
|
|||||