

植物学报 ›› 2017, Vol. 52 ›› Issue (5): 590-597.DOI: 10.11983/CBB16137
收稿日期:2016-06-21
接受日期:2017-01-10
出版日期:2017-09-01
发布日期:2017-07-10
通讯作者:
包颖
基金资助:Received:2016-06-21
Accepted:2017-01-10
Online:2017-09-01
Published:2017-07-10
Contact:
Ying Bao
摘要: 抗细胞凋亡基因(DAD)是一个高度保守的细胞凋亡抑制基因, 在植物生长发育中承担重要功能。为全面了解DAD基因在种子植物中的分布和演化规律, 该文利用31种植物的全基因组数据, 通过生物信息学手段, 深入探讨和分析了不同植物类群中DAD基因的拷贝数目、基因结构和染色体定位, 并综合另外7种裸子植物的转录组数据探讨了其在种子植物中的演化趋势。结果表明, DAD基因属于低拷贝基因, 在不同种子植物中只具有1-3个拷贝; 不同DAD基因编码的氨基酸长度在108-170 aa之间变动。同线性和系统发育分析进一步表明, 种子植物DAD基因的演化具有明显的谱系特异性。随机复制和染色体大片段复制及其随后的基因丢失可能是其维持低拷贝的重要方式。
包颖, 梅玉芹. 种子植物抗细胞凋亡DAD基因的演化. 植物学报, 2017, 52(5): 590-597.
Ying Bao, Yuqin Mei. Evolution of Defender Against Apoptotic Death (DAD) Genes in Seed Plants. Chinese Bulletin of Botany, 2017, 52(5): 590-597.
| Taxa | Gene ID | Strand | Chromosome | Duplication pattern | Gene structure | Amino acid (aa) | |
|---|---|---|---|---|---|---|---|
| Intron | Exon | ||||||
| Angiosperm | |||||||
| Dicot | |||||||
| Arabidopsis lyrata | AL1G33450 | _ | Scaffold_1 | Block | 4 | 5 | 115 |
| AL4G20890 | + | Scaffold_4 | Block | 4 | 5 | 116 | |
| A. thaliana | AT1G32210 | _ | Chr01 | Block | 4 | 5 | 115 |
| AT2G35520 | + | Chr02 | Block | 4 | 5 | 116 | |
| Brassica rapa | BR05G20160 | + | ChrA05 | Block | 4 | 5 | 115 |
| BR09G26700a | + | ChrA09 | Block | 6 | 7 | 201 (88) d | |
| Capsella rubella | CRU_001G29110 | _ | Scaffold_1 | Block | 4 | 5 | 115 |
| CRU_004G16620 | + | Scaffold_4 | Block | 4 | 5 | 115 | |
| Citrullus lanatus | CL10G00840 | + | Chr10 | 4 | 5 | 115 | |
| Cucumis melo | CM00021G01290 | _ | Scaffold00021 | 4 | 5 | 115 | |
| Eucalyptus grandis | EG0008G05930 | + | Scaffold_8 | 4 | 5 | 115 | |
| Fragaria vesca | FV2G07670 | + | LG2 | 4 | 5 | 124 | |
| Gossypium raimondii | GR03G18540 | + | Chr03 | Block | 4 | 5 | 117 |
| GR08G22400 | _ | Chr08 | Block | 4 | 5 | 117 | |
| Malus domestica | MD05G025840 | + | Chr05 | Random | 4 | 5 | 119 |
| MD10G000120 | + | Chr10 | Random | 4 | 5 | 119 | |
| Manihot esculenta | ME04430G00010 | + | Scaffold04430 | Random | 4 | 5 | 115 |
| ME07304G00010 | + | Scaffold07304 | Random | 4 | 5 | 115 | |
| Populus trichocarpa | PT01G13680 | _ | Chr01 | Block | 4 | 5 | 115 |
| PT03G09680 | + | Chr03 | Block | 4 | 5 | 115 | |
| Prunus persica | PPE_004G33460 | _ | Scaffold_4 | Block | 4 | 5 | 119 |
| PPE_008G00790 | _ | Scaffold_8 | Block | 4 | 5 | 119 | |
| Ricinus communis | RC29634G00390 | + | 29634 | Random | 4 | 5 | 115 |
| RC30068G01440 | + | 30068 | Random | 4 | 5 | 113 | |
| Solanum lycopersicum | SL08G076460 | + | Chr08 | 4 | 5 | 116 | |
| S. tuberosum | ST08G008690 | _ | Chr08 | Block | 4 | 5 | 116 |
| ST08G021840 | _ | Chr08 | Block | 4 | 5 | 116 | |
| Thellungiella parvula | TP1G27890 | _ | Chr1-1 | 4 | 5 | 115 | |
| Theobroma cacao | TC0003G30500 | + | Scaffold_3 | 4 | 5 | 116 | |
| Vitis vinifera | VV02G01690 | + | Chr02 | 4 | 5 | 115 | |
| Moncot | |||||||
| Brachypodium distachyon | BD1G50180 | + | Chr01 | 4 | 5 | 114 | |
| Hordeum vulgare | HV1571041G00020 | _ | Contig_1571041 | Random | 4 | 5 | 114 |
| HV44460G00030 | + | Contig_44460 | Random | 4 | 5 | 114 | |
| Oryza sativa | OS04G32550 | + | Chr04 | 4 | 5 | 114 | |
| Setaria italica | SI004G00880 | + | Scaffold_4 | 4 | 5 | 114 | |
| Sorghum bicolor | SB10G001000 | + | Chr10 | 4 | 5 | 114 | |
| Zea mays | ZM09G06480 | _ | Chr09 | 4 | 5 | 114 | |
| Musa acuminata | MA07G17850 | _ | Chr07 | Block | 4 | 5 | 115 |
| MA10G06540b | _ | Chr10 | Block | - | 1 | 48 (22) d | |
| MA11G08020 | + | Chr11 | Block | 4 | 5 | 170 | |
| Basal taxon | |||||||
| Amborella trichopoda | ATR_00025G00360 | + | Scaffold00025 | 4 | 5 | 121 | |
| Gymnosperm | |||||||
| Taxa | |||||||
| Ginkgo bilobac Gnetum montanumc | Gene ID | Strand | Chromosome | Duplication pattern | Gene structure | Amino acid (aa) | |
| Intron | Exon | ||||||
| Picea abies | GBI00024938 | + | 6973 | - | - | 123 | |
| GMO00028447 | _ | GTHK-0066600 | - | - | 113 | ||
| PAB00021420a | _ | MA_128105 | Random | 5 | 6 | 162 (67) d | |
| P. glaucac | PAB00041636 | _ | MA_42912 | Random | 4 | 5 | 115 |
| PAB00057214b | + | MA_8328929 | Random | 1 | 2 | 50 (45) d | |
| PGL00020840 | + | PUT-39823 PUT-39823 | - | - | 115 | ||
| P. sitchensisc | PGL00013662 | + | PUT-23726 PUT-23726 | - | - | 115 | |
| PGL00010558 | + | PUT-16509 | - | - | 115 | ||
| Pinus pinasterc | PSI00016644 | + | PUT-531483 | - | - | 148 | |
| PSI00008404 | + | PUT-21837 | - | - | 115 | ||
| Pi. sylvestrisc | PPI00061658 | _ | Unigene30039 | - | - | 115 | |
| PPI00006515 | _ | Cotig25413 | - | - | 115 | ||
| PSY00006071 | _ | Isotig31028 | - | - | 115 | ||
| Pi. taeda | PSY00026421 | + | Isotig68699 | - | - | 115 | |
| PSY00004335 | + | isotig24244 | - | - | 115 | ||
| PTA00003657 | _ | Scaffold464 | Random | 4 | 5 | 115 | |
| Pseudotsuga menziesii c | PTA00022345b | + | Scaffold214279 | Random | 2 | 3 | 86 (72) d |
| PTA00022344b | + | Scaffold214279 | Random | 1 | 2 | 44 (44) d | |
| Moss Physcomitrella patens | PME00105748 | + | Psme_598296871 | - | - | 146 | |
| PME00105747 | + | Psme_598296869 | - | - | 115 | ||
| Alga Chlamydomonas reinhardtii | PP00045G01180 | + | Scaffold_45 | 4 | 5 | 131 | |
| PP00456G00180 | - | Scaffold_456 | 4 | 5 | 114 | ||
表1 38种植物DAD同源基因的详细信息
Table 1 Detailed information of DAD homologous genes of 38 plants
| Taxa | Gene ID | Strand | Chromosome | Duplication pattern | Gene structure | Amino acid (aa) | |
|---|---|---|---|---|---|---|---|
| Intron | Exon | ||||||
| Angiosperm | |||||||
| Dicot | |||||||
| Arabidopsis lyrata | AL1G33450 | _ | Scaffold_1 | Block | 4 | 5 | 115 |
| AL4G20890 | + | Scaffold_4 | Block | 4 | 5 | 116 | |
| A. thaliana | AT1G32210 | _ | Chr01 | Block | 4 | 5 | 115 |
| AT2G35520 | + | Chr02 | Block | 4 | 5 | 116 | |
| Brassica rapa | BR05G20160 | + | ChrA05 | Block | 4 | 5 | 115 |
| BR09G26700a | + | ChrA09 | Block | 6 | 7 | 201 (88) d | |
| Capsella rubella | CRU_001G29110 | _ | Scaffold_1 | Block | 4 | 5 | 115 |
| CRU_004G16620 | + | Scaffold_4 | Block | 4 | 5 | 115 | |
| Citrullus lanatus | CL10G00840 | + | Chr10 | 4 | 5 | 115 | |
| Cucumis melo | CM00021G01290 | _ | Scaffold00021 | 4 | 5 | 115 | |
| Eucalyptus grandis | EG0008G05930 | + | Scaffold_8 | 4 | 5 | 115 | |
| Fragaria vesca | FV2G07670 | + | LG2 | 4 | 5 | 124 | |
| Gossypium raimondii | GR03G18540 | + | Chr03 | Block | 4 | 5 | 117 |
| GR08G22400 | _ | Chr08 | Block | 4 | 5 | 117 | |
| Malus domestica | MD05G025840 | + | Chr05 | Random | 4 | 5 | 119 |
| MD10G000120 | + | Chr10 | Random | 4 | 5 | 119 | |
| Manihot esculenta | ME04430G00010 | + | Scaffold04430 | Random | 4 | 5 | 115 |
| ME07304G00010 | + | Scaffold07304 | Random | 4 | 5 | 115 | |
| Populus trichocarpa | PT01G13680 | _ | Chr01 | Block | 4 | 5 | 115 |
| PT03G09680 | + | Chr03 | Block | 4 | 5 | 115 | |
| Prunus persica | PPE_004G33460 | _ | Scaffold_4 | Block | 4 | 5 | 119 |
| PPE_008G00790 | _ | Scaffold_8 | Block | 4 | 5 | 119 | |
| Ricinus communis | RC29634G00390 | + | 29634 | Random | 4 | 5 | 115 |
| RC30068G01440 | + | 30068 | Random | 4 | 5 | 113 | |
| Solanum lycopersicum | SL08G076460 | + | Chr08 | 4 | 5 | 116 | |
| S. tuberosum | ST08G008690 | _ | Chr08 | Block | 4 | 5 | 116 |
| ST08G021840 | _ | Chr08 | Block | 4 | 5 | 116 | |
| Thellungiella parvula | TP1G27890 | _ | Chr1-1 | 4 | 5 | 115 | |
| Theobroma cacao | TC0003G30500 | + | Scaffold_3 | 4 | 5 | 116 | |
| Vitis vinifera | VV02G01690 | + | Chr02 | 4 | 5 | 115 | |
| Moncot | |||||||
| Brachypodium distachyon | BD1G50180 | + | Chr01 | 4 | 5 | 114 | |
| Hordeum vulgare | HV1571041G00020 | _ | Contig_1571041 | Random | 4 | 5 | 114 |
| HV44460G00030 | + | Contig_44460 | Random | 4 | 5 | 114 | |
| Oryza sativa | OS04G32550 | + | Chr04 | 4 | 5 | 114 | |
| Setaria italica | SI004G00880 | + | Scaffold_4 | 4 | 5 | 114 | |
| Sorghum bicolor | SB10G001000 | + | Chr10 | 4 | 5 | 114 | |
| Zea mays | ZM09G06480 | _ | Chr09 | 4 | 5 | 114 | |
| Musa acuminata | MA07G17850 | _ | Chr07 | Block | 4 | 5 | 115 |
| MA10G06540b | _ | Chr10 | Block | - | 1 | 48 (22) d | |
| MA11G08020 | + | Chr11 | Block | 4 | 5 | 170 | |
| Basal taxon | |||||||
| Amborella trichopoda | ATR_00025G00360 | + | Scaffold00025 | 4 | 5 | 121 | |
| Gymnosperm | |||||||
| Taxa | |||||||
| Ginkgo bilobac Gnetum montanumc | Gene ID | Strand | Chromosome | Duplication pattern | Gene structure | Amino acid (aa) | |
| Intron | Exon | ||||||
| Picea abies | GBI00024938 | + | 6973 | - | - | 123 | |
| GMO00028447 | _ | GTHK-0066600 | - | - | 113 | ||
| PAB00021420a | _ | MA_128105 | Random | 5 | 6 | 162 (67) d | |
| P. glaucac | PAB00041636 | _ | MA_42912 | Random | 4 | 5 | 115 |
| PAB00057214b | + | MA_8328929 | Random | 1 | 2 | 50 (45) d | |
| PGL00020840 | + | PUT-39823 PUT-39823 | - | - | 115 | ||
| P. sitchensisc | PGL00013662 | + | PUT-23726 PUT-23726 | - | - | 115 | |
| PGL00010558 | + | PUT-16509 | - | - | 115 | ||
| Pinus pinasterc | PSI00016644 | + | PUT-531483 | - | - | 148 | |
| PSI00008404 | + | PUT-21837 | - | - | 115 | ||
| Pi. sylvestrisc | PPI00061658 | _ | Unigene30039 | - | - | 115 | |
| PPI00006515 | _ | Cotig25413 | - | - | 115 | ||
| PSY00006071 | _ | Isotig31028 | - | - | 115 | ||
| Pi. taeda | PSY00026421 | + | Isotig68699 | - | - | 115 | |
| PSY00004335 | + | isotig24244 | - | - | 115 | ||
| PTA00003657 | _ | Scaffold464 | Random | 4 | 5 | 115 | |
| Pseudotsuga menziesii c | PTA00022345b | + | Scaffold214279 | Random | 2 | 3 | 86 (72) d |
| PTA00022344b | + | Scaffold214279 | Random | 1 | 2 | 44 (44) d | |
| Moss Physcomitrella patens | PME00105748 | + | Psme_598296871 | - | - | 146 | |
| PME00105747 | + | Psme_598296869 | - | - | 115 | ||
| Alga Chlamydomonas reinhardtii | PP00045G01180 | + | Scaffold_45 | 4 | 5 | 131 | |
| PP00456G00180 | - | Scaffold_456 | 4 | 5 | 114 | ||
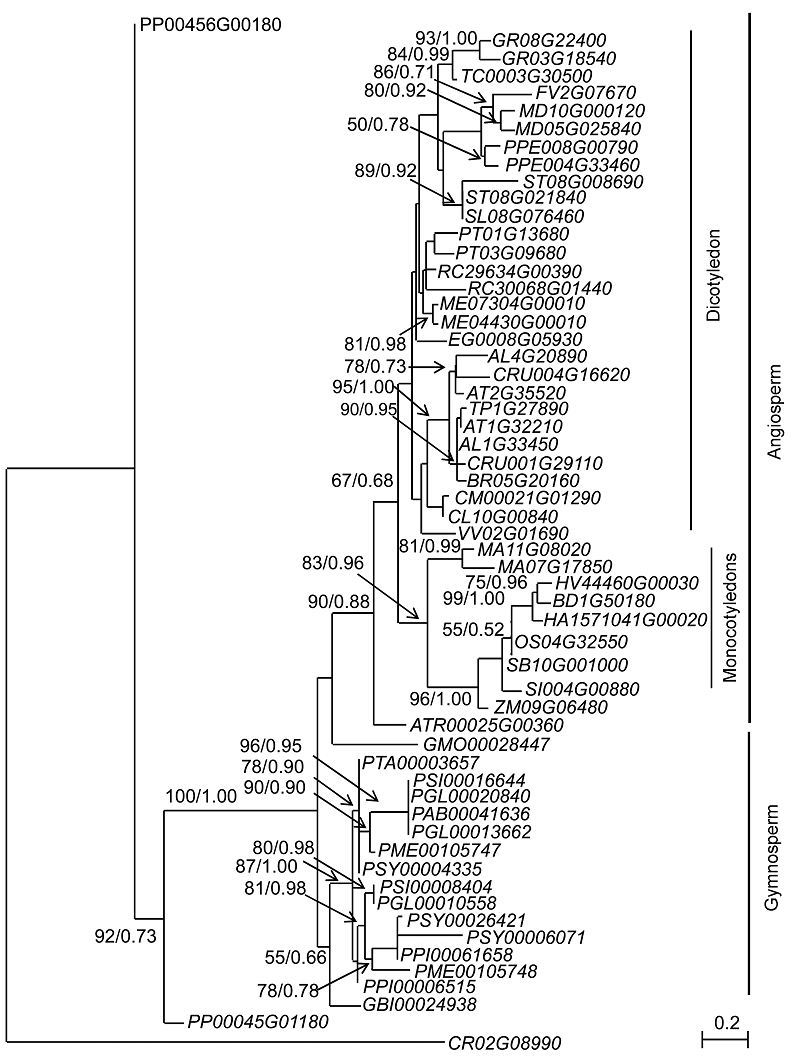
图1 基于38种植物的58条DAD基因的编码序列构建的最大似然性系统发育关系树分支附近的数值分别代表最大似然性分析中大于50%的自展支持率和贝叶斯分析中的后验率, 各基因位点同表1。
Figure 1 Maximum likelihood tree of DAD genes based on 58 amino acid sequences of 38 plantsNumbers near branches represent bootstrap values (>50%) of the maximum likelihood analysis and posterior rate of the Bayesian analysis. The gene loci in this figure are the same as those listed in Table 1.
| [1] | 龚文芳, 喻树迅, 宋美珍, 范术丽, 庞朝友, 肖水平 (2010). 棉花抗细胞凋亡基因GhDAD1的克隆、定位及表达分析. 中国农业科学 43, 3713-3723. |
| [2] | 贾志蓉, 李丹, 吕应堂 (2004). 拟南芥AtDAD1超量表达植株对H2O2抗性的研究. 武汉植物学研究 22, 373-379. |
| [3] | 杨舒雅, 史娟, 马斌芳, 赵洁, 金晓航 (2012). 抗细胞凋亡因子DAD1的研究进展. 生理科学进展 43, 315-318. |
| [4] | Apte SS, Mattei MG, Seldin MF, Olsen BR (1995). The highly conserved defender against the death 1 (DAD1) gene maps to human chromosome 14q11-q12 and mouse chromosome 14 and has plant and nematode homologs. FEBS Lett 363, 304-306. |
| [5] | Blanc G, Hokamp K, Wolfe KH (2003). A recent polyploidy superimposed on older large-scale duplications in the Arabidopsis genome.Genome Res 13, 137-144. |
| [6] | Danon A, Rotari VI, Gordon A, Mailhac N, Gallois P (2004). Ultraviolet-C overexposure induces programmed cell death in Arabidopsis, which is mediated by caspase- like activities and which can be suppressed by caspase inhibitors, p35 and defender against apoptotic death.J Biol Chem 279, 779-787. |
| [7] | Gallois P, Makishima T, Hecht V, Despres B, Laudié M, Nishimoto T, Cooke R (1997). An Arabidopsis thaliana cDNA complementing a hamster apoptosis suppressor mutant.Plant J 11, 1325-1331. |
| [8] | Gouy M, Guindon S, Gascuel O (2010). SeaView version 4: a multiplatform graphical user interface for sequence alignment and phylogenetic tree building.Mol Biol Evol 27, 221-224. |
| [9] | Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010). New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0.Syst Biol 59, 307-321. |
| [10] | Kelleher DJ, Gilmore R (1997). DAD1, the defender against apoptotic cell death, is a subunit of the mammalian oligosaccharyltransferase.Proc Natl Acad Sci USA 94, 4994-4999. |
| [11] | Kerr JFR, Wyllie AH, Currie AR (1972). Apoptosis: a basic biological phenomenon with wide ranging implications in tissue kinetics.Br J Cancer 26, 239-257. |
| [12] | Mondragón-Palomino M, Meyers BC, Michelmore RW, Gaut BS (2002). Patterns of positive selection in the com- plete NBS-LRR gene family of Arabidopsis thaliana.Genome Res 12, 1305-1315. |
| [13] | Nakashima T, Sekiguchi T, Kuraoka A, Fukushima K, Shibata Y, Komiyama S, Nishimoto T (1993). Molecular cloning of a human cDNA encoding a novel protein, DAD1, whose defect causes apoptotic cell death in hamster BHK21 cells.Mol Cell Biol 13, 6367-6374. |
| [14] | Raff M (1998). Cell suicide for beginners.Nature 396, 119-122. |
| [15] | Roboti P, High S (2012). The oligosaccharyltransferase subunits OST48, DAD1 and KCP2 function as ubiquitous and selective modulators of mammalian N-glycosylation.J Cell Sci 125, 3474-3484. |
| [16] | Ronquist F, Huelsenbeck JP (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models.Bioinformatics 19, 1572-1574. |
| [17] | Schwartzman RA, Cidlowski JA (1993). Apoptosis: the biochemistry and molecular biology of programmed cell death.Endocr Rev 14, 133-151. |
| [18] | Williams GT, Smith CA (1993). Molecular regulation of apoptosis: genetic controls on cell death.Cell 7, 777-779. |
| [19] | Wortman JR, Haas BJ, Hannick LI, Smith RK, Maiti R, Ronning CM, Chan AP, Yu CH, Ayele M, Whitelaw CA, White OR, Town CD (2003). Annotation of the Arabidopsis genome.Plant Physiol 132, 461-468. |
| [20] | Ye CY, Li T, Yin H, Weston DJ, Tuskan GA, Tschaplinski TJ, Yang X (2012). Evolutionary analyses of non-family genes in plants.Plant J 73, 788-797. |
| [1] | 景昭阳, 程可光, 舒恒, 马永鹏, 刘平丽. 全基因组重测序方法在濒危植物保护中的应用[J]. 生物多样性, 2023, 31(5): 22679-. |
| [2] | 武棒棒, 郝宇琼, 杨淑斌, 黄雨茜, 关攀锋, 郑兴卫, 赵佳佳, 乔玲, 李晓华, 刘维仲, 郑军. 山西小麦籽粒叶黄素含量变异及遗传特性分析[J]. 植物学报, 2023, 58(4): 535-547. |
| [3] | 孙蓉, 杨宇琭, 李亚军, 张会, 李旭凯. 谷子PLATZ转录因子基因家族的鉴定和分析[J]. 植物学报, 2023, 58(4): 548-559. |
| [4] | 张嘉, 李启东, 李翠, 王庆海, 侯新村, 赵春桥, 李树和, 郭强. 植物MATE转运蛋白研究进展[J]. 植物学报, 2023, 58(3): 461-474. |
| [5] | 陶洁, 李本强, 程靖华, 石迎, 刘佩红, 何桂霞, 徐伟杰, 刘惠莉. 基于宏病毒组学技术挖掘猪腹泻粪便的病毒群落组成[J]. 生物多样性, 2023, 31(11): 23170-. |
| [6] | 罗瑞, 陈娅, 张汉马. 芸薹属植物全基因组重测序研究进展[J]. 生物多样性, 2023, 31(10): 23237-. |
| [7] | 李园, 常玉洁, 王兰芬, 王述民, 武晶. 普通菜豆镰孢菌枯萎病抗性种质资源筛选及全基因组关联分析[J]. 植物学报, 2023, 58(1): 51-61. |
| [8] | 李晓明, 王兰芬, 唐永生, 常玉洁, 张菊香, 王述民, 武晶. 普通菜豆抗菜豆象性状的全基因组关联分析[J]. 植物学报, 2023, 58(1): 77-89. |
| [9] | 金京波, 梁承志. 饲草基因组学研究进展[J]. 植物学报, 2022, 57(6): 732-741. |
| [10] | 翟俊杰, 赵慧峰, 商光申, 孙振钧, 张玉峰, 王兴. 蚯蚓基因组学的研究进展: 基于全基因组及线粒体基因组[J]. 生物多样性, 2022, 30(12): 22257-. |
| [11] | 陆奇丰, 黄至欢, 骆文华. 极小种群濒危植物广西火桐、丹霞梧桐的叶绿体基因组特征[J]. 生物多样性, 2021, 29(5): 586-595. |
| [12] | 宣伟, 徐国华. 植物适应土壤氮素环境的基因选择: 以水稻为例[J]. 植物学报, 2021, 56(1): 1-5. |
| [13] | 赵宇慧, 李秀秀, 陈倬, 鲁宏伟, 刘羽诚, 张志方, 梁承志. 生物信息学分析方法I: 全基因组关联分析概述[J]. 植物学报, 2020, 55(6): 715-732. |
| [14] | 闫晨阳,陈赢男. 4种模式植物LRR VIII-2亚家族基因的鉴定和进化历史分析[J]. 植物学报, 2020, 55(4): 442-456. |
| [15] | 汪浩, 张锐, 张娇, 沈慧, 戴锡玲, 严岳鸿. 转录组测序揭示翼盖蕨(Didymochlaena trancatula)的全基因组复制历史[J]. 生物多样性, 2019, 27(11): 1221-1227. |
| 阅读次数 | ||||||
|
全文 |
|
|||||
|
摘要 |
|
|||||
